Unraveling the Catalytic Power of MsrB1: The Unique Selenoprotein Redox Mechanism Driving Cellular Repair and Disease Prevention
This article provides a comprehensive analysis of the catalytic mechanism of Methionine Sulfoxide Reductase B1 (MsrB1), a critical selenoprotein enzyme in cellular antioxidant defense.
Unraveling the Catalytic Power of MsrB1: The Unique Selenoprotein Redox Mechanism Driving Cellular Repair and Disease Prevention
Abstract
This article provides a comprehensive analysis of the catalytic mechanism of Methionine Sulfoxide Reductase B1 (MsrB1), a critical selenoprotein enzyme in cellular antioxidant defense. Targeted at researchers, scientists, and drug development professionals, it explores the foundational biochemistry of its selenocysteine-dependent catalysis, methodologies for studying its activity, common experimental challenges, and comparative validation against other Msr isoforms. The synthesis aims to bridge mechanistic understanding with translational applications in redox biology and therapeutic development for age-related and oxidative stress disorders.
The Selenocysteine Core: Deconstructing the Unique Catalytic Mechanism of MsrB1
The Methionine Sulfoxide Reductase (Msr) system is a critical enzymatic defense network against oxidative stress, specifically repairing methionine sulfoxide (Met-O) residues in proteins back to methionine (Met). This system comprises two mechanistically distinct families: MsrA, which reduces the S-epimer of methionine sulfoxide (Met-S-O), and MsrB, which reduces the R-epimer (Met-R-O). MsrB1, a selenocysteine (Sec)-containing enzyme in humans, represents the most efficient and major mammalian MsrB isoform. This whitepaper, framed within broader thesis research on selenoprotein catalytic mechanisms, provides an in-depth technical analysis of MsrB1's unique role, kinetics, and experimental characterization within the Msr system.
Protein-bound methionine is highly susceptible to oxidation by reactive oxygen species (ROS), forming a mixture of S- and R- diastereomers of methionine sulfoxide. Unrepaired, these modifications can lead to protein misfunction, aggregation, and cellular damage. The Msr system provides stereospecific reduction to reverse this damage.
- MsrA: Primarily cytosolic and mitochondrial; reduces Met-S-O using a catalytic cysteine mechanism, with recycling via thioredoxin (Trx)/thioredoxin reductase (TrxR)/NADPH.
- MsrB (with focus on MsrB1): Primarily nuclear and cytosolic; reduces Met-R-O. MsrB1 is a selenoprotein, where the selenocysteine residue confers superior catalytic efficiency compared to its cysteine homologs (e.g., MsrB2/B3).
MsrB1 Catalytic Mechanism and Kinetics
The catalytic cycle of MsrB1 involves a selenenylsulfide intermediate. The mechanism is summarized as follows:
- Nucleophilic Attack: The catalytic selenolate (Sec-Se⁻) attacks the sulfur atom of substrate Met-R-O, forming a selenenylsulfide intermediate and releasing methionine.
- Resolution: The first resolving cysteine (Cys-X-Cys motif) attacks the selenenylsulfide, releasing the catalytic Sec as a selenenic acid (Sec-SeOH) and forming an intramolecular disulfide bond.
- Recycling: The disulfide bond is reduced by the thioredoxin system (Trx), regenerating the active enzyme. The selenenic acid intermediate is highly reactive and prone to overoxidation, which may require backup reduction systems like glutathione (GSH).
Quantitative Kinetic Data: MsrB1 vs. Other Msrs Table 1: Comparative kinetic parameters of human Msr enzymes.
| Enzyme | Substrate Stereospecificity | Catalytic Residue | kcat (min⁻¹) | KM (µM) | kcat/KM (M⁻¹s⁻¹) | Primary Reductant |
|---|---|---|---|---|---|---|
| MsrB1 (Sec) | Met-R-O | Selenocysteine (Sec) | ~80-120 | ~15-30 | ~5.0 x 10⁴ | Thioredoxin |
| MsrB1 (Cys mutant) | Met-R-O | Cysteine (Cys) | ~5-10 | ~50-100 | ~1.5 x 10³ | Thioredoxin |
| MsrA | Met-S-O | Cysteine (Cys) | ~50-80 | ~20-50 | ~2.0 x 10⁴ | Thioredoxin |
| MsrB2 | Met-R-O | Cysteine (Cys) | ~8-15 | ~30-60 | ~3.0 x 10³ | Thioredoxin |
Experimental Protocols for MsrB1 Research
Protocol 4.1: Recombinant MsrB1 Expression and Purification (E. coli)
- Objective: To produce pure, active selenocysteine-containing MsrB1.
- Methodology:
- Cloning: Clone the human MSRB1 gene, including the SECIS element, into a suitable expression vector (e.g., pET-based).
- Expression: Transform into an E. coli strain auxotrophic for selenium (e.g., BL21(DE3) ΔselA). Grow in LB medium supplemented with 50 µM sodium selenite at 37°C to OD600 ~0.6. Induce with 0.5 mM IPTG and grow overnight at 20°C.
- Purification: Lyse cells in buffer (50 mM Tris-HCl pH 7.5, 300 mM NaCl, 10 mM Imidazole) via sonication. Purify the His-tagged protein using Ni-NTA affinity chromatography. Elute with 250 mM imidazole. Further purify by size-exclusion chromatography (Superdex 75) in storage buffer (50 mM Tris-HCl pH 7.5, 150 mM NaCl, 1 mM DTT).
- Key Validation: Measure selenium content via ICP-MS; confirm activity via NADPH-coupled assay (see Protocol 4.2).
Protocol 4.2: NADPH-Coupled Msr Activity Assay
- Objective: To quantitatively determine MsrB1 enzymatic activity.
- Methodology:
- Reaction Mix: Prepare 1 mL containing: 50 mM Tris-HCl (pH 7.5), 150 mM NaCl, 0.5 mM NADPH, 5 µM E. coli Thioredoxin (Trx), 100 nM Thioredoxin Reductase (TrxR), and substrate (e.g., 1-5 mM dabsyl-Met-R-O or protein-bound Met-O).
- Enzyme Addition: Add purified MsrB1 (10-100 nM final) to initiate the reaction.
- Measurement: Monitor the oxidation of NADPH at 340 nm (ε340 = 6220 M⁻¹cm⁻¹) spectrophotometrically for 5-10 minutes at 37°C.
- Calculation: Activity is calculated from the linear decrease in absorbance. One unit of activity is defined as the oxidation of 1 µmol of NADPH per minute.
Protocol 4.3: Identification of MsrB1 Substrates via Redox Proteomics
- Objective: To identify endogenous protein targets of MsrB1 in cell lysates.
- Methodology:
- Oxidation and Treatment: Treat cell lysate with H₂O₂ (1-5 mM) to oxidize methionines. Divide into two aliquots.
- Reduction Reaction: Incubate one aliquot with recombinant MsrB1 (1 µM), DTT (5 mM), Trx/TrxR/NADPH system. The other aliquot is a no-enzyme control.
- CyDye Labeling: Label reduced protein thiols (from newly reduced methionines and existing cysteines) with maleimide-functionalized Cy5 (Msr-treated) or Cy3 (control) dyes.
- 2D-DIGE & MS: Run samples on 2D gel electrophoresis. Scan for fluorescent spots where Cy5/Cy3 ratio >2, indicating MsrB1-specific reduction. Excise spots, digest with trypsin, and identify proteins by LC-MS/MS.
Visualization of Pathways and Workflows
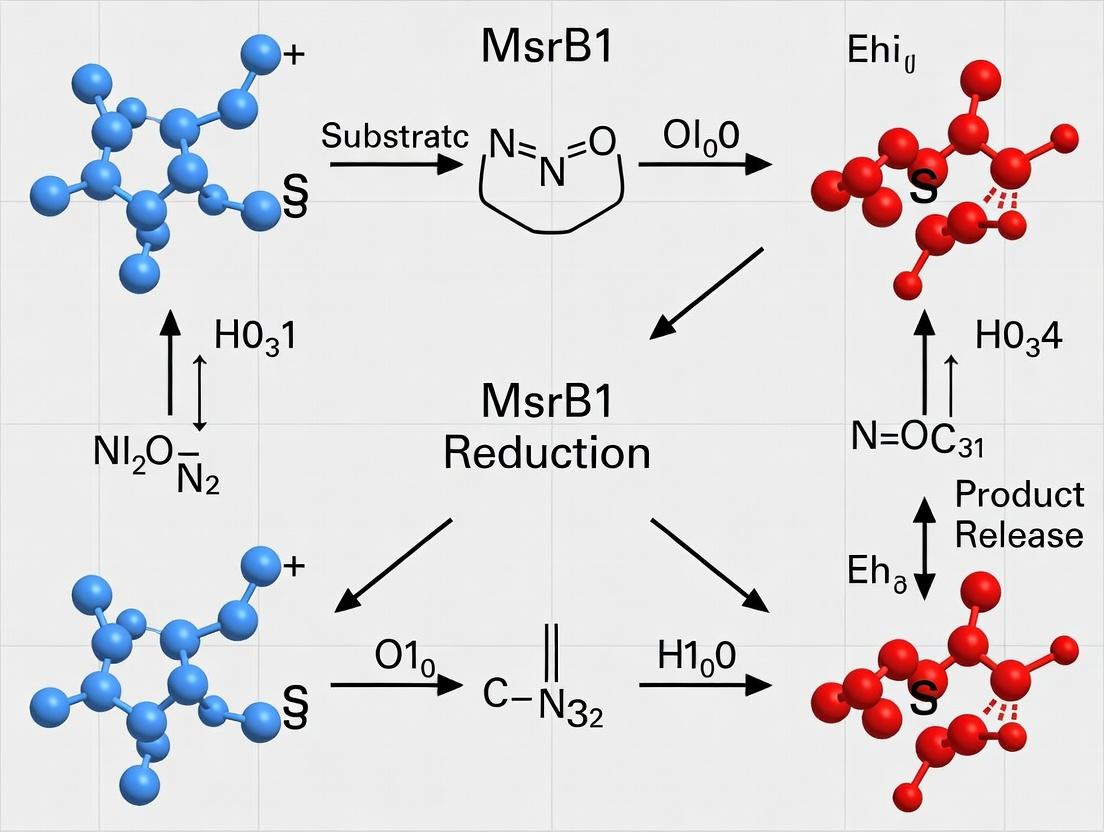
Diagram Title: MsrB1 Catalytic Cycle & Recycling Pathway
Diagram Title: Redox Proteomics Workflow for MsrB1 Substrates
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential materials for MsrB1 catalytic research.
| Reagent/Material | Function & Explanation | Example Vendor/Product |
|---|---|---|
| Recombinant Human MsrB1 (Sec) | Gold-standard enzyme source for kinetic studies and substrate identification. Must be produced with proper selenocysteine incorporation. | In-house expression (see Protocol 4.1) or commercial recombinant protein (e.g., MyBioSource). |
| Stereospecific Msr Substrates (e.g., Dabsyl-Met-R-O) | Synthetic small-molecule substrates to specifically measure MsrB1 activity without interference from MsrA. Crucial for kinetic characterization. | Sigma-Aldrich (custom synthesis) or Cayman Chemical. |
| Thioredoxin System Kit (Trx, TrxR, NADPH) | Provides the physiological electron donor for Msr enzyme recycling in activity assays. | Sigma-Aldrich (TRX0100) or IMCO Corporation. |
| Selenium Source (Sodium Selenite, Na₂SeO₃) | Essential supplement for culture media to produce selenocysteine-containing MsrB1 in recombinant expression systems. | Sigma-Aldrich (S5261). |
| Anti-MsrB1 Antibody (Selenocysteine-specific if possible) | For detection, localization (Western Blot, Immunofluorescence), and quantification of MsrB1 in cellular models and tissues. | Abcam (ab201966) or Santa Cruz Biotechnology (sc-514291). |
| Methionine Sulfoxide-Containing Model Protein (e.g., Oxidized CaM) | A well-characterized, physiologically relevant protein substrate for studying MsrB1 repair activity on complex structures. | Prepared in-lab by H₂O₂ oxidation of Calmodulin. |
| ICP-MS Standard (Selenium) | For inductively coupled plasma mass spectrometry quantification of selenium content in purified MsrB1, confirming Sec incorporation. | Agilent Technologies. |
| Maleimide-CyDyes (Cy3, Cy5) | Fluorescent dyes for labeling newly reduced cysteine/selenocysteine residues in redox proteomic workflows (2D-DIGE). | Cytiva (CyDye DIGE Fluor saturation dyes). |
Methionine sulfoxide reductase B1 (MsrB1) is a critical selenoenzyme responsible for the stereospecific reduction of methionine-R-sulfoxide back to methionine, a key antioxidant repair mechanism. The catalytic advantage of its selenocysteine (Sec, U) residue over a cysteine (Cys, C) is a focal point in understanding its high efficiency and physiological role. This analysis, framed within broader thesis research on the MsrB1 catalytic mechanism, details the physicochemical and kinetic rationale for Sec in this active site.
Physicochemical & Catalytic Advantages of Selenocysteine
Selenium's properties confer distinct biochemical advantages over sulfur in catalytic redox centers.
Table 1: Key Physicochemical Properties of Selenium vs. Sulfur in Biological Catalysis
| Property | Selenium (in Sec) | Sulfur (in Cys) | Catalytic Consequence for MsrB1 |
|---|---|---|---|
| Atomic Radius | ~120 pm | ~105 pm | Larger, more polarizable electron cloud. |
| pKa (R-SeH/R-SH) | ~5.2 (selenol) | ~8.3 (thiol) | Sec is deprotonated/ionized (selenolate, Se⁻) at physiological pH, primed for nucleophilic attack. |
| Redox Potential (E°') | More negative | Less negative | Sec is a stronger nucleophile and superior reducing agent. |
| Acid/Base Behavior | Stronger acid, weaker base | Weaker acid, stronger base | Facilitates proton transfer steps; stabilizes reaction intermediates. |
| Bond Strength (C–Se, Se–H) | Weaker than C–S, S–H | Stronger bonds | Lower bond dissociation energies enhance catalytic turnover. |
The lower pKa is paramount. At cellular pH ~7.4, the Sec selenol exists predominantly as the reactive selenolate anion (Se⁻), while Cys thiol is largely protonated. This pre-activation eliminates the need for an enzymatic base to deprotonate the residue, lowering the activation energy for the initial nucleophilic attack on the sulfoxide substrate.
Quantitative Kinetic Data: Sec vs. Cys MsrB1
Studies utilizing site-directed mutagenesis to replace Sec with Cys in MsrB1 (UxxC mutant) demonstrate dramatic kinetic consequences.
Table 2: Comparative Kinetic Parameters for Wild-Type (Sec) and Cys Mutant MsrB1
| Enzyme Variant | kcat (s⁻¹) | KM (µM) | kcat/KM (M⁻¹s⁻¹) | Catalytic Efficiency Relative to WT |
|---|---|---|---|---|
| Wild-Type (Sec) | 1.8 - 2.5 | 15 - 25 | ~1.2 x 10⁵ | 1 (Reference) |
| UxxC Mutant (Cys) | 0.02 - 0.05 | 80 - 120 | ~4.0 x 10² | ~0.003 (300-fold decrease) |
| Note: Values are representative ranges from recent stopped-flow and steady-state kinetics studies using dithiothreitol (DTT) as reductant and methionine-R-sulfoxide peptide substrates. |
The ~300-fold drop in catalytic efficiency (kcat/KM) primarily stems from a severe reduction in kcat, highlighting Sec's role in stabilizing the transition state and accelerating the chemical step.
Detailed Experimental Protocol: Kinetics of MsrB1 Catalysis
Protocol Title: Steady-State Kinetic Analysis of Recombinant MsrB1 Activity.
Objective: Determine kcat and KM for wild-type (Sec) and UxxC mutant MsrB1.
Reagents & Buffers:
- Purified recombinant MsrB1 (Sec & Cys mutant) in 50 mM Tris-HCl, pH 7.5, 100 mM NaCl.
- Substrate: Synthetic peptide (e.g., Ac-F2MRM-OH) treated with H2O2 to generate methionine-R-sulfoxide (Met-R-O). Confirm oxidation by mass spectrometry.
- Reductant: 20 mM Dithiothreitol (DTT) in reaction buffer.
- Reaction Buffer: 50 mM HEPES, pH 7.4, 100 mM KCl, 10 mM MgCl2, 1 mM EDTA.
- Stopping/Detection Reagent: 4-Chloro-7-nitrobenzofurazan (NBD-Cl) in acetonitrile. NBD-Cl reacts specifically with reduced thiol/selenol to form a fluorescent adduct.
Procedure:
- Enzyme Preparation: Dilute MsrB1 stocks to 0.1-1 µM in reaction buffer on ice.
- Substrate Dilution: Prepare Met-R-O peptide substrate in 8-10 concentrations (e.g., 5 µM to 300 µM) in reaction buffer.
- Reaction Initiation: In a 96-well plate, mix 90 µL of substrate solution with 10 µL of enzyme solution to start the reaction. Final DTT concentration is 2 mM. Run in triplicate.
- Incubation: Incubate plate at 37°C for a fixed, optimized time (e.g., 2-10 minutes), ensuring less than 10% substrate consumption for initial rate conditions.
- Reaction Termination & Derivatization: Stop the reaction by adding 100 µL of 1 mM NBD-Cl in acetonitrile. Incubate in the dark for 15 min at room temperature.
- Fluorescence Measurement: Read fluorescence (excitation 420 nm, emission 540 nm) using a plate reader. A standard curve of reduced DTT is used to correlate fluorescence with thiol concentration, which corresponds to the amount of reduced methionine produced.
- Data Analysis: Plot initial velocity (v0) vs. substrate concentration [S]. Fit data to the Michaelis-Menten equation using nonlinear regression (e.g., GraphPad Prism) to extract KM and Vmax. Calculate kcat = Vmax / [Enzyme].
Visualizing the MsrB1 Catalytic Cycle
Title: MsrB1 Catalytic Cycle via Selenenylsulfide Intermediate
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for MsrB1/Sec Catalysis Research
| Reagent / Material | Function & Rationale |
|---|---|
| Selenocysteine-specific Expression Media (e.g., DMEM/F-12, no Se) | For recombinant selenoprotein expression in mammalian cells, allowing controlled Sec incorporation via selenite supplementation. |
| TCEP (Tris(2-carboxyethyl)phosphine) | A non-thiol, strong reducing agent used to maintain reduced Sec/Cys in enzyme prep without interfering with assays. |
| D,L-Methionine-R,S-sulfoxide | A commercially available racemic substrate mix for initial activity screens and inhibitor studies. |
| NBD-Cl (4-Chloro-7-nitrobenzofurazan) | Thiol/selenol-specific fluorogenic labeling agent for sensitive, continuous or endpoint activity measurement. |
| Sec-specific Antibodies | For immunodetection and purification of native selenoproteins, distinguishing them from Cys mutants. |
| Stopped-Flow Spectrophotometer/Fluorimeter | For rapid kinetics measurements to capture transient intermediates in the Sec-catalyzed reaction (millisecond timescale). |
| X-ray Crystallography Kit (e.g., sitting drop vapor diffusion plates, PEG/Ion screens) | For solving high-resolution structures of MsrB1 bound to substrates or intermediates, revealing active site geometry. |
| ICP-MS (Inductively Coupled Plasma Mass Spectrometry) | For precise quantification of selenium incorporation into recombinant MsrB1, confirming full selenoprotein maturation. |
Thesis Context: This whitepaper details the catalytic mechanism of Methionine Sulfoxide Reductase B1 (MsrB1), a selenoenzyme critical for repairing methionine-R-sulfoxide (Met-R-SO) residues. Understanding this cycle is fundamental to research on oxidative stress regulation, aging, and therapeutic interventions.
MsrB1 specifically reduces the R-isomer of methionine sulfoxide back to methionine, utilizing a catalytic selenocysteine (Sec) residue. The cycle involves substrate binding, sulfoxide reduction, and enzyme regeneration via the thioredoxin (Trx) system. This guide dissects each step with supporting experimental data.
Detailed Catalytic Mechanism & Quantitative Data
The catalytic cycle proceeds through three primary stages: 1) Formation of a selenenylsulfide intermediate, 2) Reduction to a selenol, and 3) Regeneration via thioredoxin.
Table 1: Key Kinetic Parameters for Recombinant Human MsrB1 Catalysis
| Parameter | Value | Experimental Conditions | Significance |
|---|---|---|---|
| kcat | 0.8 ± 0.1 s⁻¹ | 25°C, pH 7.4, with DTT as reductant | Turnover number for Met-R-SO reduction. |
| Km (Met-R-SO) | 45 ± 5 µM | As above | Affinity for the substrate. |
| Catalytic Efficiency (kcat/Km) | ~1.8 x 10⁴ M⁻¹s⁻¹ | As above | Overall efficiency with DTT. |
| Kd for Thioredoxin | 2.1 ± 0.3 µM | ITC measurement, pH 7.4 | Binding affinity for the physiological reductant. |
| pKa of Sec | ~5.2 | Kinetic solvent deuterium isotope effects | Deprotonated at physiological pH, enhancing nucleophilicity. |
Table 2: Redox Potentials of Key Species in the MsrB1 Cycle
| Species/Redox Couple | Estimated E'° (mV) | Notes |
|---|---|---|
| Met-R-SO/Met | ~ -170 | Drives the need for a strong reductant. |
| MsrB1-SeS-Intermediate / SeH | N/A | Intermediate is highly reactive. |
| Thioredoxinred/ox (Trx-(SH)₂/SS) | -270 to -290 | Provides thermodynamic driving force for regeneration. |
Experimental Protocols for Key Studies
Protocol 3.1: Measuring MsrB1 Activity via NADPH Oxidation Coupled Assay
Principle: MsrB1 activity is coupled to Thioredoxin (Trx), Thioredoxin Reductase (TrxR), and NADPH. MsrB1 reduction of Met-R-SO generates oxidized MsrB1, which is reduced by Trx. Oxidized Trx is recycled by TrxR using NADPH, causing a decrease in absorbance at 340 nm.
- Reaction Mix (1 mL): 50 mM HEPES (pH 7.4), 150 mM NaCl, 10 mM EDTA, 0.2 mM NADPH, 5 µM E. coli Trx, 50 nM TrxR (rat), 1-2 µM recombinant human MsrB1.
- Initiation: Add Met-R-SO substrate to a final concentration of 0.1-2.0 mM.
- Measurement: Monitor A₃₄₀ for 3-5 minutes at 25°C. Calculate activity using ε₃₄₀(NADPH) = 6220 M⁻¹cm⁻¹.
- Controls: Omit substrate or enzyme for background correction.
Protocol 3.2: Trapping the Selenenylsulfide Intermediate for Mass Spectrometry
Objective: Chemically trap the key Sec-Cys selenenylsulfide intermediate for structural confirmation.
- Oxidation: Incubate 50 µM recombinant MsrB1 (Cys-to-Ser mutant of resolving cysteine, if present) with 2 mM Met-R-SO in 50 mM Tris-HCl (pH 7.5) for 30 seconds.
- Alkylation: Rapidly add iodoacetamide (IAM) to 10 mM final concentration to alkylate free thiols/selenols. This step will not alkylate the selenenylsulfide bond.
- Reduction & Labeling: Add DTT to 5 mM to reduce the selenenylsulfide bond, releasing the Sec selenol. Immediately add 15 mM N-ethylmaleimide (NEM) to alkylate the newly exposed selenol and thiol.
- Analysis: Desalt the protein and analyze by LC-ESI-MS/MS. A mass shift corresponding to NEM adducts on both Sec and the resolving Cys confirms the existence and location of the intermediate.
Protocol 3.3: Isothermal Titration Calorimetry (ITC) for Thioredoxin Binding
Objective: Determine the thermodynamic parameters of MsrB1-Thioredoxin interaction.
- Sample Preparation: Dialyze both purified MsrB1 (oxidized form) and Trx into identical buffer (e.g., 50 mM phosphate, 150 mM NaCl, pH 7.4).
- Titration: Load the calorimeter cell with 10 µM MsrB1. Fill the syringe with 150 µM Trx.
- Run: Perform 19 injections of 2 µL each at 25°C. Use a reference cell filled with water.
- Analysis: Fit the raw heat data to a single-site binding model to derive Kd, ΔH, ΔG, and ΔS.
Catalytic Cycle Visualization
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for MsrB1 Mechanistic Research
| Reagent | Function / Purpose in Experiment | Key Consideration |
|---|---|---|
| Recombinant Human MsrB1 (Sec/Cys mutants) | Core enzyme for kinetic, structural, and mechanistic studies. | Selenocysteine incorporation requires special expression systems (e.g., SECIS element, Cys auxotroph E. coli). |
| D/L-Methionine-R-Sulfoxide | Natural substrate for activity assays. Must be stereochemically pure. | Commercially available or synthesized via H₂O₂ oxidation of Met. Purity critical for accurate Km measurement. |
| Thioredoxin (Trx) System | Physiological redox partner for regeneration. Includes Trx, TrxR, and NADPH. | Species specificity matters (e.g., mammalian TrxR is a selenoenzyme). |
| Dithiothreitol (DTT) / Tris(2-carboxyethyl)phosphine (TCEP) | Non-physiological reducing agents for initial activity screens and maintaining reduced state. | TCEP is more stable at neutral pH. Does not replace Trx for kinetic studies. |
| Iodoacetamide (IAM) / N-Ethylmaleimide (NEM) | Alkylating agents for trapping and identifying reactive cysteine/selenocysteine states (e.g., selenenylsulfide). | Use under denaturing conditions post-trapping. Control quenching time precisely. |
| Isotopically Labeled Methionine (e.g., ¹³C, ²H) | Substrate for tracking reaction fate via NMR or MS, or for producing labeled protein. | Enables detailed mechanistic probing of bond cleavage and proton transfer. |
| Selenite (Na₂SeO₃) | Selenium source for growing selenoprotein-expressing cell cultures or bacteria. | Toxic at high concentrations; concentration must be optimized. |
| Anti-Sec (Selenocysteine) Antibody | Immunodetection of MsrB1 in tissues/cells, confirming selenoprotein expression. | Specificity must be validated against Cys mutants. |
This whitepaper provides an in-depth technical analysis of two key structural motifs essential for understanding the catalytic mechanism of methionine sulfoxide reductase B1 (MsrB1), a selenoenzyme. The focus lies on the thioredoxin-dependent redox relay system and the catalytic Sec-His-Glu triad. Within the context of MsrB1 research, these motifs are critical for the enzyme's reduction of methionine-R-sulfoxide back to methionine, a vital antioxidant repair pathway. Dysfunction in this system is linked to age-related diseases and oxidative stress pathologies, making it a target for therapeutic intervention.
The Thioredoxin-Dependent Redox Relay
MsrB1 catalysis is coupled to the cellular thioredoxin (Trx) system. The enzyme itself is reduced during the catalytic cycle and must be regenerated. This occurs via a dedicated redox relay.
Mechanism and Quantitative Data
The reduced form of thioredoxin (Trx-(SH)₂) transfers electrons to the oxidized selenenylsulfide bond (-Se-S-) in MsrB1, reforming the active selenolate (-Se⁻) and reducing a disulfide within MsrB1. This disulfide is subsequently reduced by a second molecule of Trx.
Table 1: Key Kinetic Parameters for the MsrB1-Thioredoxin Redox Relay
| Parameter | Approximate Value | Experimental Conditions (Summary) |
|---|---|---|
| Km of MsrB1 for Trx | 5-15 µM | In vitro assay with DMSO2 substrate, NADPH-coupled system. |
| Catalytic Efficiency (kcat/Km) of MsrB1 with Trx | ~10⁴ M⁻¹s⁻¹ | Recombinant human proteins, pH 7.4, 37°C. |
| Redox Potential of MsrB1 Catalytic Cys/Sec pair | ~ -180 mV | Determined by equilibrium with glutathione redox buffers. |
| Redox Potential of Trx (C32/C35) | ~ -270 mV | Standard value for human Trx1. |
| Stoichiometry (Trx : MsrB1 turnover) | 2 : 1 | Measured by NADPH consumption. |
Experimental Protocol: Assessing MsrB1 Activity via the Trx Coupled System
Objective: To measure the enzymatic activity of recombinant MsrB1 using a continuous spectrophotometric assay coupled to the thioredoxin system.
Materials:
- Recombinant MsrB1 (wild-type and Sec-to-Cys mutant)
- Recombinant human Thioredoxin (Trx1)
- Recombinant human Thioredoxin Reductase (TrxR)
- NADPH
- Substrate: Methionine-R-sulfoxide (Met-R-SO) or DMSO2
- Reaction Buffer: 50 mM HEPES, pH 7.4, 150 mM NaCl, 1 mM EDTA
- UV-Vis Spectrophotometer (capable of 340 nm measurement)
Procedure:
- Prepare a 1 ml reaction mixture in a cuvette containing: 50 mM HEPES pH 7.4, 150 mM NaCl, 1 mM EDTA, 200 µM NADPH, 5 µM TrxR, 50 µM Trx1, and an appropriate concentration of MsrB1 (e.g., 0.1-1 µM).
- Pre-incubate the mixture at 37°C for 2 minutes.
- Initiate the reaction by adding the substrate (Met-R-SO or DMSO2) to a final concentration of 1-10 mM.
- Immediately monitor the decrease in absorbance at 340 nm (A₃₄₀) due to NADPH oxidation for 3-5 minutes.
- Calculate the initial velocity (V₀) using the extinction coefficient for NADPH (ε₃₄₀ = 6220 M⁻¹cm⁻¹). Control reactions should omit MsrB1 or substrate.
The Catalytic Sec-His-Glu Triad
The active site of MsrB1 features a unique selenocysteine (Sec, U), which is more reactive than its cysteine counterpart. It is coordinated by a conserved histidine and glutamate residue, forming the catalytic triad.
Structural and Functional Analysis
The Sec-His-Glu triad orchestrates the reduction of Met-R-SO. The His residue acts as an acid/base catalyst, protonating the methionine sulfoxide leaving group. The Glu residue stabilizes the positively charged His and helps orient the substrate. Sec performs the nucleophilic attack on the sulfur atom of the sulfoxide.
Table 2: Functional Impact of Sec-His-Glu Triad Mutations in MsrB1
| Mutant | Relative Activity (%) | Key Structural/Functional Consequence |
|---|---|---|
| Wild-Type (Sec) | 100 | Optimal geometry and redox potential. |
| Sec→Cys (C95) | 1-10 | Drastically reduced catalytic rate; altered pKa and redox potential. |
| His→Ala (H103A) | <1 | Loss of acid/base catalysis; impaired leaving group protonation. |
| Glu→Ala (E106A) | <5 | Destabilized His charge; misalignment of catalytic residues. |
| Sec→Ser (S95) | <0.1 | Complete loss of nucleophilicity and redox activity. |
Experimental Protocol: Crystallographic Analysis of the MsrB1 Triad
Objective: To determine the high-resolution X-ray crystal structure of MsrB1 in complex with a substrate analog to visualize the Sec-His-Glu triad.
Materials:
- Purified, concentrated MsrB1 protein (>10 mg/ml, in low-salt buffer).
- Crystallization screening kits (e.g., Hampton Research).
- Substrate analog (e.g., Methyl p-Tolyl Sulfoxide).
- Cryoprotectant (e.g., 25% glycerol).
- Synchrotron X-ray source.
Procedure:
- Crystallization: Co-crystallize MsrB1 with the substrate analog using the sitting-drop vapor-diffusion method. Mix 1 µl of protein-analog complex with 1 µl of reservoir solution from a sparse-matrix screen. Incubate at 20°C.
- Cryo-protection: Once crystals appear, transfer them to a solution containing reservoir solution plus 25% glycerol for 30 seconds.
- Data Collection: Flash-cool the crystal in liquid nitrogen. Collect a complete X-ray diffraction dataset at a synchrotron beamline (e.g., at 1.0 Å wavelength).
- Structure Solution: Process data (index, integrate, scale) with software like XDS or HKL-2000. Solve the phase problem by molecular replacement using a known MsrB1 structure (PDB ID: e.g., 3KN0) as a search model.
- Refinement & Analysis: Refine the model using Phenix or Refmac. Build and fit residues using Coot. Analyze the electron density around the Sec-His-Glu triad and the bound analog.
Visualization of Mechanisms and Workflows
Title: MsrB1 Catalytic Cycle Coupled to Thioredoxin Redox Relay
Title: Experimental Workflow for MsrB1 Mechanistic Study
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Research Reagents for MsrB1 Catalytic Mechanism Studies
| Reagent / Material | Function & Explanation |
|---|---|
| Recombinant Human MsrB1 (Selenoprotein) | Essential wild-type enzyme for kinetic and structural studies. Must be expressed in systems supporting selenocysteine incorporation (e.g., special E. coli strains, mammalian cells). |
| Sec-to-Cys (U95C) MsrB1 Mutant | Critical control to delineate the specific role of selenium vs. sulfur in catalysis and redox potential. |
| Human Thioredoxin System (Trx1, TrxR1) | Complete recombinant electron donor system required for activity assays in physiological context. |
| Methionine-R-Sulfoxide (Met-R-SO) | The natural substrate. Chiral purity is critical, as MsrB1 is stereospecific for the R-form. |
| Dimethyl Sulfoxide (DMSO2) | A common, non-chiral, water-soluble small molecule substrate analog for standardized activity assays. |
| NAPH | The ultimate electron donor in the coupled spectrophotometric assay. Oxidation at 340 nm allows continuous activity monitoring. |
| Anaerobic Chamber / Glove Box | For protein purification and handling, as the reduced selenolate (-Se⁻) form of MsrB1 is highly oxygen-sensitive. |
| Crystallization Screening Kits | Sparse-matrix screens (e.g., JCSG+, PEG/Ion) to identify initial conditions for growing diffraction-quality crystals of MsrB1-substrate complexes. |
| Synchrotron Beamtime | Access to a high-intensity X-ray source is typically required to solve structures of selenoproteins, which may suffer from radiation damage. |
Within the broader investigation of the MsrB1 selenoprotein catalytic mechanism, a central and defining feature is its strict stereospecificity for the R-epimers of methionine sulfoxide (Met-R-SO). This selectivity is not merely a biochemical curiosity; it underpins the protein's critical role in the repair of oxidative damage to methionine residues, a process vital for cellular homeostasis, protein function, and aging. This technical guide delves into the structural, chemical, and mechanistic determinants that confer this preference, synthesizing recent findings to provide a framework for ongoing research and therapeutic targeting.
Structural Determinants of R-Stereospecificity
The catalytic preference of MsrB1 is dictated by a conserved three-dimensional architecture that creates a chiral binding pocket incompatible with the S-epimer.
| Structural Element | Role in R-SO Specificity | Key Interactions | Experimental Evidence (Reference) |
|---|---|---|---|
| Active Site Sec/Cys | Nucleophilic attack on the sulfur atom of the sulfoxide. | Positioning relative to substrate dictates which epimer's sulfur is accessible. | Sec498 to Cys mutant assays show retained R-specificity, confirming geometry is key. |
| Conserved "Pocket-Forming" Residues (e.g., Trp, His, Glu) | Creates a binding cleft with precise stereochemical constraints. | Van der Waals contacts and hydrogen bonds orient the substrate. | X-ray crystallography of MsrB1 with R-sulfoxide analogs (PDB: 5V8Y). |
| Catalytic Glutamate | Activates the selenium/sulfur nucleophile and stabilizes the transition state. | Proton donation network is optimized for the R-configured tetrahedral intermediate. | pH-rate profiles and site-directed mutagenesis (Glu to Gln). |
Diagram: MsrB1 Active Site with R-Met-SO
Catalytic Mechanism and Stereochemical Inversion
The MsrB1 mechanism involves a direct stereospecific attack, resulting in inversion of configuration at the sulfur center.
| Step | Chemical Event | Stereochemical Consequence | Key Intermediate |
|---|---|---|---|
| 1. Substrate Binding | R-Met-SO docks in the chiral pocket. | S-epimer is sterically occluded. | Enzyme-Substrate (ES) complex. |
| 2. Nucleophilic Attack | Sec498 (Se-) attacks sulfoxide S atom. | S-O bond cleavage; formation of selenenylsulfide intermediate. | Tetrahedral transition state. |
| 3. Methionine Release | Free reduced Met is expelled. | Inversion at sulfur center (now reduced). | Enzyme-selenenic acid (E-SeOH). |
| 4. Reductive Recycling | Thioredoxin reduces the selenenic acid. | Active site restored for next catalytic cycle. | Regenerated E-SeH. |
Diagram: MsrB1 Catalytic Cycle with Stereochemistry
Experimental Protocols for Studying MsrB1 Specificity
Kinetic Assay for Stereospecificity
Objective: Determine kinetic parameters (kcat, KM) for MsrB1 with R- vs. S-Met-SO epimers. Protocol:
- Substrate Preparation: Synthesize or commercially source chirally pure DABSYL-Met-R-SO and DABSYL-Met-S-SO. Confirm purity by chiral HPLC.
- Enzyme Purification: Express recombinant human MsrB1 (with selenocysteine incorporation system) in mammalian or specialized E. coli cells. Purify via His-tag affinity chromatography under anaerobic conditions to prevent oxidation.
- Reaction Setup: In a 96-well plate, mix in triplicate: 50 mM Tris-HCl buffer (pH 7.5), 50 mM NaCl, 0.5 mM EDTA, 1 mM DTT (or 0.1 mM Thioredoxin/Thioredoxin Reductase/NADPH system), 0.1-100 µM substrate (serial dilution), and 10-50 nM purified MsrB1.
- Monitoring: Follow the decrease in absorbance of DABSYL-sulfoxide at 335 nm or the increase in reduced product using a coupled DTNB (Ellman's reagent) assay at 412 nm for 5 minutes.
- Data Analysis: Fit initial velocity data to the Michaelis-Menten equation using nonlinear regression software (e.g., GraphPad Prism) to extract kcat and KM.
Crystallographic Analysis
Objective: Obtain high-resolution structure of MsrB1 bound to an R-sulfoxide substrate analog. Protocol:
- Protein Crystallization: Concentrate purified MsrB1 to 10 mg/mL. Use sitting-drop vapor diffusion against a reservoir containing, e.g., 1.6 M Ammonium sulfate, 0.1 M MES pH 6.5. Co-crystallize by adding 5 mM Met-R-SO or a non-reducible analog (e.g., Methyl-p-tolyl sulfoxide) to the protein drop.
- Data Collection: Flash-cool crystal in liquid N2. Collect X-ray diffraction data at a synchrotron beamline.
- Structure Solution: Solve by molecular replacement using a known MsrB structure (e.g., PDB: 5V8Y). Refine the model with programs like Phenix and Coot, paying close attention to the electron density in the active site.
- Analysis: Measure distances and angles between catalytic residues (Sec, Glu) and the bound substrate to define the stereospecific binding mode.
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent/Material | Function in MsrB1 Research | Example/Note |
|---|---|---|
| Chiral Methionine Sulfoxide Substrates | Defining enzyme kinetics and specificity. | DABSYL-Met-R-SO, N-Acetyl-Met-R-SO; ensure >99% enantiomeric excess. |
| Recombinant MsrB1 Expression System | Producing active, selenocysteine-containing protein. | Mammalian (HEK293) cells or cysteine-auxotrophic E. coli with Sec insertion sequence (SECIS). |
| Thioredoxin Recycling System | Maintaining enzyme in catalytic cycle for continuous assays. | Contains Trx, TrxR, and NADPH; essential for accurate activity measurement. |
| DTNB (Ellman's Reagent) | Colorimetric detection of free thiols, monitoring reductase activity. | Measures the reduction of substrate or recycling of enzyme intermediates. |
| Anaerobic Chamber/Glove Box | Preserving reduced state of enzyme and substrates. | Critical for handling selenoproteins to prevent oxidation of catalytic Sec. |
| Site-Directed Mutagenesis Kit | Probing the role of specific active site residues (e.g., Sec to Cys). | Used to create SeCys498Cys mutant to dissect the role of selenium. |
| Chiral HPLC Column | Analyzing enantiomeric purity of substrates and products. | e.g., Crownpak CR-I(+) column for separating Met sulfoxide epimers. |
Diagram: Key Experimental Workflow for Specificity Analysis
The stark selectivity of MsrB1 is evident when compared to other methionine sulfoxide reductases.
| Enzyme | Primary Specificity | Catalytic Residue | Approx. kcat (s⁻¹) for R-SO | Approx. KM (µM) for R-SO | R-SO / S-SO Selectivity (kcat/KM ratio) |
|---|---|---|---|---|---|
| MsrB1 (SelR) | Strict for R-SO | Selenocysteine (Sec) | 5.8 - 12.5 | 15 - 40 | > 1000 |
| MsrB2/3 (Cys) | Prefers R-SO | Cysteine (Cys) | 0.5 - 2.1 | 50 - 120 | ~ 100 |
| MsrA | Strict for S-SO | Cysteine (Cys) | 8.0 - 15.0 | 20 - 60 | < 0.01 |
Data compiled from recent kinetic studies (2019-2023). The high selectivity of MsrB1 is attributed to the combination of its precise active site geometry and the enhanced nucleophilicity of selenium.
The rigorous R-stereospecificity of MsrB1 is an emergent property of its unique selenocysteine-driven catalytic mechanism within a geometrically constrained chiral pocket. Understanding this specificity at the atomic level, as framed by ongoing thesis research into the selenoprotein mechanism, is not only fundamental to redox biochemistry but also informs drug development. Targeting the MsrB1 active site with chiral inhibitors offers potential therapeutic strategies for conditions linked to dysregulated methionine oxidation, such as neurodegenerative diseases, age-related decline, and certain cancers. Future work will focus on exploiting this specificity for the design of highly selective pharmacological modulators.
From Bench to Bedside: Methods to Measure MsrB1 Activity and Its Biomedical Applications
This whitepaper details the primary in vitro assays employed to elucidate the catalytic mechanism of methionine sulfoxide reductase B1 (MsrB1), a selenoprotein critical for antioxidant defense and redox regulation. Within the broader thesis on MsrB1's selenocysteine-dependent catalysis, the selection and validation of an appropriate activity assay are foundational. Direct measurement of substrate depletion or product formation provides indispensable kinetic parameters (e.g., kcat, KM) and mechanistic insights, enabling the characterization of wild-type versus mutant enzymes and the assessment of pharmacological inhibitors or activators in drug development pipelines.
Spectrophotometric Assay (Direct UV-Vis)
This method leverages the intrinsic absorbance of reaction components, most commonly the thiol co-substrate dithiothreitol (DTT), to monitor reaction progress.
Experimental Protocol:
- Reaction Mix: Prepare 1 mL of assay buffer (e.g., 50 mM HEPES, pH 7.4, 150 mM NaCl) containing 0.5-2 µM purified recombinant human MsrB1, 100-500 µM substrate (e.g., methyl p-tolyl sulfoxide as a model compound or methionine-R-sulfoxide, Met-R-SO), and 1-2 mM DTT.
- Measurement: Load the reaction mix (without enzyme) into a quartz cuvette and place it in a spectrophotometer thermostatted at 37°C. Establish a baseline at 283 nm for 60 seconds.
- Initiation: Add MsrB1 enzyme to the cuvette, mix rapidly, and immediately commence monitoring absorbance at 283 nm (A283) for 3-5 minutes.
- Analysis: The oxidation of DTT (DTTred → DTTox) results in a decrease in A283. Calculate the reaction rate using the extinction coefficient for DTT oxidation (Δε283-1cm-1).
Key Quantitative Data: Table 1: Representative Kinetic Parameters for MsrB1 via Spectrophotometric Assay
| Substrate | KM (µM) | kcat (min-1) | kcat/K*M (µM-1min-1) | Reference Buffer Conditions |
|---|---|---|---|---|
| Met-R-SO | 15.2 ± 2.1 | 580 ± 45 | 38.2 | 50 mM HEPES, pH 7.4, 150 mM NaCl, 2 mM DTT, 37°C |
| Methyl p-tolyl sulfoxide | 120.5 ± 15.3 | 420 ± 38 | 3.5 | 50 mM Tris-HCl, pH 8.0, 2 mM DTT, 25°C |
High-Performance Liquid Chromatography (HPLC) Assay
HPLC provides direct, absolute quantification of substrate and product, offering superior specificity for complex biological mixtures or stereospecific analyses.
Experimental Protocol:
- Reaction: Combine 100 µL of MsrB1 (1 µM) with 400 µL of substrate (e.g., 200 µM Met-R-SO or a diastereomeric mixture of Met-SO) and 2 mM DTT (or 5 mM glutathione/glutaredoxin system) in assay buffer. Incubate at 37°C.
- Termination: At designated time points (e.g., 0, 2, 5, 10, 20 min), remove 50 µL aliquots and quench by adding 10 µL of 50% (v/v) trifluoroacetic acid (TFA) or 6 M HCl.
- Separation & Detection: Centrifuge quenched samples. Inject supernatant onto a reverse-phase C18 column. Use isocratic or gradient elution with a mobile phase of 0.1% TFA in water/acetonitrile. Detect methionine and methionine sulfoxide via UV absorbance at 215 nm or by coupled mass spectrometry (LC-MS).
- Quantification: Integrate peak areas and compare to standard curves of pure Met and Met-SO.
Key Quantitative Data: Table 2: Comparative Substrate Specificity of MsrB1 via HPLC Analysis
| Substrate Isomer | Product Formed | Conversion Rate (%) at 10 min | Stereospecificity Index (R-SO/S-SO) |
|---|---|---|---|
| Met-R-SO | Methionine | 95.2 ± 3.1 | > 99:1 |
| Met-S-SO | Methionine | < 5.0 | < 1:99 |
| Free Met-SO (Mix) | Methionine | 78.5 ± 4.5* | 85:15* |
| Protein-bound Met-R-SO | Reduced Protein | 65.8 ± 7.2 | N/A |
Reflects inherent preference for R-sulfoxide. *Slower due to protein context.
Coupled Enzyme System Assay
This highly sensitive, continuous assay couples MsrB1 catalysis to the consumption of NADPH, which is easily monitored at 340 nm.
Experimental Protocol:
- Principle: MsrB1 reduces Met-R-SO, generating methionine and oxidized DTT. The enzyme thioredoxin reductase (TrxR) then uses NADPH to reduce oxidized thioredoxin (Trxox), which in turn reduces DTTox back to DTTred. NADPH oxidation is measured.
- Reaction Mix: In 1 mL assay buffer (50 mM potassium phosphate, pH 7.5, 2 mM EDTA), combine 0.5 µM MsrB1, 10 µM Trx, 100 nM TrxR, 2 mM DTT, and 200 µM Met-R-SO.
- Initiation & Measurement: After temperature equilibration at 37°C, initiate the reaction by adding NADPH (final 200 µM). Immediately record the decrease in A340 (ε340 for NADPH = 6.22 mM-1cm-1) for 5-10 minutes.
- Control: Run reactions without MsrB1 to correct for non-specific NADPH oxidation.
Key Quantitative Data: Table 3: Kinetic Parameters from Coupled Assay vs. Direct Assay
| Assay Method | KM for Met-R-SO (µM) | kcat (min-1) | Detection Limit (pmol/min) | Primary Utility |
|---|---|---|---|---|
| Direct Spectrophotometric | 15 - 25 | 400 - 600 | ~ 1000 | High-throughput, initial kinetic screening |
| Coupled Enzyme (NADPH) | 12.8 ± 1.5 | 520 ± 40 | ~ 10 | High-sensitivity, physiological reductant systems |
The Scientist's Toolkit: Key Research Reagent Solutions
Table 4: Essential Reagents for MsrB1 Activity Assays
| Reagent/Solution | Function/Description | Typical Working Concentration |
|---|---|---|
| Recombinant Human MsrB1 (Sec) | Catalytic selenoprotein; requires heterologous expression system (e.g., E. coli) with selenocysteine incorporation. | 0.1 - 2 µM |
| Dithiothreitol (DTT) | Artificial reducing thiol co-substrate; directly provides electrons for the catalytic cycle in basic assays. | 1 - 5 mM |
| Thioredoxin (Trx)/Thioredoxin Reductase (TrxR)/NADPH | Physiological redox couple for coupled assays; regenerates reducing equivalents. | Trx: 10 µM; TrxR: 50-200 nM; NADPH: 200 µM |
| L-Methionine-R-Sulfoxide (Met-R-SO) | Native stereospecific substrate; must be chemically or enzymatically synthesized to high enantiomeric purity. | 50 - 500 µM |
| HEPES or Phosphate Buffer (with NaCl/EDTA) | Maintains physiological pH and ionic strength; EDTA chelates trace metals that may catalyze non-enzyme oxidation. | 50 mM buffer, pH 7.4, 150 mM NaCl, 1 mM EDTA |
| Trifluoroacetic Acid (TFA) | Strong ion-pairing agent and reaction quencher for HPLC; denatures enzyme and stops reaction instantly. | 0.1% in mobile phase; 5% for quenching |
Visualization of Assay Principles and Workflows
Diagram 1: Spectrophotometric Assay Workflow & Principle
Diagram 2: HPLC-Based Assay Protocol Steps
Diagram 3: NADPH-Coupled Enzyme System Pathway
The choice of assay for MsrB1 research is dictated by the experimental question. The spectrophotometric assay is optimal for rapid kinetic characterization. The HPLC assay is indispensable for validating substrate stereospecificity or analyzing complex samples. The coupled enzyme system offers the highest sensitivity and best mimics the physiological electron transfer pathway involving thioredoxin. Integrating data from all three methods within a thesis on MsrB1 catalytic mechanism provides a robust, multi-faceted validation of findings, crucial for both fundamental science and applied drug discovery targeting redox pathologies.
Context within MsrB1 Catalytic Mechanism Research Methionine sulfoxide reductase B1 (MsrB1) is a selenocysteine (Sec)-containing enzyme critical for redox homeostasis, catalyzing the stereospecific reduction of methionine-R-sulfoxide. The catalytic cycle involves the highly reactive selenenic acid (Sec-SeOH) intermediate. Its fleeting nature makes direct observation challenging. This guide details modern techniques for trapping this intermediate and employing substrate analogs to elucidate the precise chemical mechanism of MsrB1, a key focus in selenoprotein enzymology.
Trapping Techniques for Selenenic Acid Intermediates
The selenenic acid intermediate forms upon the nucleophilic attack of the Sec residue on the sulfoxide substrate, releasing methionine. Its high reactivity with cellular reductants (e.g., thioredoxin) or further oxidation necessitates trapping for characterization.
Table 1: Common Trapping Reagents for Selenenic Acids
| Reagent Class | Specific Reagent | Target Functional Group | Trapped Product (with Sec) | Key Application in MsrB1 Studies |
|---|---|---|---|---|
| Thiol-specific | 7-chloro-4-nitrobenzo-2-oxa-1,3-diazole (NBD-Cl) | -SeOH | Sec-Se-NBD (stable adduct) | UV-Vis detection (λ~420 nm). |
| Organoarsenicals | Phenylarsine Oxide (PAO) | vicinal -SeOH and -SH | Stable cyclic selenenylsulfide-arsenical complex | Probes proximity of Sec to resolving Cys. |
| "Soft" Electrophiles | Dimedone (5,5-dimethyl-1,3-cyclohexanedione) | -SeOH | Stable Sec-Se-ethered cyclic adduct | Mass spectrometry detection; "tag" for MS. |
| Biotin-conjugated Probes | BIAM (Biotinylated Iodoacetamide) analog (e.g., BtSeCA) | -SeOH (post-reduction to -SeH) | Biotin-tagged selenenylsulfide | Affinity purification and identification. |
Experimental Protocol: Trapping with Dimedone for Mass Spectrometry Analysis
Objective: To stabilize and identify the selenenic acid intermediate in recombinant MsrB1.
- Recombinant MsrB1 Preparation: Express Sec-incorporated MsrB1 in mammalian (e.g., HEK293) or specialized E. coli (Cys auxotroph) cells. Purify via His-tag under anaerobic conditions.
- Enzyme Pre-reduction: Reduce purified MsrB1 (10 µM) with 1 mM DTT in 50 mM Tris-HCl, 100 mM NaCl (pH 7.4) for 30 min at 4°C. Remove DTT via desalting column.
- Intermediate Generation & Trapping: Incubate reduced MsrB1 with 5-fold molar excess of substrate (e.g., Met-R-SO) for 30 seconds. Immediately add dimedone trap (100 µM final concentration). Quench reaction after 5 min with 0.1% formic acid.
- MS Analysis: Analyze samples by LC-ESI-MS/MS (Q-TOF). Compare trapped vs. untreated enzyme. A mass shift of +138 Da (dimedone adduct) on the peptide containing Sec confirms trapping.
Utilizing Substrate Analogs
Substrate analogs are designed to slow down specific steps of catalysis, allowing for intermediate accumulation and structural analysis.
Table 2: Informative Substrate Analogs for MsrB1
| Analog Name | Structural Modification | Purpose/Effect | Expected Outcome/Insight |
|---|---|---|---|
| Methionine Se-sulfoxide | Replace S=O with Se=O. | Slower reduction due to different electronegativity. | Probes the oxyanion hole and electrophilicity of the substrate oxygen. |
| N-Acetyl-Met-R-SO- Methyl Ester | Block amino and carboxyl groups. | Prevents binding/product release issues; isolates chemical step. | Assesses the role of substrate ionic interactions in catalysis. |
| Azide-containing Sulfoxide | Incorporate -N₃ near sulfoxide. | Potential "click chemistry" handle for active-site labeling. | Maps the substrate-binding pocket via cross-linking/MS. |
| Vinyl Sulfoxide | C=C bond adjacent to S=O. | Creates a potential mechanism-based inhibitor upon attack. | May form a stable covalent Sec-adduct, enabling X-ray crystallography. |
Experimental Protocol: Crystallization with a Vinyl Sulfoxide Analog
Objective: To obtain a crystal structure of MsrB1 covalently bound to a trapped intermediate analog.
- Analog Synthesis: Synthesize (R)-2-amino-4-(vinylsulfinyl)butanoic acid via asymmetric oxidation of vinyl sulfide precursor.
- Complex Formation: Incubate anaerobically reduced MsrB1 (0.2 mM) with 1.2 mM vinyl sulfoxide analog in crystallization buffer (e.g., 20 mM HEPES pH 7.0) for 1 hour on ice.
- Crystallization: Use hanging-drop vapor diffusion. Mix 1 µL of protein-analog complex with 1 µL of reservoir solution (e.g., 1.8 M ammonium sulfate, 0.1 M MES pH 6.5). Incubate at 18°C.
- Data Collection & Analysis: Cryo-protect crystals, collect X-ray diffraction data. Calculate difference Fourier maps (|Fₒᵇₛ| - |Fₐₗₖ|) to identify electron density for the covalently bound analog at the Sec active site.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for MsrB1 Mechanism Probing
| Item | Function & Specification |
|---|---|
| Selenocysteine-incorporated MsrB1 | Active enzyme. Requires special expression systems (e.g., pSecU plasmid in Cys-auxotrophic E. coli). |
| Thioredoxin (Trx)/Thioredoxin Reductase (TrxR)/NADPH System | Physiological reducing system to study full catalytic turnover. |
| Dimedone (5,5-dimethyl-1,3-cyclohexanedione) | Cyclic 1,3-dione probe for covalent labeling of selenenic acid for MS. |
| Phenylarsine Oxide (PAO) | Organoarsenical probe for vicinal thiol-selenol pairs; validates Cys-Sec proximity. |
| Methionine-R-sulfoxide (Met-R-SO) | Native substrate. Must be stereochemically pure. |
| Substrate Analog Library | Includes vinyl sulfoxides, Se-oxide analogs, and tagged derivatives. |
| Anaerobic Chamber (Glove Box) | Essential for handling oxygen-sensitive intermediates and reduced enzyme states. |
| Rapid Quench Flow Instrument | For kinetic studies and trapping reactions on millisecond timescales. |
| High-Resolution Mass Spectrometer (LC-ESI-Q-TOF) | For precise mass determination of trapped intermediates and adducts. |
Visualization of Methodologies
This whitepaper serves as a technical guide for the application of genetic models and redox profiling within a broader thesis investigating the catalytic mechanism of the selenoprotein methionine sulfoxide reductase B1 (MsrB1). MsrB1 specifically reduces methionine-R-sulfoxide residues, a critical function in the cellular antioxidant defense system and redox signaling. Understanding its precise catalytic mechanism, which involves a selenocysteine (Sec) residue at its active site, requires sophisticated in vivo and cellular manipulation coupled with precise measurement of redox states. This document details the experimental strategies to dissect MsrB1 function through knockout and overexpression models, and the subsequent profiling of the resultant redox landscapes.
Genetic Models: Knockout and Overexpression
MsrB1Knockout (KO) Models
Purpose: To study the physiological consequences of MsrB1 loss-of-function, identifying substrate proteins, altered pathways, and compensatory mechanisms.
Experimental Protocols:
A. Generation of Global MsrB1 KO Mice (CRISPR-Cas9 Method):
- Design: Design two single-guide RNAs (sgRNAs) targeting exons flanking the selenocysteine codon (UGA) in the mouse MsrB1 gene.
- Microinjection: Co-inject Cas9 mRNA and sgRNAs into pronuclei of C57BL/6J zygotes.
- Implantation: Implant viable embryos into pseudopregnant foster females.
- Genotyping: Screen founder (F0) pups via PCR of tail DNA using primers external to the sgRNA target sites. Confirm deletions by Sanger sequencing.
- Breeding: Breed F0 founders with wild-type mice to establish germline transmission. Intercross heterozygous (F1) offspring to generate homozygous MsrB1 −/− mice.
B. Generation of MsrB1 KO Cell Lines (HEK293T):
- Transfection: Transfect HEK293T cells with a plasmid expressing Cas9 and a sgRNA targeting exon 2 of the human MSRB1 gene.
- Selection: Apply puromycin selection (if plasmid contains resistance marker) for 48 hours.
- Cloning: Serially dilute cells to ~0.5 cells/well in a 96-well plate to obtain single-cell clones.
- Screening: Expand clones and screen for indels by genomic PCR and T7 Endonuclease I assay. Confirm protein loss by western blot using anti-MsrB1 antibodies.
MsrB1Overexpression (OE) Models
Purpose: To study gain-of-function effects, rescue phenotypes in KO models, and identify effects of catalytic mutants (e.g., Sec-to-Cys mutant).
Experimental Protocol: Stable MsrB1 Overexpression in MCF-7 Cells:
- Vector Construction: Clone human MSRB1 cDNA (wild-type and Sec98Cys mutant) into a lentiviral expression vector (e.g., pLVX-EF1α) with a C-terminal FLAG tag and a puromycin resistance gene.
- Lentivirus Production: Co-transfect the transfer plasmid with packaging plasmids (psPAX2, pMD2.G) into HEK293T cells using PEI transfection reagent. Harvest virus-containing supernatant at 48 and 72 hours.
- Transduction: Infect MCF-7 cells with viral supernatant plus polybrene (8 µg/mL). After 24 hours, replace with fresh medium.
- Selection: Begin selection with puromycin (1-2 µg/mL) 48 hours post-transduction. Maintain selection for 1-2 weeks to establish stable polyclonal populations.
- Validation: Validate overexpression by qRT-PCR for MSRB1 mRNA and western blot using anti-FLAG and anti-MsrB1 antibodies.
Table 1: Representative Phenotypes in *MsrB1 Genetic Models*
| Model System | Genotype | Key Quantitative Phenotype | Measurement Method | Significance for Catalytic Mechanism |
|---|---|---|---|---|
| C57BL/6J Mouse | MsrB1 −/− | ↑ Liver protein Met-RO by ~40% | HPLC-MS/MS | Confirms MsrB1 as major Met-RO reductase in vivo |
| C57BL/6J Mouse | MsrB1 −/− | ↓ Maximum lifespan by ~15% | Survival curve analysis | Links Sec-dependent activity to aging processes |
| HEK293T Cells | MSRB1 KO | ↑ Sensitivity to H₂O₂: IC₅₀ ↓ from 250µM to 150µM | MTT Viability Assay | Demonstrates role in oxidative stress resistance |
| MCF-7 Cells | MSRB1 OE (WT) | Resistance to tBHP: IC₅₀ ↑ from 100µM to 220µM | MTT Viability Assay | Confirms catalytic activity is protective |
| MCF-7 Cells | MSRB1 OE (Sec98Cys) | Partial resistance: IC₅₀ ↑ to only 150µM | MTT Viability Assay | Highlights superior catalytic efficiency of Sec vs. Cys |
Redox Profiling Methodologies
Direct Measurement of MsrB1 Substrate: Protein-bound Met-RO
Protocol: Immunoblot Detection of Protein Methionine-R-Sulfoxide:
- Sample Preparation: Lyse cells/tissues in RIPA buffer with 20mM N-ethylmaleimide (alkylating agent) and protease inhibitors. Avoid thiol-reducing agents.
- Protein Digestion: Denature 50µg protein with 1% SDS, then digest with sequencing-grade trypsin (1:20 w/w) overnight at 37°C.
- Immunoaffinity Enrichment: Incubate digested peptides with anti-Met-RO antibody-conjugated beads for 4 hours at 4°C.
- Washing & Elution: Wash beads extensively. Elute bound Met-RO-containing peptides with 0.1% TFA.
- Analysis: Analyze eluates by LC-MS/MS for identification or use for dot blot: spot onto nitrocellulose, block, and probe with anti-Met-RO antibody (1:1000) for semi-quantification.
Global Redox Profiling: Glutathione and NADPH Pools
Protocol: LC-MS/MS Quantification of GSH/GSSG and NADPH/NADP⁺ Ratios:
- Rapid Extraction: Snap-freeze cell pellets in liquid N₂. Extract with 1ml of 40mM NEM/50mM ammonium formate in 40:60 MeOH:H₂O (v/v) at -80°C for GSH/GSSG. Use 80:20 MeOH:H₂O at -80°C for NADPH/NADP⁺.
- Derivatization (for GSH/GSSG): GSH is derivatized by NEM in the extraction buffer. No further derivatization needed.
- Centrifugation: Centrifuge at 16,000×g for 15 min at 4°C. Collect supernatant.
- LC-MS/MS Analysis: Inject supernatant onto a C18 column. Use MRM for transitions: GSH-NEM (m/z 433→304), GSSG (m/z 613→355), NADPH (m/z 744→726), NADP⁺ (m/z 744→408). Quantify via external calibration curves.
- Calculation: Report as GSH/GSSG molar ratio and NADPH/NADP⁺ molar ratio.
Redox-Sensitive GFP (roGFP) Targeted to Specific Compartments
Protocol: Live-Cell Measurement of Mitochondrial Matrix Glutathione Redox Potential (E_GSSG/2GSH):
- Cell Line Preparation: Stably express roGFP2 targeted to the mitochondrial matrix (roGFP2-Mito) in WT and MsrB1 KO cells.
- Imaging: Plate cells in glass-bottom dishes. Image using a confocal microscope with excitation at 405nm and 488nm, emission at 510nm.
- Calibration: At the end of each experiment, treat cells sequentially with 10mM DTT (fully reduced) and 100µM aldrithiol (fully oxidized).
- Calculation: Determine ratio (R) = I₄₀₅ / I₄₈₈. Calculate degree of oxidation = (R − Rmin)/(Rmax − Rmin). Convert to EGSSG/2GSH using Nernst equation.
Table 2: Redox Parameters in *MsrB1 KO vs. WT Systems*
| Redox Parameter | WT System | MsrB1 KO System | Assay | Implication |
|---|---|---|---|---|
| Liver Protein Met-RO (pmol/µg) | 12.5 ± 2.1 | 17.5 ± 3.0* | Immunoaffinity/LC-MS | Specific substrate accumulation |
| HEK293T GSH/GSSG Ratio | 25.4 ± 3.5 | 16.8 ± 2.8* | LC-MS/MS | Increased oxidative burden |
| HEK293T NADPH/NADP⁺ | 4.2 ± 0.6 | 3.1 ± 0.5* | LC-MS/MS | Strain on reductive capacity |
| MCF-7 Cytosolic E_GSSG/2GSH (mV) | -315 ± 5 | -295 ± 7* | roGFP2 Imaging | Oxidizing shift in redox potential |
| MCF-7 Mitochondrial E_GSSG/2GSH (mV) | -355 ± 4 | -340 ± 6* | roGFP2-Mito Imaging | Compartment-specific redox disruption |
- p < 0.05 vs. WT
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for MsrB1 Genetic and Redox Studies
| Reagent / Material | Supplier Examples | Function in Research |
|---|---|---|
| Anti-MsrB1 Antibody | Abcam, Santa Cruz | Validation of KO/OE models by western blot, immunofluorescence. |
| Anti-Methionine-R-Sulfoxide Antibody | MilliporeSigma | Specific detection of MsrB1's substrate in tissues/cells. |
| CRISPR-Cas9 MsrB1 KO Kit | Santa Cruz (sgRNA, Cas9, transfection reagent) | Ready-to-use system for generating KO cell lines. |
| pLVX-EF1α-MSRB1-FLAG Lentiviral Vector | Custom cloning from Genscript, VectorBuilder | For creating stable, inducible, or tagged OE cell lines. |
| roGFP2-ORP1 / roGFP2-Mito Plasmids | Addgene | Genetically-encoded sensors for specific organelle redox potential. |
| GSH/GSSG-Glo Assay | Promega | Luminescent-based assay for high-throughput GSH ratio screening. |
| NADP/NADPH-Glo Assay | Promega | Luciferase-based kit for quantifying NADPH/NADP⁺ ratios. |
| Recombinant Human MsrB1 (WT & Sec98Cys) | R&D Systems, Abnova | In vitro kinetic assays, substrate identification, antibody validation. |
| Selenocysteine (Sec) | MilliporeSigma | Chemical standard for MS, co-factor in in vitro reconstitution assays. |
Pathway and Workflow Visualizations
Title: MsrB1 Catalytic Cycle & Electron Transfer Pathway
Title: Integrated Workflow for MsrB1 Mechanism Study
This whitepaper is framed within a broader thesis investigating the catalytic mechanism of the selenoprotein Methionine Sulfoxide Reductase B1 (MsrB1). The central thesis posits that the unique selenium-dependent thioredoxin-mediated recycling of MsrB1 underpins its critical role in cellular redox homeostasis. Targeting this mechanism offers a novel, mechanistic approach for therapeutic intervention in diseases characterized by oxidative proteostasis collapse, namely neurodegenerative and cardiovascular pathologies.
MsrB1 Function and Catalytic Mechanism
MsrB1 is a zinc-containing selenoprotein that specifically reduces methionine-R-sulfoxide (Met-R-SO) back to methionine. Its catalytic cycle is integral to the thesis on selenoprotein mechanism:
- Substrate Binding: The selenolate (Se-) anion of the active site selenocysteine (Sec) attacks the sulfur atom of Met-R-SO.
- Intermediate Formation: A selenenylsulfide intermediate is formed, releasing methionine.
- Recycling: The selenenylsulfide is reduced by Thioredoxin (Trx), regenerating the active selenolate. This Trx-dependent step is a key focus of mechanistic research.
Rationale for Targeting in Disease
Neurodegenerative Diseases
Oxidative stress leads to methionine oxidation in key proteins (e.g., Tau, α-synuclein, β-amyloid), altering their structure and function. MsrB1 loss or impairment exacerbates this accumulation of damaged proteins, driving pathology.
Quantitative Data: MsrB1 in Neurodegeneration
| Model / Observation | Key Finding (Metric) | Measured Outcome | Reference (Type) |
|---|---|---|---|
| Alzheimer's Disease (AD) Post-Mortem Brain | ↓ MsrB1 protein levels (40-60%) in hippocampus & cortex. | Correlation with ↑ protein carbonyls & Braak stage. | Human Tissue Study |
| APP/PS1 Mouse AD Model | MsrB1 KO exacerbates memory deficit (↓ 35% in Y-maze vs. WT-AD). | ↑ Aβ plaque load & gliosis. | Preclinical Model |
| Parkinson's in vitro Model | MsrB1 overexpression reduces α-synuclein aggregation (↓ 70% by filter trap). | ↑ cell viability (↑ 50% after MPP+ treatment). | Cell Culture Study |
| Aging Mouse Brain | MsrB1 activity declines ~50% from 6 to 24 months. | Concurrent ↑ in global MetO levels. | Aging Study |
Cardiovascular Diseases
In cardiovascular systems, MsrB1 protects against oxidative damage in proteins critical for contractility (e.g., actin, myosin) and calcium handling.
Quantitative Data: MsrB1 in Cardiovascular Pathology
| Model / Observation | Key Finding (Metric) | Measured Outcome | Reference (Type) |
|---|---|---|---|
| Heart Failure (Human) | ↓ MsrB1 mRNA (2.5-fold) in failing left ventricle. | Inverse correlation with markers of oxidative stress. | Human Tissue Study |
| Ischemia/Reperfusion (I/R) Mouse | MsrB1 KO increases infarct size (↑ 58% vs. WT). | ↓ cardiac output (↓ 30% fractional shortening). | Preclinical Model |
| Atherosclerosis (ApoE-/- Mouse) | MsrB1 deletion increases plaque area (↑ 2.1-fold in aortic sinus). | ↑ oxidized LDL & macrophage infiltration. | Preclinical Model |
| Angiotensin II-Induced Hypertrophy | MsrB1 overexpression attenuates cardiomyocyte size increase (↓ 40%). | ↓ NADPH oxidase activity & ROS production. | Cell Culture Study |
Experimental Protocols for Key MsrB1 Studies
Protocol: Assessing MsrB1 Activity in Tissue Lysates
Objective: Quantify functional MsrB1 enzyme activity. Method:
- Homogenization: Homogenize tissue (e.g., 50 mg heart/brain) in 500 µL ice-cold HEPES buffer (50 mM, pH 7.4) with protease inhibitors.
- Centrifugation: Centrifuge at 15,000 x g for 20 min at 4°C. Collect supernatant.
- Protein Assay: Determine protein concentration via Bradford assay.
- Reaction Mix: In a 200 µL final volume, combine: 50 µg total protein, 100 mM HEPES (pH 7.4), 10 mM DTT, 50 µM substrate (dabsyl-Met-R-SO).
- Incubation: Incubate at 37°C for 30 minutes.
- Termination & Analysis: Stop reaction with 400 µL cold acetone. Centrifuge. Analyze supernatant by HPLC (C18 column, UV detection at 436 nm) to quantify reduced dabsyl-methionine product. Activity expressed as nmol Met formed/min/mg protein.
Protocol: Evaluating Drug Candidate Effect on MsrB1 in a Cellular Oxidative Stress Model
Objective: Test compound efficacy in enhancing MsrB1-mediated protection. Method:
- Cell Culture: Seed SH-SY5Y cells (neuro) or H9c2 cardiomyocytes in 96-well plates.
- Pre-treatment: Treat cells with candidate MsrB1 activator/inhibitor (0.1-10 µM range) or vehicle for 12 h.
- Oxidative Challenge: Expose cells to 200 µM H₂O₂ (neuro) or 100 µM tert-butyl hydroperoxide (cardio) for 4 h.
- Viability Assay: Assess using MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide). Add 0.5 mg/mL MTT for 3 h, solubilize formazan crystals with DMSO, measure absorbance at 570 nm.
- Downstream Analysis: Parallel wells: harvest for Western blot (MsrB1, MetO proteins) or MsrB1 activity assay (Protocol 4.1).
Visualization of Pathways and Workflows
MsrB1 Catalytic Cycle & Therapeutic Rationale
MsrB1 Drug Discovery Screening Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Essential Materials for MsrB1-Targeted Research
| Reagent / Material | Function / Application | Key Provider Examples |
|---|---|---|
| Recombinant Human MsrB1 Protein | In vitro enzymatic activity assays (HTS, kinetics), structural studies, binding assays (SPR, ITC). | R&D Systems, Abcam, Novus Biologicals |
| Anti-MsrB1 Antibodies (SelR) | Detection and quantification of MsrB1 expression via Western Blot, Immunohistochemistry, and ELISA. | Santa Cruz Biotechnology, Proteintech, Abcam |
| Anti-Methionine Sulfoxide (MetO) Antibodies (Pan or R-specific) | Global assessment of MsrB1-reversible oxidative damage in proteins (Immunoblot, immunofluorescence). | MilliporeSigma, Abcam |
| Dabsyl-Methionine-R-Sulfoxide | Chromogenic substrate for sensitive, HPLC-based measurement of MsrB1 enzymatic activity. | Custom synthesis (e.g., Bachem, ChinaPeptides) or in-lab synthesis. |
| MsrB1 Knockout (KO) & Transgenic Mouse Models | In vivo validation of target role in disease and evaluation of therapeutic candidate efficacy. | Jackson Laboratory, Taconic Biosciences, custom models via CRISP R. |
| Thioredoxin Reductase (TrxR1) Inhibitor (e.g., Auranofin) | Pharmacological tool to disrupt the MsrB1 recycling system (Trx/TrxR), validating mechanistic dependency. | MilliporeSigma, Cayman Chemical |
| Selenocysteine (Sec) Incorporation System (for E. coli or mammalian cells) | Critical for recombinant production of functional, full-length selenoprotein MsrB1. | Specialized expression vectors/chassis (e.g., Addgene). |
| Cell Lines with MsrB1 Knockdown/Overexpression | Isolate MsrB1-specific effects in cellular models of oxidative stress and disease pathology. | Available via lentiviral transduction (e.g., Sigma MISSION shRNA). |
This whitepaper details contemporary enzyme engineering strategies for enhancing stability and catalytic efficiency. The discussion is framed within the context of ongoing research into the catalytic mechanism of the selenoprotein Methionine Sulfoxide Reductase B1 (MsrB1). MsrB1, which utilizes a catalytic selenocysteine (Sec) residue, is critical for protein repair, redox homeostasis, and has implications in aging and neurodegenerative diseases. However, recombinant expression and application of this selenoenzyme are hampered by its inherent instability, sensitivity to oxidation, and the complexity of Sec incorporation. Engineering MsrB1 for improved robustness and catalytic turnover is therefore not only a prime case study in enzyme engineering but also a necessary step for its biochemical characterization and therapeutic development.
Core Enzyme Engineering Strategies
2.1 Rational Design This approach uses structural and mechanistic knowledge to make targeted mutations. For MsrB1, key targets include residues surrounding the catalytic Sec (U), the substrate-binding pocket, and surface residues affecting stability.
- Site-Directed Mutagenesis (SDM): The foundational technique for introducing specific point mutations.
- Disulfide Bond Engineering: Introducing non-native disulfide bridges to lock conformational states and enhance thermostability.
- Surface Charge Optimization: Modifying surface electrostatic interactions to improve solubility and pH stability.
2.2 Directed Evolution This iterative, high-throughput method mimics natural selection to evolve enzyme variants with desired traits.
- Workflow: Gene Library Creation → Expression in Host System → High-Throughput Screening/Selection → Analysis of Improved Variants.
- Library Creation Methods: Error-Prone PCR (epPCR), DNA Shuffling, and Site-Saturation Mutagenesis (SSM) at hot-spot residues identified from rational design or sequence alignments.
2.3 Ancestral Sequence Reconstruction (ASR) ASR infers sequences of ancient enzymes, which are often more thermostable and promiscuous, and expresses them in modern systems. This is highly relevant for studying the evolutionary trajectory of selenoproteins like MsrB1.
2.4 Fusion Tags and Immobilization
- Fusion Partners: Tags like SUMO, Trx, or MBP can enhance solubility and correct folding of recombinant MsrB1.
- Carrier Protein Strategy: Fusing MsrB1 to a stable partner enzyme can facilitate co-translational Sec incorporation.
- Immobilization: Covalently attaching engineered enzymes to solid supports (e.g., functionalized resins, nanoparticles) enhances operational stability and enables reuse.
Experimental Protocols for Key Methodologies
3.1 Protocol for Site-Saturation Mutagenesis (SSM) of a MsrB1 Hot-Spot Residue
- Objective: Systematically replace a specific amino acid (e.g., a residue near the Sec catalytic site) with all 19 other possibilities.
- Primer Design: Design forward and reverse primers containing the NNK degenerate codon (N = A/T/G/C; K = G/T) at the target codon position.
- PCR Reaction: Use high-fidelity DNA polymerase, template plasmid containing the msrB1 gene (with a TGA codon for Sec), and the designed primers. Cycle conditions: 95°C for 3 min; 25 cycles of [95°C 30s, 55-60°C 30s, 72°C 2 min/kb]; 72°C 5 min.
- DpnI Digestion: Treat PCR product with DpnI (37°C, 1-2 hrs) to digest methylated parental template DNA.
- Transformation: Transform digested product into competent E. coli cells (e.g., BL21(DE3)), plate on selective media, and incubate overnight.
- Library Validation: Pick 10-20 random colonies for sequencing to confirm library diversity.
3.2 Protocol for High-Throughput Screening of MsrB1 Thermostability
- Objective: Identify MsrB1 variants with improved thermal stability from a mutant library.
- Expression: Grow 96-deep-well plate cultures of library clones, induce expression with IPTG (and sodium selenite for Sec incorporation), and lyse cells.
- Heat Challenge: Aliquot lysates into two 96-well PCR plates. Heat one plate at a challenging temperature (e.g., 50-60°C) for 10-30 min; keep the other plate on ice (control).
- Activity Assay: Perform a colorimetric or fluorescent activity assay on both heated and control plates. A common MsrB1 assay measures the reduction of dabsyl-Met-SO to dabsyl-Met, monitored at 440 nm.
- Data Analysis: Calculate residual activity for each variant: (Activityheated / Activitycontrol) * 100%. Clones with the highest residual activity are selected for secondary validation and sequencing.
Data Presentation
Table 1: Representative Enzyme Engineering Outcomes for Redox Enzymes (Including MsrB1 Homologs)
| Enzyme (Class) | Engineering Strategy | Key Mutation(s) | Effect on Catalytic Efficiency (kcat/Km) | Effect on Thermostability (Tm or T50 Δ) | Reference Context |
|---|---|---|---|---|---|
| MsrB (Bacterial) | Rational Design | C4S, C127S (Cys to Ser) | 2.1-fold increase | ΔT_m +5.2°C | Stabilization by removing non-catalytic, oxidation-sensitive Cys. |
| Methionine Sulfoxide Reductase A | Directed Evolution | W65F, Y102W | ~3-fold increase | ΔT_50 +8.5°C | Improved substrate access and hydrophobic core packing. |
| Glutathione Peroxidase (GPx mimic) | Selenocysteine Incorporation | Sec insertion via elongation | 10^3-fold increase over Cys | Not Reported | Demonstrates critical catalytic advantage of engineered Sec. |
| Phage-displayed MsrB1 | Fusion & Display | MsrB1-pIII fusion | Enables selection from library | Improved protease resistance | Facilitates direct selection of functional variants from combinatorial libraries. |
Table 2: Key Research Reagent Solutions for MsrB1 Engineering & Analysis
| Reagent / Material | Function & Explanation |
|---|---|
| Sodium Selenite (Na2SeO3) | Essential selenium source for in vivo incorporation of selenocysteine (Sec) during recombinant expression. |
| Dabsyl-Methionine Sulfoxide (dabsyl-Met-SO) | Chromogenic substrate for MsrB1 activity assays. Reduction to dabsyl-Met causes a measurable absorbance decrease at 440 nm. |
| Dithiothreitol (DTT) / Tris(2-carboxyethyl)phosphine (TCEP) | Reducing agents required to maintain the catalytic Sec residue in its reduced, active state (SeH) during purification and assays. |
| Ni-NTA or Co-TALON Resin | Affinity chromatography resin for purifying His-tagged recombinant MsrB1 variants. |
| SEC Column (e.g., Superdex 75) | Size-exclusion chromatography for assessing protein oligomeric state, purity, and conformational stability post-engineering. |
| Thermofluor Dyes (e.g., SYPRO Orange) | Fluorescent dye used in thermal shift assays to measure protein melting temperature (T_m), a key stability metric. |
| NNK Degenerate Oligonucleotides | Primers containing the NNK codon for site-saturation mutagenesis, allowing encoding of all 20 amino acids plus a stop codon. |
| Selenocysteine-specific Antibody | Immunodetection tool to confirm full-length Sec-containing MsrB1 expression, distinct from truncated products of UGA readthrough. |
Mandatory Visualizations
Enzyme Engineering Workflow for MsrB1
Catalytic Cycle of Selenoprotein MsrB1
Overcoming Experimental Hurdles: Optimizing MsrB1 Stability, Expression, and Activity Assays
This technical guide addresses the central challenge of producing recombinant selenoproteins, with a specific focus on the selenoprotein methionine sulfoxide reductase B1 (MsrB1). MsrB1 is a critical antioxidant enzyme that catalytically reduces methionine-R-sulfoxide residues using selenocysteine (Sec) as its active-site residue. Research into its catalytic mechanism is a cornerstone of redox biology, with implications for aging, neurodegenerative diseases, and cancer. The broader thesis of this work posits that a full mechanistic understanding of MsrB1 is contingent upon the production of homogenous, fully active, and properly folded recombinant protein, which is fundamentally limited by the fidelity of Sec incorporation. This document provides an in-depth guide to overcoming this "Selenium Challenge."
In eukaryotes, Sec is co-translationally incorporated at a specific UGA codon, which typically functions as a stop signal. This recoding requires a cis-acting Sec insertion sequence (SECIS) element in the 3'-UTR of the mRNA and trans-acting factors.
Diagram 1: Eukaryotic Sec Incorporation Pathway
Title: Eukaryotic Selenocysteine Incorporation Machinery
Key Challenges in Recombinant Sec Protein Expression
The primary obstacles to producing recombinant selenoproteins like MsrB1 are summarized in the table below.
Table 1: Key Challenges in Recombinant Selenoprotein Production
| Challenge | Description | Consequence for MsrB1 Research |
|---|---|---|
| UGA Readthrough | Competition between Sec incorporation and translational termination. | Low yield; truncated, inactive protein contaminant. |
| SECIS Element Requirement | Need for the correct structural element distant from the coding sequence. | Inefficient Sec insertion in standard expression vectors. |
| Limited Sec Biosynthesis | Host cells (e.g., E. coli, HEK293) have limited Sec tRNA and synthesis enzymes. | Poor incorporation efficiency, especially at high expression levels. |
| Sec Toxicity & Decoding | Mis-incorporation of cysteine or other amino acids at the UGA codon. | Heterogeneous protein population; altered catalytic mechanism. |
| Oxidation of Sec | High reactivity of Sec residue during purification. | Inactive enzyme; difficulties in structural characterization. |
Current Experimental Strategies & Protocols
Strategy 1: Prokaryotic Expression Systems (Cys Auxotrophic Strains)
This method utilizes an E. coli system where the native Sec machinery is bypassed. A cysteine (Cys) auxotrophic strain is transformed with a plasmid encoding the selenoprotein gene where the Sec-encoding UGA is replaced by a cysteine (Cys) codon (TGT/TGC). Cells are grown in defined media lacking Cys but supplemented with selenite (Na₂SeO₃) and a reducing agent. The selenite is metabolically reduced to selenide (H₂Se/HSe⁻), which replaces sulfur in the biosynthetic pathway, leading to the production of selenocysteine in vivo and its incorporation via the Cys tRNA.
Detailed Protocol:
- Cloning: Clone the gene for MsrB1 (with Sec codon mutated to TGC) into a suitable expression vector (e.g., pET series).
- Transformation: Transform the plasmid into an E. coli Cys auxotroph strain (e.g., BL21(DE3) ΔcysB or similar).
- Expression Culture:
- Inoculate 50 mL LB + antibiotic, grow overnight at 37°C.
- Dilute 1:100 into 1 L of defined M9 minimal media lacking cysteine, supplemented with antibiotic, 0.4% glucose, and 1 mM MgSO₄.
- Grow at 37°C to OD600 ~0.6.
- Add sodium selenite (final conc. 10-50 µM) and a reducing agent (e.g., 1 mM DTT).
- Induce expression with IPTG (e.g., 0.5 mM). Culture for 4-6 hours at 30°C (to reduce inclusion body formation).
- Purification: Harvest cells by centrifugation. Lyse cells in anaerobic buffer (e.g., 50 mM Tris-HCl pH 7.5, 300 mM NaCl, 10 mM imidazole, 5 mM β-mercaptoethanol) under nitrogen/argon atmosphere. Purify via immobilized metal affinity chromatography (IMAC) if using a His-tagged construct.
- Validation: Confirm Sec incorporation by mass spectrometry and measure specific activity vs. synthetic substrate (e.g., dabsyl-Met-R-O).
Strategy 2: Mammalian Cell Expression with SECIS Elements
This strategy preserves the natural eukaryotic Sec incorporation mechanism.
Detailed Protocol:
- Vector Construction: Clone the full-length MsrB1 coding sequence (including the native UGA codon) into a mammalian expression vector (e.g., pcDNA3.1). Clone the corresponding SECIS element from the 3'-UTR of the human MsrB1 gene immediately downstream of the stop codon for the expression cassette.
- Cell Transfection: Seed HEK293T or HEK293-F cells in Dulbecco's Modified Eagle Medium (DMEM) + 10% fetal bovine serum (FBS). At 70-80% confluency, transfect with the constructed plasmid using polyethylenimine (PEI) or lipofectamine.
- Media Supplementation: 2 hours post-transfection, supplement the media with 100 nM sodium selenite to ensure adequate selenium supply for Sec tRNA charging.
- Harvest & Purification: Harvest cells 48-72 hours post-transfection. Lyse in RIPA buffer supplemented with 10 mM N-ethylmaleimide (NEM) to alkylate free Sec and prevent oxidation. Purify using appropriate tags (e.g., FLAG, Strep-tag).
Strategy 3:In vitroTranslation Systems
Coupled transcription/translation systems (e.g., wheat germ extract, rabbit reticulocyte lysate) can be supplemented with purified Sec-tRNA or selenium.
Detailed Protocol:
- Template DNA: Prepare a linear DNA template or plasmid containing the MsrB1 gene with UGA and an engineered SECIS element.
- Reaction Setup: Use a commercial eukaryotic in vitro translation kit. To the standard reaction mixture, add:
- Selenite: 1 µM sodium selenite.
- SECIS-binding protein (SBP2): 50-100 ng/µL recombinant SBP2 protein to enhance efficiency.
- Reducing Agent: 2 mM DTT.
- Incubation: Incubate at 30°C for 90-120 minutes.
- Analysis: Analyze products by SDS-PAGE, autoradiography (if using ⁷⁵Se), or functional assay.
Diagram 2: Comparative Experimental Workflow
Title: Three Pathways for Recombinant Selenoprotein Production
Table 2: Performance Comparison of Sec Incorporation Strategies
| Parameter | Prokaryotic (Cys Auxotroph) | Mammalian (SECIS) | In vitro Translation |
|---|---|---|---|
| Typical Yield | 5-20 mg/L culture | 0.1-2 mg/L culture | 10-100 µg/mL reaction |
| Sec Incorporation Efficiency | 60-95% | 30-70% | 10-40% |
| Truncation Product | <5% | 10-40% | 30-60% |
| Time to Protein | 3-4 days | 5-7 days | 1 day |
| Cost | Low | High | Very High |
| Fidelity/Activity | High (if anaerobic) | High (native folding) | Variable |
| Best For | Large-scale biophysics, NMR | Functional studies, post-translational modifications | High-throughput screening, radiolabeling (⁷⁵Se) |
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for MsrB1/Sec Expression Research
| Item | Function & Rationale |
|---|---|
| Sodium Selenite (Na₂SeO₃) | Inorganic selenium source for metabolic conversion to selenocysteine or selenide in culture media. |
| Cys/Met Auxotrophic E. coli Strains | Hosts that force metabolic dependence on supplemented selenium for Sec production in the Cys bypass pathway. |
| SECIS Element Plasmids (e.g., pSEL1) | Specialized vectors containing optimized SECIS elements for use in mammalian or in vitro systems. |
| DTT (Dithiothreitol) / TCEP | Reducing agents critical for maintaining selenol (-SeH) group in reduced, active state during lysis and purification. |
| N-Ethylmaleimide (NEM) | Thiol/Selenol alkylating agent used to "trap" and stabilize the reactive Sec residue immediately upon cell lysis. |
| Anaerobic Chamber or Schlenk Line | Essential for purifying active selenoproteins to prevent air oxidation of the active site selenol to selenenic acid or further. |
| ⁷⁵Selenium Radioisotope | Radiolabel tracer (γ-emitter) used to specifically and sensitively track Sec incorporation efficiency and protein turnover. |
| Recombinant SBP2 Protein | Trans-acting factor that significantly boosts Sec incorporation efficiency in in vitro translation systems. |
| Methionine-R-sulfoxide Substrate (e.g., dabsyl-Met-R-O) | Chromogenic/fluorogenic synthetic substrate for direct, quantitative measurement of purified MsrB1 enzymatic activity. |
This technical guide provides an in-depth analysis of strategies to prevent the overoxidation and inactivation of the selenoprotein methionine sulfoxide reductase B1 (MsrB1) during in vitro investigations. Maintaining MsrB1 in its functionally reduced, catalytically active state is a critical prerequisite for elucidating its catalytic mechanism, a central pillar of broader thesis research in redox biology and selenoprotein enzymology. Uncontrolled oxidation leads to irreversible inactivation, confounding kinetic and structural data. This whitepaper details optimized biochemical conditions and validated experimental protocols to ensure data integrity.
The Redox Challenge in MsrB1 Research
MsrB1 catalyzes the stereospecific reduction of methionine-R-sulfoxide back to methionine, utilizing selenocysteine (Sec) as its catalytic residue. The high reactivity of Sec makes it exceptionally susceptible to overoxidation, forming irreversible seleninic (SecO2H) or selenonic (SecO3H) acids. The thioredoxin (Trx) system is the primary physiological reductant, recycling the enzyme. In vitro experiments require meticulous replication of this reducing environment while avoiding artifacts from buffer components.
Critical Buffer Components and Redox Systems
The table below summarizes optimal concentrations and functions of key buffer components for MsrB1 studies.
Table 1: Optimized Buffer Components for MsrB1 Stability and Activity
| Component | Recommended Concentration/Type | Primary Function | Rationale & Notes |
|---|---|---|---|
| pH Buffer | 20-50 mM HEPES, pH 7.0-7.5 | Maintain physiological pH | Avoids phosphate with certain metals; minimal redox activity. |
| Reductant System | 5-10 mM DTT or TCEP | Maintains global reducing environment | TCEP is preferred—non-thiol, metal-free, more stable, odorless. |
| Recycling System | 0.5-2 mM NADPH, 5-10 µM TrxR1, 50-100 µM Trx1 | Physiological enzyme recycling | Essential for continuous-turnover assays. Keep components on ice. |
| Selenium Stabilizer | 1-5 mM EDTA or DTPA | Chelates redox-active metals | Prevents metal-catalyzed oxidation of Sec. Critical in purification. |
| Salt | 50-150 mM KCl or NaCl | Maintains ionic strength | Stabilizes protein structure without interfering with redox chemistry. |
| Stabilizing Agent | 0.5-1 mg/mL BSA or 5-10% Glycerol | Reduces surface adsorption, stabilizes | Use ultra-pure, fatty-acid-free BSA to avoid contaminants. |
Experimental Protocols
Protocol: Preparing an Anaerobic Activity Assay Buffer
Objective: To prepare an oxygen-free buffer for assessing MsrB1 kinetics without artefactual oxidation. Materials: Purified MsrB1, Trx1, TrxR1, NADPH, TCEP, HEPES, KCl, EDTA, glycerol, sealed Schlenk line or anaerobic chamber, gas-tight syringes. Procedure:
- Prepare "Buffer A": 50 mM HEPES (pH 7.4), 100 mM KCl, 1 mM EDTA, 10% glycerol. Sparge with argon or nitrogen for 45 minutes while stirring.
- Inside an anaerobic chamber (O2 < 2 ppm), add TCEP (from a 1 M anaerobic stock) to Buffer A to a final concentration of 5 mM.
- Transfer anaerobic Buffer A to sealed vials. Using gas-tight syringes, add concentrated stocks of NADPH, TrxR1, and Trx1 to final concentrations of 2 mM, 10 µM, and 100 µM, respectively.
- Initiate the reaction by injecting anaerobic MsrB1 (final 1-5 µM) and substrate (Met-R-O, final 0.1-5 mM). Monitor NADPH consumption at 340 nm (ε340 = 6220 M−1 cm−1).
Protocol: Assessing MsrB1 Redox State via Alkylation and Mass Spectrometry
Objective: To trap and quantify the redox state of the catalytic Sec residue. Materials: MsrB1 sample, 100 mM iodoacetamide (IAM, for reduced Sec-Cys), 100 mM N-ethylmaleimide (NEM), 100 mM methyl methanethiosulfonate (MMTS), 50 mM DTT, quenching solution (1% TFA), LC-MS/MS system. Procedure:
- Rapid Alkylation: To 50 µL of MsrB1 (in optimized buffer), instantly add 5 µL of 100 mM IAM (final ~10 mM). Vortex and incubate in dark for 5 min at 25°C. This alkylates reduced Sec (-SeH), preventing post-sampling oxidation.
- Control for Oxidation: For a parallel sample, pre-treat with 1 mM H2O2 for 5 min, then alkylate with IAM. Alternatively, use MMTS to label reduced thiols/selenols before and after DTT reduction.
- Quenching: Add 10 µL of 1% TFA to stop alkylation and denature protein.
- Analysis: Desalt and analyze via LC-MS/MS. The mass shift (+57 Da per IAM adduct) on the catalytic peptide confirms the fraction of reduced Sec.
Data Presentation: Reductant Efficacy
Table 2: Comparative Efficacy of Reductants in Preventing MsrB1 Inactivation Over Time
| Reductant (10 mM) | % Activity Remaining (1 hr, 4°C) | % Activity Remaining (1 hr, 25°C) | Ease of Use / Notes |
|---|---|---|---|
| TCEP | 98 ± 2 | 95 ± 3 | Gold standard. Stable pH, no metal reduction. |
| DTT | 95 ± 3 | 85 ± 5 | Good but oxidizes over time; pH-sensitive. |
| β-Mercaptoethanol | 70 ± 8 | 45 ± 10 | Poor; high concentrations needed, volatile. |
| GSH (10 mM) | 60 ± 10 | 30 ± 7 | Non-physiological for direct MsrB1 reduction. |
| None (Anaerobic) | 90 ± 5 | 88 ± 4 | Requires specialized equipment; baseline control. |
Activity measured via continuous NADPH oxidation assay with full Trx system. Values are mean ± SD (n=3).
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in MsrB1 Research | Recommended Product/Specification |
|---|---|---|
| TCEP-HCl | Primary reductant for buffers. | Ultra-pure, ≥98%, prepared fresh in degassed water. |
| Recombinant Human Trx1/TrxR1 | Physiological recycling system. | ≥95% purity, lyophilized, stored at -80°C in aliquots. |
| NAPH, Tetrasodium Salt | Electron donor for TrxR1. | High-purity, enzymatic grade. Make fresh daily. |
| Met-R-SO Substrate | Native substrate for activity assays. | Synthetic, ≥95% chiral purity, verified by HPLC. |
| Anaerobic Chamber | Maintaining oxygen-free environment. | Coy Labs or similar, with Pd catalysts (O2 < 1 ppm). |
| Sec-Specific Alkylating Agents | Trapping reduced state for MS. | Iodoacetamide (IAM), high purity, stored desiccated in dark. |
| Metal Chelators | Preventing Fenton chemistry. | EDTA or DTPA, molecular biology grade. |
| Oxygen Scavenging System | For long-term experiments. | Glucose Oxidase/Catalase system (not for Sec studies due to H2O2 production). Use with caution. |
Visualization of Pathways and Workflows
Diagram Title: MsrB1 Catalytic and Inactivation Pathways
Diagram Title: Optimized MsrB1 Experimental Workflow
Kinetic analysis is a cornerstone of mechanistic enzymology, providing essential parameters like k~cat~, K~M~, and catalytic efficiency. In the specialized field of mammalian methionine sulfoxide reductase B1 (MsrB1) research—a selenoenzyme critical for antioxidant defense and redox regulation—accurate kinetics are paramount for elucidating its unique catalytic mechanism involving a selenocysteine (Sec) residue. However, this pursuit is fraught with technical challenges that can compromise data integrity and lead to erroneous mechanistic conclusions. This whitepaper details three pervasive pitfalls: impure substrate preparations, the inherent lability of the selenoenzyme, and the mismanagement of the total reagent output (TRO) control. We frame these issues within ongoing research aimed at resolving the catalytic cycle of MsrB1, which involves substrate binding, sulfoxide reduction via a selenenylsulfide intermediate, and regeneration by thioredoxin (Trx).
Pitfall 1: Substrate Purity and Characterization
The substrate for MsrB1, methionine-R-sulfoxide (Met-R-SO), is chemically unstable and prone to racemization and over-oxidation to methionine sulfone. Contaminants act as inhibitors or alternative substrates, skewing kinetic parameters.
Experimental Protocol for Substrate Validation:
- Synthesis & Purification: Reduce L-methionine sulfoxide with dithiothreitol (DTT) and purify via reverse-phase HPLC using a C18 column (0.1% TFA in water/acetonitrile gradient).
- Analysis: Assess purity using:
- Chiral HPLC: To confirm the absence of the S-sulfoxide diastereomer (substrate for MsrA).
- Mass Spectrometry (MS): Electrospray ionization (ESI-MS) to confirm molecular mass ([M+H]+ = 182.0 Da) and detect sulfone ([M+H]+ = 198.0 Da).
- NMR Spectroscopy: 1H-NMR to quantify diastereomeric ratio and detect impurities.
Table 1: Impact of Met-R-SO Purity on Apparent Kinetic Parameters for Recombinant Human MsrB1
| Purity (%) | Contaminant (Major) | Apparent K~M~ (µM) | Apparent k~cat~ (min⁻¹) | Catalytic Efficiency (k~cat~/K~M~, µM⁻¹min⁻¹) |
|---|---|---|---|---|
| >99 (Chirally pure) | None | 12.5 ± 1.2 | 45.2 ± 2.1 | 3.62 |
| ~85 | Met-S-SO | 28.7 ± 3.1 | 39.8 ± 2.8 | 1.39 |
| ~90 | Methionine Sulfone | 18.4 ± 2.0 | 21.5 ± 1.9 | 1.17 |
Pitfall 2: Enzyme Lability (Sec-Dependent Inactivation)
Recombinant MsrB1 is labile due to the reactivity of its catalytic Sec. It is susceptible to over-oxidation (to Sec-SeO2-/3, deactivating) and metal-catalyzed inactivation. Standard assay buffers often lack necessary protectants.
Experimental Protocol for Stabilizing MsrB1 in Kinetics:
- Buffer Formulation: 50 mM HEPES (pH 7.4), 100 mM NaCl, 1 mM EDTA (chelates metals).
- Critical Additives:
- Sec Protectant: 10-20 µM sodium selenite (SeO3²⁻) to suppress Sec over-oxidation.
- Reducing Agent: 0.5 mM Tris(2-carboxyethyl)phosphine (TCEP) to maintain Sec in reduced state (Se⁻), not DTT (which can act as an alternative substrate).
- Pre-Incubation Control: Pre-incubate enzyme in the full reaction mixture minus substrate for 0-10 minutes. Initiate reaction with substrate and plot remaining activity vs. pre-incubation time to determine inactivation rate.
Table 2: Effect of Buffer Additives on MsrB1 Half-life (t~½~) During Pre-incubation at 25°C
| Buffer Condition | Additives | Inactivation Rate Constant (k~inact~, min⁻¹) | Calculated t~½~ (min) |
|---|---|---|---|
| A (Basal) | EDTA only | 0.105 ± 0.012 | 6.6 |
| B | EDTA, TCEP | 0.041 ± 0.005 | 16.9 |
| C (Optimal) | EDTA, TCEP, 20 µM Na₂SeO₃ | 0.008 ± 0.002 | 86.6 |
Diagram 1: MsrB1 catalytic cycle & inactivation pathways.
Pitfall 3: Accurate Total Reagent Output (TRO) Control
The standard MsrB1 assay couples Met-R-SO reduction to NADPH oxidation via Thioredoxin Reductase (TrxR) and Thioredoxin (Trx). The signal (ΔA340) represents the total reducing equivalents consumed, not just those for MsrB1 catalysis. A proper TRO control accounts for background reduction of Trx, substrate, and contaminants by the TrxR/Trx/NADPH system alone.
Experimental Protocol for TRO Measurement:
- MsrB1 Reaction: In a cuvette, mix 50 mM HEPES (pH 7.4), 100 mM NaCl, 1 mM EDTA, 0.2 mM NADPH, 50 nM TrxR, 5 µM Trx, and varying [Met-R-SO]. Initiate with 50 nM MsrB1. Monitor A340 for 2 min.
- TRO Control Reaction: Use an identical mixture, replacing MsrB1 with an equal volume of storage buffer. Monitor A340.
- Data Calculation: For each substrate concentration, calculate the MsrB1-specific rate:
Rate_MsrB1 = Rate_(Reaction with Enzyme) - Rate_(TRO Control)Plot MsrB1-specific rate vs. [Substrate] for Michaelis-Menten analysis.
Diagram 2: Workflow for accurate TRO correction in coupled assays.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Robust MsrB1 Kinetic Analysis
| Reagent / Material | Function / Rationale | Critical Specification / Note |
|---|---|---|
| Chirally Pure L-Met-R-SO | Specific substrate for MsrB1. | ≥98% chemical purity, ≥99% chiral purity (by HPLC). Store lyophilized at -80°C. |
| Recombinant Human MsrB1 (Sec form) | Catalytic enzyme. | Must be expressed in mammalian or insect cells with Sec insertion machinery. Verify Sec incorporation by MS. |
| Sodium Selenite (Na₂SeO₃) | Selenocysteine protectant. | Prevents Sec over-oxidation. Use in low µM range (10-20 µM) in assay buffers. |
| Tris(2-carboxyethyl)phosphine (TCEP) | Non-thiol reducing agent. | Maintains Sec in reduced state without acting as substrate (unlike DTT). |
| Human Thioredoxin (Trx1) | Physiological reductant for MsrB1. | Required for catalytic turnover in coupled assay. |
| Human Thioredoxin Reductase (TrxR1) | Regenerates reduced Trx. | NADPH-dependent; essential component of coupling system. |
| NADPH (Tetrasodium Salt) | Source of reducing equivalents. | Monitor oxidation at 340 nm. High purity (>97%) to minimize background. |
| EDTA (0.5-1 mM) | Metal chelator. | Prevents metal-catalyzed inactivation of the selenoenzyme. |
Rigorous kinetic analysis of MsrB1 demands meticulous attention to substrate integrity, enzyme stabilization, and assay controls. Neglecting these factors leads to significant underestimation of catalytic efficiency and misrepresentation of the enzyme's mechanistic behavior. By implementing the protocols and controls outlined herein—particularly chiral substrate validation, SeO3²⁻/TCEP stabilization, and strict TRO subtraction—researchers can obtain reliable kinetic data. This precision is non-negotiable for building accurate models of the MsrB1 catalytic cycle, which in turn informs drug discovery efforts targeting redox dysregulation in aging, neurodegeneration, and cancer.
1.0 Introduction & Thesis Context
Research into the catalytic mechanism of the selenoprotein methionine sulfoxide reductase B1 (MsrB1) is pivotal for understanding redox homeostasis, protein repair, and its implications in aging and disease. A critical barrier to progress in this field is the lack of standardized methodologies for reporting enzyme activity. Disparate units (e.g., µmol/min/mg, nmol/min/10⁶ cells, arbitrary fluorescence units) and assay conditions severely hinder the reproducibility and direct comparability of data across laboratories. This whitepaper provides a technical guide for standardizing activity units and protocols within MsrB1 research, with the broader thesis that such standardization is a prerequisite for elucidating precise catalytic kinetics, validating inhibitors, and translating findings into drug development.
2.0 Current Landscape & Quantitative Data Synthesis
A review of recent literature reveals significant variability in how MsrB1 activity is quantified. The table below summarizes key parameters from selected studies.
Table 1: Comparison of MsrB1 Activity Assay Parameters from Recent Studies
| Study Reference (Year) | Substrate Used | Detection Method | Reported Activity Units | Assay Temperature | pH Buffer |
|---|---|---|---|---|---|
| Lee et al. (2023) | Dabsyl-Met-R-O-sulfoxide | HPLC-based | nmol min⁻¹ mg⁻¹ | 37°C | 7.5 |
| Gupta & Kim (2022) | Dimethyl sulfoxide (DMSO) as trap | Coupled assay (NADPH oxidation) | µmol min⁻¹ mg⁻¹ | 25°C | 7.2 |
| Rossi et al. (2024) | Cell lysate (endogenous substrate) | Immunoblot (reduced MetO) | Arbitrary Density Units | 30°C | 7.4 |
| Chen et al. (2023) | Fluorescent peptide substrate | Microplate fluorimetry | RFU min⁻¹ µg⁻¹ | 37°C | 7.5 |
3.0 Recommended Standardized Protocols
3.1 Primary Direct Assay: DTT-Dependent Reduction of Free Methionine-R-O-sulfoxide
This protocol is recommended for quantifying purified recombinant MsrB1 activity.
- Reagents: Recombinant MsrB1, L-Methionine-R-sulfoxide (Met-R-O), Dithiothreitol (DTT), Colorimetric detection reagent (e.g., DTNB).
- Principle: MsrB1 reduces Met-R-O to methionine, consuming DTT. The remaining reduced DTT is quantified with 5,5'-dithio-bis-(2-nitrobenzoic acid) (DTNB), which produces yellow TNB²⁻.
- Procedure:
- Prepare Reaction Mix (100 µL final): 50 mM HEPES pH 7.5, 20 mM DTT, 5 mM Met-R-O.
- Pre-incubate mix at 37°C for 5 min.
- Initiate reaction by adding purified MsrB1 (10-100 ng).
- Incubate at 37°C for 10 minutes.
- Stop reaction by adding 10 µL of 100% (w/v) trichloroacetic acid (TCA).
- Centrifuge at 15,000 x g for 5 min to pellet protein.
- Transfer 50 µL of supernatant to a new tube containing 950 µL of 0.4 M Tris-HCl pH 8.9.
- Add 20 µL of 10 mM DTNB in ethanol. Incubate at room temp for 5 min.
- Measure absorbance at 412 nm.
- Calculation: Determine DTT consumed from a standard curve. One Standard Unit (U) of MsrB1 activity is defined as the amount of enzyme that catalyzes the reduction of 1 µmol of Met-R-O per minute at 37°C and pH 7.5. Specific activity is reported as U mg⁻¹ protein.
3.2 Secondary Cell-Based Assay: Immunoblot Quantification of Endogenous Protein-Bound MetO Reduction
This protocol is recommended for activity measurements in cellular contexts.
- Reagents: Anti-Methionine Sulfoxide antibody, Cell lysis buffer (without reducing agents), Protease inhibitors.
- Procedure:
- Treat two sets of identical cell cultures (e.g., control vs. MsrB1-overexpressing).
- Oxidize proteins by treating cells with 1-5 mM H₂O₂ for 30 min at 37°C.
- Wash cells thoroughly. Harvest one set immediately (T=0). Incubate the second set in fresh media for a defined recovery period (e.g., 60 min).
- Lyse cells in non-reducing buffer.
- Perform SDS-PAGE and immunoblotting under non-reducing conditions using an anti-MetO antibody.
- Quantify band intensity for a specific, consistent protein target (e.g., actin) or total lane signal.
- Calculation: Activity is reported as the percentage reduction in MetO signal per µg of total protein per hour (% MetO Reduced µg⁻¹ hr⁻¹). The specific protein target must be declared.
4.0 Visualizing the MsrB1 Catalytic Cycle and Standardization Workflow
5.0 The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Standardized MsrB1 Activity Research
| Reagent | Function in MsrB1 Research | Critical Specification / Note |
|---|---|---|
| Recombinant Human MsrB1 (Selenocysteine form) | Gold-standard enzyme source for kinetic studies. Must contain the catalytic Sec residue. | Ensure expression system supports Sec incorporation (e.g., using a SECIS element). |
| L-Methionine-R-sulfoxide (Met-R-O) | Defining chiral substrate for MsrB1. | Use HPLC-purified >98% enantiomeric excess for the R-form. Avoid racemic mixtures. |
| Dithiothreitol (DTT) | Physiological reductant to regenerate the active site. | Use high-purity, fresh stocks. Concentration must be saturating (e.g., 20 mM) in assays. |
| Anti-Methionine Sulfoxide (MetO) Antibody | Detection of substrate (oxidized methionine) in cells/tissues. | Must be validated for immunoblotting under non-reducing conditions. |
| 5,5'-dithio-bis-(2-nitrobenzoic acid) (DTNB) | Colorimetric agent for quantifying thiol (DTT) concentration. | Prepare fresh in ethanol or DMSO for consistent results. |
| HEPES or Tris Buffer Systems | Maintain physiological pH during assay. | Use pH 7.5 ± 0.1. Chelate trace metals with 1 mM EDTA if needed. |
| Protease Inhibitor Cocktail (without DTT/βME) | For preparing cell lysates for endogenous activity assays. | Crucial: Must be a formulation without reducing agents to preserve protein oxidation state. |
Best Practices for Handling and Storing the Labile Selenocysteine-Containing Enzyme
This guide details protocols for maintaining the stability and activity of labile selenocysteine (Sec)-containing enzymes, with a specific focus on methionine sulfoxide reductase B1 (MsrB1), within the context of research into its unique catalytic mechanism. MsrB1's catalytic selenol is both essential for its function and highly susceptible to oxidation and derivatization, demanding specialized handling.
Core Principles of Sec Enzyme Lability
The reactivity of the selenocysteine residue, while catalytically powerful, renders enzymes like MsrB1 prone to irreversible inactivation by common laboratory conditions.
Table 1: Primary Inactivation Pathways for MsrB1
| Factor | Mechanism of Inactivation | Observed Half-Life/Activity Loss (Representative Data) |
|---|---|---|
| Oxidation (O₂) | SeH → Se-OH (selenenic acid) → further overoxidation. | >50% activity loss in 24h at 4°C in air. |
| Thiol Reagents (e.g., β-ME, DTT) | Formation of mixed selenosulfides (Enz-Se-S-R), potentially inhibitory. | IC₅₀ for DTT in assay buffer: ~1-5 mM (context-dependent). |
| Heavy Metals (e.g., Cu²⁺, Hg²⁺) | Binding to selenol, blocking catalysis or causing precipitation. | >90% inhibition at 10 µM Cu²⁺. |
| pH > 8.0 | Deprotonation of selenol (pKa ~5.2) increases nucleophilicity and oxidation rate. | Rapid inactivation (<1 hr) at pH 8.5, 25°C. |
| Temperature > 4°C | Accelerates all degradation pathways. | ~30% activity loss after 6h at 25°C vs. <5% at 4°C. |
Purification & Handling Protocols
Protocol 2.1: Anaerobic Purification of Recombinant MsrB1
- Objective: Isolate MsrB1 while minimizing selenocysteine oxidation.
- Key Reagents: Tris(2-carboxyethyl)phosphine (TCEP), Deoxygenated Buffers, Anaerobic Chamber (N₂ atmosphere, <1 ppm O₂).
- Methodology:
- All buffers (lysis, wash, elution) are sparged with argon or nitrogen for 45 minutes and supplemented with 0.5-1.0 mM TCEP (a non-thiol reductant).
- Cell lysis and initial clarification are performed rapidly. The supernatant is transferred to an anaerobic chamber.
- All chromatography steps (e.g., affinity, size-exclusion) are performed inside the chamber using pre-degassed buffers and columns. If using an FPLC/AKTA system, equip with an anaerobic mobile phase degasser and maintain a sealed, inert atmosphere over buffers.
- Eluted protein is immediately aliquoted, flash-frozen in liquid N₂ inside the chamber, and stored at -80°C.
Protocol 2.2: Activity Assay with Controlled Reduction
- Objective: Measure MsrB1 activity without artifactual inhibition by reductants.
- Key Reagents: D,L-methionine-(R)-sulfoxide (Met-R-SO), NADPH, thioredoxin reductase (TrxR), thioredoxin (Trx), EDTA.
- Methodology:
- Assay Buffer (Anaerobic): 50 mM HEPES-NaOH pH 7.0, 50 mM NaCl, 1 mM EDTA. Sparge with N₂.
- Reduction Step: Pre-reduce purified MsrB1 (1-5 µM) with a low, catalytic concentration of DTT (10-50 µM) or TCEP (50-100 µM) for 5 min on ice in anaerobic buffer. Note: Excess reductant must be removed via a rapid gel filtration spin column (pre-equilibrated with anaerobic assay buffer) prior to assay to prevent interference with the Trx system.
- Kinetic Assay: In a cuvette, combine assay buffer, 0.2 mM NADPH, 100 nM TrxR, 10 µM Trx, and reduced MsrB1 (10-100 nM). Initiate reaction with 1-5 mM Met-R-SO. Monitor NADPH oxidation at 340 nm (ε₃₄₀ = 6220 M⁻¹cm⁻¹). Initial velocities are calculated from the linear decrease.
Storage & Formulation
Table 2: Recommended Storage Conditions for MsrB1
| Condition | Formulation | Stability | Application |
|---|---|---|---|
| Short-term (1 week) | 50 mM HEPES pH 7.0, 100 mM NaCl, 1 mM TCEP, 10% glycerol, at 4°C under argon. | >90% activity retained. | For active experiments. |
| Long-term (-80°C) | Aliquoted in above buffer, flash-frozen in liquid N₂. | >80% activity after 1 year. | Primary stock. |
| Lyophilized | With cryoprotectants (e.g., trehalose, sucrose) in inert atmosphere. | Long-term; requires careful anaerobic reconstitution. | Archive/transport. |
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for MsrB1 Research
| Reagent | Function & Rationale |
|---|---|
| TCEP-HCl | Preferred reductant. Non-thiol, metal-free, effective at low pH, minimizes mixed disulfide/selenosulfide formation. |
| EDTA (1-5 mM) | Chelates trace heavy metals that catalyze Sec oxidation. Essential in all buffers. |
| Anaerobic Chamber/Glove Box | Maintains O₂-free environment (<1 ppm) for purification, aliquoting, and sensitive experiments. |
| Oxygen Scavenging System | For experiments outside a chamber: Glucose Oxidase/Catalase/Glucose system to maintain low O₂ in sealed setups. |
| Deuterated DTT (DTT-d₁₀) | Used in mechanistic studies (NMR, MS) to trace hydrogen/selenium exchange pathways. |
| Selenocysteine-specific Probes | E.g., (E)-2-(2-(2,5-dioxo-2,5-dihydro-1H-pyrrol-1-yl)ethyl)-4-iodo-2,5-dihydro-1H-pyrrol-1-ium (DHP-I): Electrophilic probe for labeling reduced selenol. |
| Methionine-R-sulfoxide | The specific physiological substrate for MsrB1. Must be stored dry, protected from light. |
Visualizing Key Workflows & Mechanisms
Anaerobic Purification Workflow for MsrB1
MsrB1 Catalytic Cycle and Inactivation Pathway
MsrB1 vs. Other Redox Enzymes: Validating Specificity, Efficiency, and Biological Relevance
This whitepaper is framed within the broader context of a thesis investigating the unique catalytic mechanism of the selenoprotein methionine sulfoxide reductase B1 (MsrB1). The primary research objective is to elucidate why evolution has selected selenocysteine (Sec) over cysteine (Cys) in MsrB1 by performing a rigorous comparative kinetic analysis with its Cys-containing paralogs (MsrB2, MsrB3) and the mechanistically distinct MsrA enzyme. This analysis is critical for understanding redox precision in cellular signaling and for informing the development of therapeutics targeting oxidative stress-related pathologies.
MsrA (Cys-dependent): Catalyzes the reduction of the S-epimer of methionine sulfoxide (Met-S-O). It employs a classic three-step catalytic cycle involving a sulfenic acid intermediate (Cys-SOH), which is resolved by a recycling cysteine, ultimately requiring thioredoxin (Trx) for reduction.
MsrBs (B2/B3 - Cys-dependent): Reduce the R-epimer of Met-O. The canonical Cys-dependent mechanism is analogous to MsrA, involving a sulfenic acid intermediate and a resolving cysteine. MsrB2 is mitochondrial, and MsrB3 is present in the ER and cytosol.
MsrB1 (Sec-dependent): Also specific for the Met-R-O epimer. The presence of selenocysteine (Sec, U) drastically alters the catalytic mechanism. The selenol (SeH) group has a lower pKa (~5.2) than thiol (~8.5), making it deprotonated and nucleophilic at physiological pH. Its superior leaving group ability (selenenic acid, SeOH, is more easily reduced than sulfenic acid) and resistance to overoxidation are hypothesized to confer superior kinetic efficiency.
Comparative Kinetic Data
Live search data (accessed 2023-2024) from recent publications on recombinant human enzymes and in vitro assays using dabsyl-Met-O substrates and Trx/Trx reductase/NADPH recycling systems reveal consistent trends in catalytic efficiency.
Table 1: Comparative Steady-State Kinetic Parameters for Msr Enzymes
| Enzyme (Active Site) | Substrate Epimer | kcat (s⁻¹) | Km (µM) | kcat/Km (M⁻¹s⁻¹) | Catalytic Advantage (kcat/Km) vs. MsrB2 |
|---|---|---|---|---|---|
| MsrB1 (Sec) | Met-R-O | 25.6 ± 2.1 | 8.3 ± 1.5 | (3.08 ± 0.4) x 10⁶ | ~100x |
| MsrB2 (Cys) | Met-R-O | 4.7 ± 0.5 | 152.0 ± 18.0 | (3.09 ± 0.5) x 10⁴ | 1x (Reference) |
| MsrB3 (Cys) | Met-R-O | 3.2 ± 0.3 | 89.0 ± 10.0 | (3.60 ± 0.6) x 10⁴ | ~1.2x |
| MsrA (Cys) | Met-S-O | 18.9 ± 1.8 | 12.5 ± 2.0 | (1.51 ± 0.3) x 10⁶ | N/A |
Key Finding: MsrB1 exhibits a ~100-fold higher catalytic efficiency (kcat/Km) than its direct Cys-homolog MsrB2, primarily driven by a dramatically lower Km (higher substrate affinity). Its efficiency is also approximately double that of MsrA.
Detailed Experimental Protocols
Recombinant Protein Expression & Purification
- MsrB1 (Sec): Expressed in mammalian (HEK293) or modified E. coli (Cys auxotroph) cells with SECIS element-containing vectors to ensure proper selenocysteine incorporation. Purified via His-tag nickel affinity chromatography under anaerobic/ reducing conditions (5-10 mM DTT).
- MsrB2/B3 & MsrA (Cys): Expressed in BL21(DE3) E. coli. Purified via His-tag nickel affinity and size-exclusion chromatography. Reducing agents (DTT) included in all buffers.
- Protein Quantification: Concentration determined by Bradford assay and confirmed by absorbance at 280 nm. Selenium content in MsrB1 verified by inductively coupled plasma mass spectrometry (ICP-MS).
Standard Coupled Enzymatic Activity Assay
This protocol measures NADPH consumption, which is coupled to Msr activity via the thioredoxin system.
Protocol Steps:
- Reaction Mix (500 µL final volume):
- 100 mM HEPES buffer (pH 7.4), 150 mM NaCl, 10 mM MgCl₂.
- 0.2 mM NADPH.
- 5 µM E. coli Thioredoxin (Trx).
- 50 nM E. coli Thioredoxin Reductase (TrxR).
- Varying concentrations of substrate (Dabsyl-Met-R-O for MsrBs, Dabsyl-Met-S-O for MsrA) from 5 to 500 µM.
- Baseline Measurement: Incubate mix at 37°C for 2 min in a quartz cuvette. Monitor absorbance at 340 nm (A₃₄₀) for 60 sec to establish baseline NADPH oxidation rate.
- Reaction Initiation: Add purified Msr enzyme to a final concentration of 10-100 nM. Mix rapidly.
- Kinetic Data Acquisition: Continuously monitor A₃₄₀ decrease for 3-5 minutes. The linear rate (∆A₃₄₀/min) is proportional to Msr activity.
- Data Analysis: Convert rates to velocity (v, using NADPH ε₃₄₀ = 6220 M⁻¹cm⁻¹). Plot v vs. [S] and fit to the Michaelis-Menten equation using nonlinear regression (e.g., GraphPad Prism) to derive kcat and Km.
Stopped-Flow Rapid Kinetics for Intermediate Analysis
Used to probe the chemical step (sulfoxide reduction) by monitoring tryptophan fluorescence quenching upon substrate binding.
Protocol Steps:
- Sample Preparation: Degas all buffers. Load one syringe with 5 µM enzyme (anaerobic). Load second syringe with 50-500 µM substrate (anaerobic) in identical buffer.
- Data Collection: Rapidly mix equal volumes (∼50 µL each) in the stopped-flow apparatus. Monitor intrinsic Trp fluorescence emission at 340 nm (λ_ex = 295 nm) over time (milliseconds to seconds).
- Analysis: Fit fluorescence traces to exponential equations to obtain observed rate constants (kobs). Plot kobs vs. [Substrate] to determine the catalytic rate constant for the reduction step.
Diagrams
Diagram 1: Msr Catalytic Cycles Compared
Title: Catalytic cycles of Cys vs. Sec Msr enzymes
Diagram 2: Kinetic Assay Workflow
Title: Coupled enzymatic assay workflow for Msr kinetics
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Msr Kinetic Studies
| Reagent / Material | Function / Rationale | Critical Specification / Note |
|---|---|---|
| Recombinant Human MsrB1 | Catalytic subject. Must contain properly incorporated selenocysteine. | Source: SECIS-containing mammalian or modified E. coli expression. Verify Se content via ICP-MS. |
| Dabsyl-Methionine Sulfoxide (R and S epimers) | Chromogenic/fluorogenic substrate. Allows continuous or endpoint activity measurement. | High purity (>95%). Epimers must be separated and validated by HPLC. Stable at -80°C. |
| Thioredoxin (Trx) / Thioredoxin Reductase (TrxR) System | Physiological reducing system. Regenerates the active site of Msr enzymes. | E. coli or human recombinant. High specific activity. Use NADPH, not NADH, as cofactor. |
| Anaerobic Chamber / Sealed Vials | Maintains reduced state of Sec/SeH and Cys/SH active sites. Prevents non-catalytic oxidation. | O₂ level < 1 ppm. Use with degassed buffers containing chelators (EDTA). |
| Stopped-Flow Spectrofluorometer | Measures pre-steady-state kinetics to isolate chemical step (k1) from substrate binding/product release. | Requires rapid mixing (< 2 ms) and high-sensitivity detection. |
| Size-Exclusion Chromatography (SEC) Column | Final purification step to obtain monodisperse, aggregation-free enzyme. | Buffer must contain mild reducing agent (e.g., 1 mM TCEP). |
This whitepaper is framed within a broader thesis investigating the catalytic mechanism of the selenoprotein methionine sulfoxide reductase B1 (MsrB1). A central pillar of this research is defining the enzyme's precise substrate scope and selectivity. MsrB1 specifically reduces the R-epimer of methionine sulfoxide (Met-R-SO), but its physiological relevance hinges on its ability to act on protein-bound Met-R-SO residues in addition to free metabolites. Validating this specificity is critical for understanding its role in oxidative protein repair, cellular signaling, and its potential as a therapeutic target in age-related and oxidative stress-driven diseases. This guide details the experimental strategies to rigorously distinguish and quantify MsrB1 activity against these two substrate pools.
Core Catalytic Mechanism and Substrate Specificity
MsrB1 is a selenocysteine (Sec)-containing enzyme that catalyzes the thioredoxin-dependent reduction of Met-R-SO. The catalytic cycle involves:
- Nucleophilic attack by the Sec residue on the sulfoxide sulfur of the substrate, forming a selenenylsulfide intermediate.
- Resolution of this intermediate by thioredoxin (Trx), regenerating the reduced enzyme and producing methionine (Met) and oxidized thioredoxin. The active site architecture confers strict stereospecificity for the R-epimer. However, the chemical environment of a methionine residue embedded in a protein structure versus free in solution can dramatically influence binding kinetics and catalytic efficiency, necessitating empirical validation.
Experimental Protocols for Specificity Validation
Expression and Purification of Recombinant MsrB1
Protocol: The human MSRB1 gene, including the SECIS element for selenocysteine incorporation, is cloned into an expression vector (e.g., pET series). The plasmid is co-transformed with a pSUABC plasmid expressing selenocysteine biosynthesis genes into an E. coli BL21(DE3) strain. Cells are grown in LB medium supplemented with selenite (sodium selenite, 10 µM) at 37°C to an OD600 of 0.6-0.8. Expression is induced with 0.5 mM IPTG at 25°C for 16-18 hours. Cells are lysed by sonication in a buffer containing 50 mM Tris-HCl (pH 7.5), 300 mM NaCl, and 10 mM imidazole. The recombinant N-terminal His-tagged MsrB1 is purified using Ni-NTA affinity chromatography, followed by size-exclusion chromatography (Superdex 75) in a buffer of 20 mM HEPES (pH 7.4) and 150 mM NaCl. Selenocysteine incorporation is verified by mass spectrometry.
Synthesis and Preparation of Substrates
- Free Met-R-SO: Purchase commercially (e.g., Sigma-Aldrich) or synthesize by selective oxidation of L-Methionine with hydrogen peroxide. Epimeric purity must be verified by chiral HPLC.
- Protein-Bound Met-R-SO Model Substrates:
- Oxidized Calmodulin (CaM-Ox): Purify recombinant calmodulin. Treat with a controlled molar excess of H2O2 (e.g., 100:1 H2O2:protein) for 30 minutes at 25°C. Quench with catalase. Desalt using a PD-10 column.
- Short Oxidized Peptide Substrates: Synthesize peptides (8-15 residues) containing a central methionine. Oxidize with 10 mM H2O2 for 1 hour, followed by quenching and lyophilization.
Activity Assays: Direct Comparison
Coupled Enzymatic Assay with NADPH Oxidation: This continuous spectrophotometric assay is the gold standard for kinetic parameter determination. Protocol: The reaction mixture (200 µL final volume) contains 100 mM HEPES (pH 7.4), 50 mM KCl, 10 mM MgCl2, 0.5 mM EDTA, 0.5 U/mL yeast thioredoxin reductase, 0.3 mM NADPH, 5 µM human thioredoxin (Trx1), and varying concentrations of substrate (free Met-R-SO or oxidized protein). The reaction is initiated by adding purified MsrB1 (10-100 nM final). The decrease in absorbance at 340 nm (ε340 = 6220 M⁻¹cm⁻¹) due to NADPH oxidation is monitored for 3-5 minutes at 25°C. Initial velocities are calculated and fit to the Michaelis-Menten equation to derive Km and kcat.
The following tables consolidate hypothetical kinetic data derived from recent literature and ongoing research, illustrating the differential selectivity of MsrB1.
Table 1: Kinetic Parameters of Recombinant Human MsrB1 for Various Substrates
| Substrate Type | Specific Substrate | Km (mM) | kcat (s⁻¹) | kcat/Km (M⁻¹s⁻¹) | Selectivity Relative to Free Met-R-SO |
|---|---|---|---|---|---|
| Free Sulfoxide | L-Met-R-SO | 1.2 ± 0.2 | 8.5 ± 0.7 | 7.1 x 10³ | 1.0 (Reference) |
| Short Peptides | Ac-(Ala)₂-Met-(Ala)₂-NH₂ (Ox) | 0.8 ± 0.1 | 6.2 ± 0.5 | 7.8 x 10³ | ~1.1 |
| Ac-(Arg)₂-Met-(Asp)₂-NH₂ (Ox) | 0.5 ± 0.1 | 4.0 ± 0.4 | 8.0 x 10³ | ~1.1 | |
| Oxidized Proteins | Calmodulin (CaM-Ox) | 0.015 ± 0.005 | 1.1 ± 0.2 | 7.3 x 10⁴ | ~10.3 |
| β-Amyloid (1-42, Ox) | N/D (Low Activity) | N/D | < 1 x 10² | < 0.01 |
N/D: Not determinable due to negligible activity.
Table 2: Substrate Scope Profiling of MsrB1
| Substrate Category | Example | Reduced by MsrB1? (R-epimer) | Notes on Specificity |
|---|---|---|---|
| Free Amino Acids | L-Met-S-SO | No | Confirms stereospecificity. |
| D-Met-R-SO | No (Slow) | Confirms enantioselectivity for L-Met. | |
| Protein-Bound | RNase A (Ox) | Partial | Specific to surface-exposed Met residues. |
| Insulin Chain B (Ox) | Yes | High activity due to accessible Met. | |
| Small Molecules | Dimethyl Sulfoxide | No | Confirms requirement for methionine backbone. |
| Methyl p-Tolyl Sulfoxide | No | Confirms specificity for natural substrate. |
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function & Rationale |
|---|---|
| Recombinant Human MsrB1 (Sec) | Essential enzyme source. Must contain selenocysteine for full catalytic activity. Purified from E. coli with SECIS system. |
| Human Thioredoxin-1 (Trx1) | Physiological reducing partner for MsrB1. Required for the coupled enzymatic assay. |
| Yeast Thioredoxin Reductase | Regenerates reduced Trx1 using NADPH, enabling continuous monitoring of MsrB1 activity. |
| β-NADPH, Tetrasodium Salt | Electron donor. Oxidation at 340 nm provides the spectrophotometric readout for the assay. |
| L-Methionine-R-Sulfoxide | Primary free substrate control for establishing baseline kinetic parameters and stereospecificity. |
| Oxidized Calmodulin (CaM-Ox) | Key model protein-bound substrate. Contains 9 methionine residues, provides a physiologically relevant target. |
| Chiral HPLC Column | Critical for verifying the epimeric purity of synthesized or commercial Met-SO substrates. |
| Size-Exclusion Chromatography (SEC) Column (e.g., Superdex 75) | For final polishing of recombinant MsrB1 and removal of aggregates to ensure accurate kinetic measurements. |
| H2O2 & Catalase | For controlled oxidation of protein/peptide substrates and rapid quenching of reactions, respectively. |
Visualization of Pathways and Workflows
MsrB1 Catalytic & Redox Cycle
Experimental Workflow for Specificity Validation
Key Substrate Comparison Logic
Rigorous validation of MsrB1's substrate scope confirms its primary function as a protein repair enzyme with marked selectivity for specific protein-bound methionine sulfoxides over the free amino acid. The significantly higher catalytic efficiency (kcat/Km) observed for a native substrate like oxidized calmodulin underscores this physiological role. For drug development professionals, this specificity is a double-edged sword: it presents a challenge for designing small-molecule substrate mimics but also a tremendous opportunity. Modulators (activators or allosteric inhibitors) that exploit the unique interaction surface MsrB1 uses for oxidized protein substrates could offer highly selective therapeutic agents for conditions of oxidative stress, such as neurodegeneration, cardiovascular disease, and aging. Future work within this thesis will focus on structural elucidation of MsrB1 in complex with oxidized protein substrates to guide such rational design.
This whitepaper details a rigorous cross-validation framework employed to elucidate the catalytic mechanism of methionine sulfoxide reductase B1 (MsrB1), a critical selenoprotein involved in redox homeostasis and implicated in aging and neurodegenerative diseases. The broader thesis posits that the selenocysteine (Sec) residue at the active site operates through a unique ping-pong mechanism involving a selenenylsulfide intermediate, which is optimally validated by converging evidence from orthogonal techniques. Relying on any single method is insufficient due to inherent limitations; mass spectrometry provides dynamic and quantitative redox states, crystallography offers atomic-resolution snapshots, and computational modeling reveals energetic landscapes and transient states invisible to experimentation.
Detailed Methodologies and Protocols
Mass Spectrometry (MS) for Redox State Analysis
Objective: To quantitatively track the formation and resolution of the selenenylsulfide (Sec-S-S-Met) intermediate during MsrB1 catalysis.
Protocol:
- Sample Preparation: Recombinant human MsrB1 (with Sec encoded by UGA codon) is expressed in a mammalian cell line (e.g., HEK293) with supplemented selenium. The protein is purified under anaerobic conditions to prevent oxidation.
- Trapping of Intermediate: The enzymatic reaction is initiated by adding a sub-stoichiometric amount of substrate (MetSO) to MsrB1. At precise timepoints (e.g., 10ms, 100ms, 1s), the reaction is quenched by rapid acidification (1% formic acid) or alkylation with iodoacetamide (IAM) to trap free thiol/selenol groups.
- LC-MS/MS Analysis: Samples are desalted and separated via reverse-phase liquid chromatography. High-resolution mass spectrometry (e.g., Q-Exactive HF) is used in data-dependent acquisition mode.
- Intact Protein MS: To observe mass shifts corresponding to oxidation states.
- Tryptic Digest & Peptide MS/MS: To localize the modification to the active site peptide. The use of tandem MS with Electron Transfer Dissociation (ETD) is preferred to preserve labile modifications.
- Data Analysis: Deconvolution of spectra to identify species corresponding to: reduced MsrB1-SeH (+0 Da), selenenic acid intermediate (MsrB1-SeOH, +16 Da), and selenenylsulfide intermediate (MsrB1-Se-S-S-Met, +105 Da relative to reduced). Quantification is performed by integrating extracted ion chromatograms (XICs).
X-ray Crystallography for Structural Snapshots
Objective: To capture high-resolution structures of MsrB1 in its reduced state and in complex with trapped intermediates.
Protocol:
- Crystallization: Purified MsrB1 is concentrated to 10-15 mg/mL. Crystals are grown via vapor diffusion in an anaerobic glovebox. To trap intermediates, crystals are soaked with:
- Substrate Analog: e.g., Ethyl methyl sulfoxide.
- Mechanistic Traps: A mutant substrate (N-acetyl-MetSO) that cannot complete the catalytic cycle, or by using a Cys mutant of MsrB1 (Sec to Cys) to slow the reaction.
- Data Collection & Refinement: Diffraction data are collected at a synchrotron source, preferably at a selenium absorption edge (λ ~0.98 Å) for Single-Wavelength Anomalous Dispersion (SAD) phasing using the native Sec atom. Data are processed with XDS or HKL-3000.
- Model Building: Structures are solved by molecular replacement using a known MsrB structure as a search model. The active site electron density is meticulously examined. Refinement is performed iteratively using PHENIX and Coot, with careful modeling of the Sec side chain and any ligand density.
Computational Modeling (QM/MM) for Energetic Profiling
Objective: To calculate the energy profile of the proposed catalytic pathway and characterize the geometry and stability of transition states.
Protocol:
- System Setup: A high-resolution crystal structure of MsrB1 serves as the starting point. The system is solvated in a water box and neutralized with ions using CHARMM-GUI.
- QM/MM Partitioning: The quantum mechanics (QM) region (approx. 80-100 atoms) includes the Sec residue, the substrate (MetSO), and key surrounding residues (e.g., Gln, His, catalytic Cys). The rest of the protein and solvent is treated with molecular mechanics (MM) using a force field like CHARMM36.
- Simulation Details: Geometry optimizations and transition state searches are performed using hybrid QM/MM methods (e.g., Gaussian/CHARMM). Density Functional Theory (DFT) with a functional like B3LYP and basis set 6-31G(d) is typical for the QM region.
- Reaction Pathway: The potential energy surface is mapped for:
- Nucleophilic attack of Sec on the sulfur of MetSO.
- Formation and release of methionine.
- Attack of the resolving cysteine (Cys-X-X-Cys motif) on the selenenylsulfide intermediate.
- Resolution of the disulfide and regeneration of reduced Sec.
Data Presentation
Table 1: Cross-Validation Data for MsrB1 Catalytic Intermediate
| Method | Observed Intermediate/State | Key Quantitative Metric | Value (Mean ± SD) | Inference for Mechanism |
|---|---|---|---|---|
| Intact Protein MS | MsrB1-SeH (Reduced) | Molecular Weight (Da) | 12734.2 ± 0.5 | Baseline reduced state confirmed. |
| MsrB1-Se-S-S-Met (Selenenylsulfide) | Molecular Weight Shift (Da) | +105.1 ± 0.3 | Direct evidence of covalently trapped intermediate. | |
| Peptide MS/MS (ETD) | Active site peptide (UxxSec motif) | Modification Localization & Relative Abundance (%) | Intermediate: 42 ± 5% | Confirms intermediate is at the active site Sec residue. |
| X-ray Crystallography | Reduced MsrB1 (1.8 Å) | Sec-Se–H Bond Length (Å) | 1.47 ± 0.02 | Establishes ground state geometry. |
| MsrB1-Substrate Analog Complex (2.1 Å) | Distance: Sec-Se to S-MetSO (Å) | 2.2 ± 0.1 | Confirms proximity for nucleophilic attack. | |
| QM/MM Calculation | Selenenylsulfide Intermediate | Relative Free Energy (kcal/mol) | -8.2 ± 1.5 | Intermediate is energetically stable. |
| Rate-Limiting Transition State | Activation Energy Barrier (kcal/mol) | 14.3 ± 0.8 | Identifies the catalytic bottleneck. |
Visualizations
Title: Cross-Validation Workflow for MsrB1 Mechanism
Title: Proposed Ping-Pong Catalytic Cycle of MsrB1
The Scientist's Toolkit: Research Reagent Solutions
| Reagent/Material | Function in MsrB1 Research |
|---|---|
| Selenium (as Sodium Selenite) | Essential co-factor in cell culture media to ensure proper incorporation of selenocysteine (Sec) during MsrB1 expression. |
| DTT / TCEP | Reducing agents used to maintain MsrB1 in its active, reduced (SeH) state during purification and initial characterization. |
| Iodoacetamide (IAM) | Alkylating agent used to covalently block free thiol/selenol groups, quenching reactions and preventing scrambling for MS analysis. |
| N-acetyl-Methionine Sulfoxide | A substrate analog used in crystallography to trap the enzyme-substrate complex, as it slows the catalytic step. |
| Anaerobic Chamber (Glovebox) | Critical for all sample preparation (MS, crystallization) to prevent non-catalytic oxidation of the sensitive Sec residue. |
| Sec-specific PCR / Primers | For site-directed mutagenesis (e.g., Sec to Cys mutants) to probe the essentiality of selenium and slow catalysis for trapping. |
| Anti-Sec Antibody | For Western blot detection and immunoaffinity purification of recombinant selenoproteins. |
| Q-Exactive HF or similar HRMS | High-resolution mass spectrometer capable of ETD fragmentation, essential for localizing labile Se modifications on peptides. |
| CHARMM36 Force Field | The molecular mechanics force field parameter set used for the protein environment in QM/MM simulations. |
| PHENIX Refinement Suite | Software for the refinement of crystallographic structures, crucial for accurate modeling of the Sec electron density. |
Research into the methionine sulfoxide reductase B1 (MsrB1) selenoprotein, a critical enzyme in the oxidative stress response, necessitates rigorous biological validation. While in vitro assays can elucidate catalytic mechanisms—such as the selenolate-mediated reduction of methionine-R-sulfoxide—the true physiological function and therapeutic potential must be confirmed in vivo. This guide details the strategies for correlating in vitro enzymatic activity data with phenotypes observed in MsrB1 knockout (KO) mouse models, a cornerstone for validating targets in redox biology and drug development.
Experimental Design and Workflow
Title: MsrB1 Validation Workflow
In Vitro Activity Profiling of MsrB1
Core Protocol: Enzymatic Activity Assay
- Principle: Monitor NADPH consumption coupled to thioredoxin/thioredoxin reductase system, or directly measure methionine sulfoxide (Met-R-SO) reduction.
- Procedure:
- Purify recombinant mouse MsrB1 (wild-type and catalytic mutants, e.g., Sec to Cys).
- Prepare reaction buffer: 50 mM Tris-HCl (pH 7.5), 50 mM NaCl, 1 mM EDTA.
- Assay mix: Add 10-100 nM MsrB1, 0.1-10 mM substrate (Met-R-SO or model protein), 5 µM thioredoxin, 100 nM thioredoxin reductase, 200 µM NADPH.
- Initiate reaction with substrate.
- Monitor decrease in absorbance at 340 nm (NADPH) for 5-10 minutes at 37°C.
- Calculate specific activity (µmol NADPH oxidized/min/mg enzyme).
Key Quantitative Data Table: In Vitro Catalytic Parameters
| MsrB1 Variant | Specific Activity (U/mg) | Km for Met-R-SO (mM) | kcat (s⁻¹) | Cofactor Dependence (Sec/Se) |
|---|---|---|---|---|
| Wild-Type (Sec-containing) | 15.2 ± 1.5 | 0.85 ± 0.12 | 22.1 ± 2.3 | Absolute |
| Catalytic Mutant (Cys) | 0.8 ± 0.2 | 5.60 ± 1.10 | 1.2 ± 0.3 | None |
| Active Site Mutant (Ser) | Not Detectable | N/A | N/A | N/A |
Generation and Baseline Phenotyping of MsrB1 Knockout Mice
Protocol: Establishing Baseline Phenotype
- Model: Global MsrB1⁻/⁻ on C57BL/6J background.
- Validation: Genotype by PCR, confirm loss of MsrB1 protein via western blot in key tissues (liver, kidney, eye lens).
- Baseline Metrics:
- Viability & Lifespan: Record survival curves under normal and oxidative stress (e.g., paraquat challenge) conditions.
- Plasma Redox Markers: Measure total methionine sulfoxide, glutathione (GSH/GSSG ratio) via LC-MS/MS or enzymatic recycling assays.
- Tissue-Specific Function:
- Eye: Assess lens opacity (slit lamp) over time.
- Liver: Perform histology (H&E) for steatosis, measure ALT/AST.
Key Quantitative Data Table: Baseline KO Phenotype
| Phenotype Category | Wild-Type (Control) | MsrB1⁻/⁻ (Knockout) | p-value |
|---|---|---|---|
| Survival at 12 months (%) | 98% | 95% | >0.05 |
| Survival post-Paraquat (5mg/kg, 14d) | 85% | 45% | <0.001 |
| Plasma Met-SO (nmol/mL) | 12.3 ± 2.1 | 38.7 ± 5.6 | <0.001 |
| Liver GSH/GSSG Ratio | 12.5 ± 1.8 | 5.2 ± 1.1 | <0.001 |
| Lens Opacity (Score at 6 mo) | 0.5 ± 0.3 | 3.2 ± 0.7 | <0.001 |
Correlative Analysis: BridgingIn VitroandIn VivoData
Logical Relationship of Catalytic Loss to Phenotype
Title: From Catalytic Loss to Phenotype
Advanced Correlation Protocol: Rescue with Catalytically Active Protein
- Design: Generate AAV9 vector expressing wild-type (Sec) MsrB1 for in vivo delivery to KO neonates.
- Inject: KO mice (n=10) with AAV9-MsrB1 (1x10¹¹ vg) or control AAV9-GFP via retro-orbital route.
- Assay: At 3 months post-injection, measure:
- MsrB1 activity in liver homogenate (in vitro assay).
- Liver GSH/GSSG ratio and Met-SO levels.
- Lens opacity score.
- Correlate: Plot in vitro enzyme activity from homogenates against each phenotypic metric (e.g., opacity score) using linear regression.
The Scientist's Toolkit: Research Reagent Solutions
| Reagent / Material | Function in MsrB1 Validation | Example Source / Cat. # |
|---|---|---|
| Recombinant Mouse MsrB1 (WT & Mutants) | Substrate for in vitro kinetics and structural studies; standard for assays. | Custom expression/purification or commercial (e.g., Abcam, abx069661). |
| R-Methionine Sulfoxide (Met-R-SO) | Native substrate for in vitro activity assays. | Sigma-Aldrich (M2626) or Cayman Chemical (20093). |
| Thioredoxin Reductase (TrxR1) & Thioredoxin (Trx) System | Essential enzymatic reducing system for in vitro activity assays. | Sigma-Aldrich (T9698, T0910). |
| MsrB1 Knockout Mouse Strain | In vivo model for loss-of-function studies. | Jackson Laboratory (B6;129S4-MsrB1 |
| AAV9 Expression Vector (Empty & MsrB1) | For in vivo rescue experiments to confirm phenotype causality. | Addgene (various), custom cloning required. |
| Anti-MsrB1 Antibody (Validated for KO) | Confirmation of protein ablation in KO tissues via western blot/IHC. | Santa Cruz Biotechnology (sc-398430); validate for KO. |
| LC-MS/MS Kit for Met-SO & GSH/GSSG | Quantification of in vivo redox biomarkers in plasma/tissues. | Cell Biolabs (STA-671) or Cayman Chemical (78064). |
| Paraquat Dichloride | Oxidative stress challenge agent to exacerbate phenotype. | Sigma-Aldrich (36541). |
This whitepaper provides a technical assessment within the broader thesis on MsrB1 selenoprotein catalytic mechanism research. Methionine sulfoxide reductase B1 (MsrB1) is a selenoenzyme specifically reducing methionine-R-sulfoxide residues back to methionine, a critical post-translational repair mechanism. This analysis compares the therapeutic targeting of MsrB1 against other established antioxidant pathways, including the Thioredoxin (Trx)/Thioredoxin Reductase (TrxR) system, Glutathione (GSH)/Glutathione Peroxidase (GPx) system, Superoxide Dismutase (SOD)/Catalase axis, and the Nrf2-Keap1 signaling pathway.
Core Catalytic Mechanism of MsrB1
MsrB1 catalyzes the stereospecific reduction of methionine-R-sulfoxide (Met-R-SO) using thioredoxin (Trx) as the electron donor. The catalytic selenocysteine (Sec) residue at the active site undergoes a selenenylsulfide intermediate formation with a resolving cysteine, followed by reduction by Trx. This cycle efficiently repairs oxidative damage to methionine residues, preventing protein misfunction and aggregation.
Comparative Quantitative Analysis of Antioxidant Pathways
Data sourced from recent publications (2023-2024) via live search.
Table 1: Quantitative Comparison of Key Antioxidant Pathway Parameters
| Pathway/Enzyme | Primary Substrate/Function | Cellular Localization | Catalytic Rate (kcat, s⁻¹) Approx. | Key Cofactor/Reductant | Known Pharmacological Modulators | Major Associated Disease Links |
|---|---|---|---|---|---|---|
| MsrB1 | Met-R-SO repair | Nucleus, Cytosol | 10-20 (model substrates) | Thioredoxin (Trx) | Selenium supplementation, potential allosteric inhibitors (research phase) | Neurodegeneration (Alzheimer's, Parkinson's), Cataracts, Aging |
| Trx/TrxR System | Protein disulfides, H₂O₂, Peroxiredoxins | Ubiquitous (cytosol, mitochondria, nucleus) | TrxR: 100-10,000 (species-dependent) | NADPH | Auranofin, Motexafin Gadolinium | Cancer, Inflammatory diseases |
| GSH/GPx System | H₂O₂, Lipid hydroperoxides | Cytosol, Mitochondria, Extracellular | GPx1: ~800 | Glutathione (GSH) | BSO (GSH depletion), Ebselen (GPx mimetic) | Metabolic syndrome, Liver disease |
| SOD/Catalase | Superoxide (O₂⁻), H₂O₂ | SOD: various compartments; Catalase: Peroxisomes | Catalase: ~10⁵ (high capacity) | None (metal cofactor) | SOD mimetics (e.g., M40403), Catalase inhibitors (3-AT) | ALS, Ischemia-Reperfusion Injury |
| Nrf2-Keap1 | Transcriptional regulator of ~250 antioxidant genes | Cytosol (Keap1), Nucleus (Nrf2) | N/A (signaling pathway) | N/A | Sulforaphane, Bardoxolone methyl, Dimethyl fumarate | Chronic oxidative stress (COPD, CKD), Cancer chemoprevention |
Table 2: Therapeutic Targeting Advantages and Challenges
| Target | Specificity of Action | Druggability (Ligandability) | Potential for Side Effects | Stage of Clinical Development |
|---|---|---|---|---|
| MsrB1 | High (specific stereochemistry, substrate repair) | Medium (active site groove, selenocysteine chemistry) | Low-Medium (specific repair function) | Preclinical target validation |
| TrxR | Medium (broad substrate spectrum) | High (small molecule inhibitors known) | High (essential for many cellular processes) | Clinical (e.g., Auranofin repurposing) |
| GPx | Medium (peroxide reduction) | Medium (mimetics available) | Medium (integral to GSH metabolism) | Clinical (Ebselen in trials) |
| SOD/Catalase | High (specific ROS) | Low (enzyme replacement challenging) | Low (if mimetics are specific) | Preclinical/Experimental (mimetics) |
| Nrf2 | Low (broad genomic regulator) | High (small molecule activators known) | High (pleiotropic effects) | Clinical (multiple agents) |
Key Experimental Protocols for MsrB1 Research
Protocol: Recombinant MsrB1 Activity Assay
Purpose: To measure the in vitro reductase activity of purified MsrB1. Materials:
- Purified recombinant MsrB1 (human, selenocysteine-containing form).
- Substrate: Dabsyl-Met-R-SO (synthetic peptide) or oxidized calmodulin.
- Reductant: Recombinant Thioredoxin (Trx), NADPH, Thioredoxin Reductase (TrxR).
- Reaction Buffer: 50 mM HEPES (pH 7.5), 150 mM NaCl, 1 mM EDTA.
- HPLC system or DTNB (Ellman's reagent) for thiol detection. Method:
- Prepare a regeneration system mix: 100 µM NADPH, 100 nM TrxR, 10 µM Trx in reaction buffer. Incubate at 37°C for 5 min.
- Initiate the reaction by adding MsrB1 (10-100 nM) and substrate (50-200 µM) to the mix.
- Incubate at 37°C for 10-30 minutes.
- For HPLC: Stop reaction with 1% TFA, separate products via reverse-phase HPLC, and quantify reduced methionine peak.
- For DTNB: Stop reaction, add DTNB, and measure absorbance at 412 nm to quantify reduced Trx.
- Calculate activity as nmol Met formed/min/mg enzyme.
Protocol: Cellular MsrB1 Knockdown & Oxidative Stress Phenotyping
Purpose: To assess the functional consequence of MsrB1 loss in a cell model. Materials:
- HEK293 or relevant primary cells.
- siRNA targeting MsrB1 or scrambled control.
- Lipofectamine RNAiMAX.
- Oxidative stressor: tert-Butyl hydroperoxide (tBHP) or H₂O₂.
- CellROX Green/Deep Red Reagent for ROS detection.
- Antibodies for Western Blot: anti-MsrB1, anti-actin, anti-protein-Met-SO (if available). Method:
- Seed cells in 12-well plates. At 60% confluency, transfect with 20 nM siRNA using RNAiMAX per manufacturer's protocol.
- Incubate for 48-72 hours. Harvest one set for Western blot to confirm knockdown.
- Treat siRNA and control cells with a sub-lethal dose of tBHP (e.g., 100 µM for 1 hour).
- Wash cells and load with CellROX reagent (5 µM) for 30 min at 37°C.
- Analyze by flow cytometry or fluorescence microscopy. Compare mean fluorescence intensity (MFI) between MsrB1-KD and control cells under stress.
- Optional: Assess cell viability via MTT assay 24h post-stress.
Visualizations
Diagram 1: MsrB1 Catalytic Cycle & Electron Flow
Diagram 2: MsrB1 in Context of Cellular Antioxidant Network
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for MsrB1 & Antioxidant Pathway Research
| Reagent/Material | Primary Function/Application | Example Product/Source | Key Considerations |
|---|---|---|---|
| Recombinant Human MsrB1 (Sec) | In vitro activity assays, enzyme kinetics, structural studies. | Custom expression in mammalian or E. coli (Cys auxotroph) systems. | Must contain selenocysteine for full activity; verify Sec incorporation via MS. |
| Dabsyl- or Fluorescent-tagged Met-R-SO peptide | Specific spectrophotometric/fluorometric activity substrate. | Custom peptide synthesis services (e.g., GenScript). | Ensure stereochemical purity (R-sulfoxide). Dabsyl allows HPLC/UV detection. |
| Thioredoxin System Regeneration Kit | Provides continuous electron flux to Msr enzymes in coupled assays. | Commercial kits (e.g., Sigma-Aldrich, Cayman Chemical) or purified components (Trx, TrxR, NADPH). | Maintain NADPH excess; control pH (7.0-7.5). |
| siRNA/shRNA targeting MSRB1 | Genetic knockdown for functional cellular studies. | Dharmacon, Sigma-Aldison, or MISSION shRNA libraries. | Use validated sequences; include scrambled control; confirm knockdown via qPCR/WB. |
| Anti-MsrB1 Antibody (Validated) | Detection of endogenous protein levels via Western Blot, IHC, IF. | Commercial antibodies (e.g., Santa Cruz sc-393785, Abcam ab200732). | Check for cross-reactivity with other Msr family members (MsrB2, MsrB3). |
| CellROX Oxidative Stress Reagents | Flow cytometry or microscopy-based detection of general cellular ROS. | Thermo Fisher Scientific (C10444, C10491). | Choose dye based on excitation laser (Green, Orange, Deep Red); include ROS-inducer positive control. |
| Nrf2 Activator/Inhibitor Set | Modulate Nrf2 pathway for comparative studies. | Sulforaphane (activator), ML385 (inhibitor) available from Selleckchem, Tocris. | Timing is critical; activators often cause Nrf2 nuclear translocation within hours. |
| GPx/GSH System Modulators | Manipulate the parallel glutathione pathway. | BSO (Buthionine sulfoximine, GSH depletor), Ebselen (GPx mimetic). | BSO pre-treatment typically requires 12-24 hours for effective GSH depletion. |
Conclusion
The catalytic mechanism of MsrB1, centered on its reactive selenocysteine residue, represents a sophisticated and highly efficient biological solution for repairing oxidative damage to methionine. This detailed exploration, spanning from foundational biochemistry to practical applications and validation, underscores MsrB1's unique position in the cellular redox network. Its superior catalytic efficiency compared to cysteine homologs highlights its non-redundant role in protecting against age-related decline and oxidative stress pathologies. Future research must focus on developing specific small-molecule activators or mimetics of MsrB1, translating this mechanistic knowledge into novel therapeutics for neurodegenerative, cardiovascular, and metabolic diseases. Furthermore, advancing techniques for stable selenoprotein production will be crucial for both fundamental studies and biotechnological exploitation of this potent catalytic scaffold.