Quantifying Cellular Redox: FLIM Biosensing for Thiol-Disulfide Dynamics in Health and Disease
This article provides a comprehensive guide for researchers on applying Fluorescence Lifetime Imaging Microscopy (FLIM) to monitor thiol-disulfide redox states in live cells and tissues.
Quantifying Cellular Redox: FLIM Biosensing for Thiol-Disulfide Dynamics in Health and Disease
Abstract
This article provides a comprehensive guide for researchers on applying Fluorescence Lifetime Imaging Microscopy (FLIM) to monitor thiol-disulfide redox states in live cells and tissues. We first establish the critical role of redox homeostasis in cellular signaling, stress response, and disease pathology. The core methodological section details the design, implementation, and application of genetically encoded and small-molecule FLIM biosensors for glutathione and thioredoxin systems. We then address common experimental challenges, including sensor calibration, environmental artifacts, and data interpretation. Finally, we validate FLIM against established biochemical assays, compare it with intensity-based ratiometric imaging, and highlight its unique advantages for spatiotemporal quantification. This resource empowers scientists to leverage FLIM's precision for advancing redox biology and therapeutic discovery.
The Redox Code: Why Thiol-Disulfide Balance is a Fundamental Cellular Metric
Redox homeostasis is the dynamic equilibrium between the production of reactive oxygen/nitrogen species (ROS/RNS) and their elimination by antioxidant defenses. This balance is critical for cellular signaling, metabolism, and survival. Key to this regulation are redox-sensitive cysteine thiols (-SH), which act as molecular switches. Their reversible oxidation to disulfides (-S-S-) or other oxidative post-translational modifications (PTMs) transduces redox signals, modulating protein function, pathway activity, and gene expression.
Fluorescence Lifetime Imaging Microscopy (FLIM) is a powerful tool for quantifying thiol-disulfide redox states in live cells. Unlike intensity-based methods, FLIM measures the nanosecond decay time of fluorescence, a parameter independent of probe concentration, excitation intensity, or photobleaching, providing robust, quantitative biosensing of the cellular redox environment.
Quantitative Landscape of Thiol Redox Potential
The following table summarizes key quantitative parameters and measurements in cellular thiol redox research, highlighting targets for FLIM-based sensing.
Table 1: Quantitative Metrics in Thiol-Disulfide Redox Homeostasis
| Parameter | Typical Physiological Range (Compartment) | Measurement Method | Significance |
|---|---|---|---|
| Glutathione Redox Potential (Eh) | -260 to -200 mV (Cytosol) | HPLC, Redox-sensitive GFP (roGFP) | Master indicator of cellular redox buffer capacity. |
| GSSG/GSH Ratio | 1:100 to 1:10 (Cytosol) | Enzymatic recycling assay, MS | Increased ratio indicates oxidative stress. |
| Cysteine Sulfenic Acid (-SOH) | Low nM, transient | Dimedone-based probes, MS | Initial, reversible oxidative PTM; signaling intermediate. |
| Protein S-glutathionylation | Variable, condition-dependent | Biotin-switch assays, MS | Protective/regulatory PTM; mixed disulfide with GSH. |
| H2O2 Concentration | 1-100 nM (steady-state) | Genetically encoded sensors (e.g., HyPer) | Primary signaling ROS molecule. |
| FLIM Donor Lifetime | 1.0 - 4.0 ns (for common dyes) | Time-correlated single photon counting (TCSPC) | Direct readout of molecular environment, FRET efficiency, or quenching. |
Core Experimental Protocols
Protocol 1: FLIM-Based Measurement of Glutathione Redox Potential Using roGFP2-Orp1
This protocol uses a FRET-based sensor where the redox-sensitive roGFP2 is fused to the H2O2-sensing protein Orp1. Oxidation-induced disulfide formation alters FRET efficiency, detected via FLIM.
I. Materials & Reagents
- Plasmid: pCMV-roGFP2-Orp1.
- Cell Culture: Appropriate media, transfection reagent (e.g., Lipofectamine 3000).
- Treatment Solutions: 100 µM - 1 mM H2O2 in PBS (freshly diluted); 5 mM Dithiothreitol (DTT) in PBS.
- Imaging Buffer: Hanks' Balanced Salt Solution (HBSS), pH 7.4.
- Equipment: Confocal or multiphoton microscope equipped with TCSPC FLIM module, 405 nm or 920 nm (2P) pulsed laser.
II. Procedure
- Cell Preparation: Seed cells onto 35 mm glass-bottom dishes. At 60-70% confluence, transfect with the roGFP2-Orp1 plasmid per manufacturer's protocol. Incubate for 24-48h.
- FLIM Acquisition:
- Mount dish on microscope stage maintained at 37°C/5% CO2.
- Excite roGFP2 using a 405 nm pulsed laser (or 920 nm for 2P).
- Collect emission through a 525/50 nm bandpass filter.
- Acquire FLIM images until >1000 photons per pixel are collected in the peak channel for sufficient decay curve fitting.
- Experimental Treatments:
- Acquire a baseline FLIM image (Control).
- Gently add H2O2 (e.g., 200 µM final) to the dish. Incubate for 5-10 min, then acquire a second FLIM image (Oxidized).
- Wash cells with HBSS. Add DTT (5 mM final). Incubate for 10 min, acquire a third FLIM image (Reduced).
- Data Analysis:
- Fit fluorescence decay curves per pixel to a double-exponential model using software (e.g., SPCImage, FLIMfit).
- Calculate the amplitude-weighted average lifetime (τavg).
- Generate pseudocolor lifetime maps. A decrease in τavg indicates increased FRET due to sensor oxidation (disulfide formation).
- Quantify mean lifetime per cell/region across conditions.
Protocol 2: Assessing Specific Protein S-Glutathionylation via Biotin-Switch Assay
This biochemical protocol identifies proteins that form mixed disulfides with glutathione (GSH).
I. Materials & Reagents
- Lysis Buffer: HEN buffer (250 mM HEPES pH 7.7, 1 mM EDTA, 0.1 mM neocuproine) + 1% Triton X-100, supplemented with 50 mM N-ethylmaleimide (NEM) and protease inhibitors.
- Blocking Reagent: NEM (from stock).
- Reducing Agent: Sodium ascorbate (fresh 100 mM stock).
- Biotinylation Reagent: EZ-Link HPDP-Biotin (1 mg/mL in DMSO).
- NeutrAvidin Beads: For pull-down.
- Detection: Streptavidin-HRP (western) or mass spectrometry buffers.
II. Procedure
- Cell Lysis & Free Thiol Blocking: Lyse treated cells in NEM-containing lysis buffer. Incubate 30 min at 50°C. This alkylates all free thiols.
- Protein Cleanup: Remove excess NEM by acetone precipitation or desalting column.
- Reduction of S-Glutathionylation: Resuspend protein pellet in HEN + 1% SDS. Reduce glutathionylated disulfides by adding sodium ascorbate to 25 mM final. Incubate 1h at room temperature.
- Biotin Labeling: Add HPDP-Biotin to label the newly reduced thiols. Incubate 1h at RT.
- Pull-down & Analysis: Remove excess biotin. Incubate samples with NeutrAvidin beads overnight at 4°C. Wash beads stringently. Elute proteins with Laemmli buffer containing β-mercaptoethanol for western blot analysis, or process for tryptic digest and LC-MS/MS.
Visualizing Pathways and Workflows
Diagram 1: Cysteine Thiols as Molecular Switches in Signaling
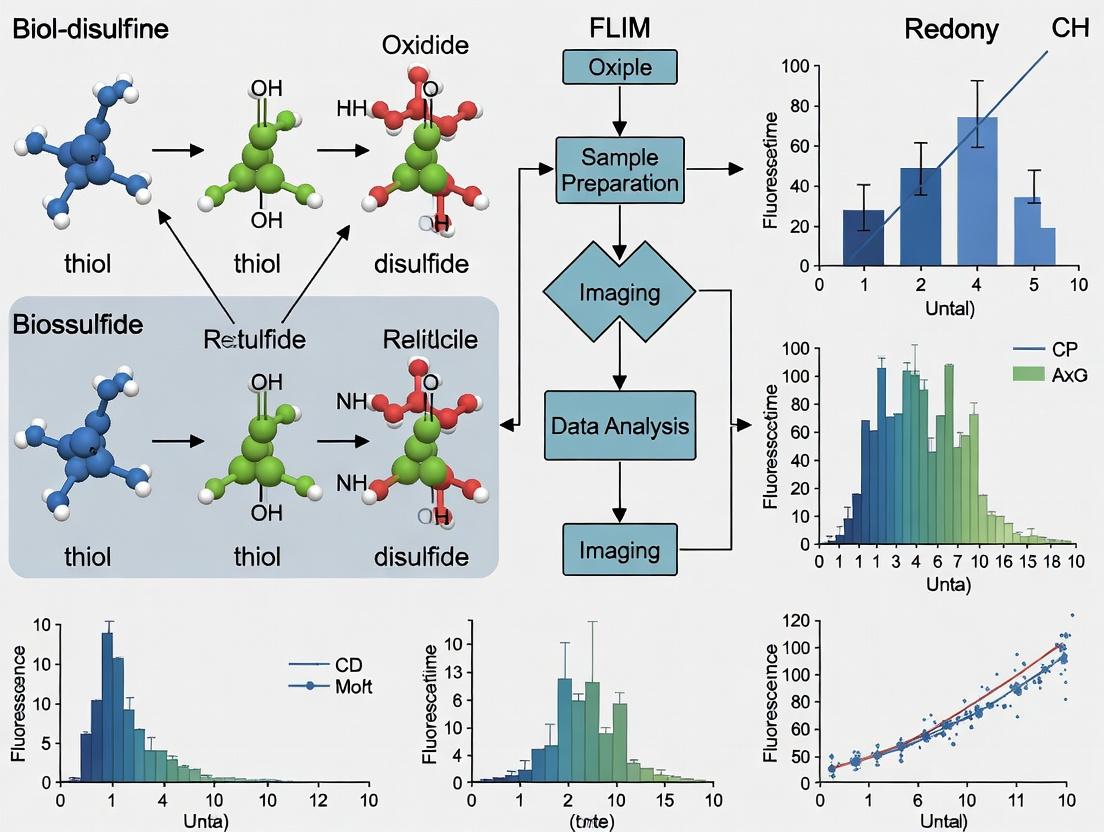
Diagram 2: FLIM Workflow for Redox Biosensing
The Scientist's Toolkit: Key Reagent Solutions
Table 2: Essential Research Reagents for Thiol-Disulfide Redox Studies
| Reagent / Tool | Category | Primary Function |
|---|---|---|
| roGFP2 (redox-sensitive GFP) | Genetically Encoded Biosensor | Conformation changes upon thiol oxidation/reduction, altering fluorescence excitation peaks. Ratio-metric readout. |
| roGFP2-Orp1 / Grx1-roGFP2 | Genetically Encoded Biosensor (FLIM optimized) | FRET-based sensors for specific detection of H22 (Orp1) or glutathione redox potential (Grx1). Ideal for FLIM. |
| N-Ethylmaleimide (NEM) | Alkylating Agent | Irreversibly blocks free thiol (-SH) groups, "snapshot" of redox state at moment of lysis. Critical for biotin-switch assays. |
| Iodoacetamide (IAM) | Alkylating Agent | Alternative to NEM for blocking free thiols; used in proteomic sample prep to prevent scrambling. |
| Diamide | Thiol-Oxidizing Agent | Chemically oxidizes glutathione and protein thiols, inducing disulfide stress in a controlled manner. |
| Dithiothreitol (DTT) / Tris(2-carboxyethyl)phosphine (TCEP) | Reducing Agents | Chemically reduce disulfide bonds (-S-S-) back to free thiols (-SH). TCEP is more stable and acts at a wider pH range. |
| Sodium Ascorbate | Selective Reducing Agent | Specifically reduces S-nitrosothiols and some mixed disulfides (like -SSG) but not all disulfides, used in selective labeling. |
| EZ-Link HPDP-Biotin | Biotinylation Reagent | Thiol-reactive, used to label newly reduced cysteine thiols in biotin-switch assays for pull-down and detection. |
| Recombinant Thioredoxin (Trx) / Glutaredoxin (Grx) Systems | Enzymatic Reductants | Specific physiological systems for reducing protein disulfides and deglutathionylating proteins, respectively. |
| Monobromobimane (mBBr) | Thiol-Specific Fluorescent Probe | Cell-permeable dye that forms adducts with thiols for detection via fluorescence (not ratio-metric). |
The glutathione (GSH/GSSG) and thioredoxin (Trx) systems are the principal cellular thiol-disulfide redox buffers, maintaining a reducing intracellular environment. Dysregulation of these systems is a hallmark of oxidative stress, linked to aging, neurodegenerative diseases, cancer, and metabolic disorders. Fluorescence Lifetime Imaging Microscopy (FLIM) is a powerful tool for studying these redox couples in vivo and in real-time. Unlike intensity-based methods, FLIM measures the exponential decay rate of fluorescence, a parameter that is inherently concentration-independent and sensitive to the molecular microenvironment. This makes it ideal for biosensing using redox-sensitive fluorescent proteins (roGFPs, rxYFPs) or small-molecule dyes, enabling precise, compartment-specific quantification of redox potentials (Eh).
Core System Biochemistry & Quantitative Landscape
The Glutathione System
Glutathione exists as reduced monomeric glutathione (GSH) and oxidized glutathione disulfide (GSSG). The balance is maintained by NADPH-dependent glutathione reductase (GR). The redox potential (Eh) is calculated via the Nernst equation.
Key Reaction: 2GSH ⇌ GSSG + 2H+ + 2e-
The Thioredoxin System
Thioredoxin (Trx) is a small redox-active protein with a dithiol/disulfide active site. It is reduced by thioredoxin reductase (TrxR) using NADPH. The Trx system regulates target proteins via post-translational thiol-disulfide exchange.
Key Reaction: Trx-(SH)2 + Protein-S2 ⇌ Trx-S2 + Protein-(SH)2
Table 1: Key Quantitative Parameters of Major Cellular Redox Buffers
| Parameter | Glutathione System | Thioredoxin System (Trx1) |
|---|---|---|
| Total Pool Size | 1-10 mM (cytosol) | ~10 µM (cytosol) |
| Redox Potential (Eh) | -260 to -200 mV (cytosol) | ~-280 mV (cytosol) |
| [Reduced]/[Oxidized] Ratio | 30:1 to 100:1 (healthy cells) | >100:1 (reduced/oxidized) |
| Primary Reductase | Glutathione Reductase (GR) | Thioredoxin Reductase (TrxR) |
| Electron Donor | NADPH | NADPH |
| Primary Compartments | Cytosol, Mitochondria, Nucleus, ER | Cytosol, Nucleus, Mitochondria (Trx2) |
Table 2: Common FLIM-Compatible Redox Biosensors
| Biosensor Name | Redox Target | Excitation (nm) | Emission (nm) | Redox-Sensitive FLIM Readout |
|---|---|---|---|---|
| roGFP2 | GSH/GSSG (via Grx1) | 400 / 490 | 510 | Lifetime change with oxidation state |
| rxYFP | Trx Family | 514 | 527 | Lifetime sensitive to dithiol-disulfide status |
| Grx1-roGFP2 | GSH/GSSG (direct) | 400 / 490 | 510 | Highly specific for glutathione potential |
| Mrx1-roGFP | Mycothiol (bacteria) | 400 / 490 | 510 | Prokaryotic redox sensing |
Detailed Protocols for FLIM-Based Redox Sensing
Protocol 3.1: Live-Cell Imaging of GSH/GSSG Redox Potential using Grx1-roGFP2 FLIM
Objective: To measure compartment-specific glutathione redox potential (Eh) in living cells.
Materials:
- Cell line expressing targeted Grx1-roGFP2 (e.g., cytosol, mitochondria).
- Confocal/FLIM microscope system with time-correlated single photon counting (TCSPC).
- Imaging medium (e.g., phenol red-free, with 25mM HEPES).
- Redox modulators: 2mM DTT (reducing control), 1mM Diamide (oxidizing control).
- Calibration buffer set (pH 7.2) with 10mM GSH/GSSG mixtures for in situ calibration.
Procedure:
- Cell Preparation: Seed cells on imaging dishes 24-48h prior. Transfect or infect with Grx1-roGFP2 construct 24h before imaging.
- Microscope Setup: Use a 405nm or 440nm pulsed laser for excitation. Collect emission at 500-520nm using a bandpass filter. Set TCSPC acquisition for sufficient photons (>1000 at peak) for lifetime fitting.
- In-situ Calibration (Crucial for Eh calculation): a. Permeabilize cells with 0.05% digitonin in calibration buffers. b. Incubate for 5 min in each of 5 buffers containing fixed total GSH+GSSG (e.g., 10mM) but varying GSH/GSSG ratios (e.g., 100:1, 10:1, 1:1, 1:10, 1:100). c. Acquire FLIM data at each ratio.
- Experimental Imaging: Acquire baseline FLIM images. Add experimental compounds (e.g., H2O2, drugs) and perform time-lapse FLIM.
- Data Analysis:
a. Fit fluorescence decay curves per pixel to a double-exponential model:
I(t) = α1 exp(-t/τ1) + α2 exp(-t/τ2) + C. b. Calculate the amplitude-weighted mean lifetime: τm = (α1τ1 + α2τ2) / (α1 + α2). c. Plot τm from calibration steps against known Eh (calculated via Nernst equation) to generate a standard curve. d. Convert experimental τm values to Eh (mV) using this curve.
Protocol 3.2: Assessing Thioredoxin Redox Status with rxYFP FLIM
Objective: To monitor the oxidation state of the cytosolic thioredoxin system.
Materials:
- Cells expressing rxYFP (targeted as needed).
- FLIM microscope with 514nm excitation capability.
- Modulators: 10µM Auranofin (TrxR inhibitor), 1mM H2O2.
- Quencher: 1mM KI for assessing accessibility (collisional quenching).
Procedure:
- Express rxYFP in the cellular compartment of interest.
- Acquire FLIM images using 514nm pulsed laser excitation, collect emission at 525-550nm.
- Treat cells with inhibitors/oxidants and perform time-lapse FLIM.
- For data validation, perform a KI quenching experiment. A longer lifetime that is sensitive to KI indicates a more reduced, solvent-accessible state of the rxYFP dithiol.
- Analyze mean fluorescence lifetime (τm) maps. A shift to a shorter τm typically indicates oxidation of the sensor.
The Scientist's Toolkit: Essential Reagents & Materials
Table 3: Key Research Reagent Solutions for Thiol-Disulfide Redox Research
| Reagent / Material | Function & Application |
|---|---|
| roGFP2 (or Grx1-roGFP2) Plasmids | Genetically encoded biosensor for GSH/GSSG redox potential. |
| rxYFP Plasmids | Genetically encoded biosensor for thioredoxin-family redox states. |
| Auranofin | Potent and specific inhibitor of Thioredoxin Reductase (TrxR). |
| BSO (Buthionine sulfoximine) | Inhibitor of glutathione synthesis (GCL). Depletes cellular GSH. |
| Diamide | Thiol-specific oxidant; rapidly converts GSH to GSSG. |
| DTT (Dithiothreitol) | Strong reducing agent; used as a positive control for reduction. |
| Monochlorobimane | Cell-permeable dye forming fluorescent adduct with GSH; used for total GSH estimation. |
| NADPH/NADP+ Assay Kits | Quantify the NADPH pool, the electron source for both GSH and Trx systems. |
| TCSPC FLIM Module | Time-Correlated Single Photon Counting hardware/software for precise lifetime measurement. |
| Mathematical Fitting Software | e.g., SPCImage, FLIMfit, for analyzing lifetime decay curves. |
Visualization Diagrams
Fig 1: Core Thiol Redox Systems & NADPH Link
Fig 2: FLIM Workflow for Redox Biosensing
Fig 3: roGFP Redox Sensing Mechanism
This application note explores the central role of thiol-disulfide redox dysregulation across pathologies, emphasizing the use of Fluorescence Lifetime Imaging (FLIM) biosensing for quantitative, spatiotemporally resolved measurements. The content supports a thesis focused on advancing FLIM-based biosensors for dynamic redox research in live cells and tissues.
The dynamic equilibrium between reduced (thiol, -SH) and oxidized (disulfide, -S-S-) groups in proteins and low-molecular-weight compounds (e.g., glutathione, GSH/GSSG) is a fundamental cellular regulatory mechanism. Dysregulation of this redox balance is a common feature in diverse diseases, driving pathological signaling, metabolic reprogramming, and cell death.
Quantitative Data on Redox Dysregulation in Disease
Table 1: Characteristic Redox Perturbations in Major Disease Classes
| Disease Category | Key Redox Metric | Typical Change vs. Healthy State | Primary Compartment Affected | Associated Consequences |
|---|---|---|---|---|
| Cancer (Solid Tumors) | Glutathione (GSH) | ↑ 2- to 10-fold | Cytosol, Mitochondria | Chemoresistance, Proliferation, Metastasis |
| Cysteine (Cys/CySS) Pool (Eh) | More oxidized (~+40 to +60 mV) | Extracellular | Pro-survival signaling | |
| ROS (H₂O₂) | Chronic, moderate ↑ | Perinuclear, Focal | Pro-tumorigenic signaling (e.g., HIF-1α, NF-κB) | |
| Neurodegeneration (e.g., AD, PD) | Glutathione (GSH) | ↓ 30-70% | Mitochondria, Neuronal Cytosol | Oxidative damage, Protein aggregation |
| Protein-S-glutathionylation | ↑ | Mitochondria, Synapses | Synaptic dysfunction, Metabolic failure | |
| Lipid Peroxidation (4-HNE) | ↑ 2- to 5-fold | Neuronal Membranes | Ferroptosis, Loss of membrane integrity | |
| Cardiovascular (e.g., HF, Atherosclerosis) | NADPH/NADP+ Ratio | ↓ | Cytosol | Reduced antioxidant capacity (GR, Trx) |
| Thioredoxin-1 (Trx1) Redox State | Oxidized | Cytosol/Nucleus | Apoptosis, Hypertrophy | |
| Metabolic (e.g., T2D) | Mitochondrial H₂O₂ | ↑ | Mitochondrial Matrix | Insulin resistance, β-cell dysfunction |
Table 2: FLIM-Based Redox Biosensors: Key Performance Metrics
| Biosensor Name | Redox Pair Sensed | Dynamic Range (Lifetime Change) | Response Time | Primary Cellular Target |
|---|---|---|---|---|
| roGFP (e.g., roGFP2-Orp1) | GSH/GSSG, H₂O₂ | ~0.8-1.2 ns (Ratiometric) | Seconds to minutes | Cytosol, Mitochondria, ER |
| HyPer | H₂O₂ | ~0.5-0.7 ns | Seconds | Cytosol, Nucleus |
| GRX1-roGFP2 | Glutathione Redox Potential (Eh) | ~1.5 ns | Minutes | Sub-compartments (e.g., Mito matrix) |
| Custom FLIM-FRET Sensors | Protein-specific thiol oxidation | ~0.3-2.0 ns (Donor lifetime) | Seconds | Defined protein targets |
Experimental Protocols
Protocol 1: FLIM Measurement of Glutathione Redox Potential in Live Cells Using roGFP2-Grx1
Objective: Quantify the glutathione redox potential (EGSSG/2GSH) in the mitochondrial matrix of live cancer cells. Principle: roGFP2 is a redox-sensitive GFP. Grx1 specifically equilibrates it with the GSH/GSSG pool. Oxidation/reduction alters its excitation spectrum, detectable via FLIM as a change in fluorescence lifetime, which is rationetric and insensitive to probe concentration.
Materials:
- Cells stably expressing mito-roGFP2-Grx1.
- Confocal/FLIM microscope with time-correlated single photon counting (TCSPC) capability.
- 405 nm and 488 nm pulsed laser lines.
- Imaging chamber with environmental control (37°C, 5% CO₂).
- Redox Modulators:
- 1 mM DTT (strong reductant, negative control).
- 200 µM Diamide (thiol oxidant, positive control).
- 10 mM N-Ethylmaleimide (NEM, thiol alkylator, for fixation control).
Procedure:
- Cell Preparation: Seed cells on glass-bottom dishes. Transfect/infect with mito-roGFP2-Grx1 construct 24-48h prior.
- Microscope Setup:
- Set TCSPC parameters: laser repetition rate ≤ 40 MHz, acquisition time to achieve ~10⁶ photons in the brightest pixel.
- Configure two excitation channels: 405 nm and 488 nm.
- Use a 500-550 nm bandpass emission filter.
- FLIM Acquisition:
- Acquire a baseline FLIM image using both 405 nm and 488 nm excitation.
- Perfuse cells with culture medium containing 200 µM Diamide. Acquire sequential FLIM images every 2 minutes for 20 minutes.
- Wash with fresh medium. Perfuse with 1 mM DTT. Acquire images every 2 minutes for 15 minutes.
- Data Analysis:
- Fit fluorescence decay curves per pixel to a double-exponential model: I(t) = α₁exp(-t/τ₁) + α₂exp(-t/τ₂) + C.
- Calculate the amplitude-weighted mean fluorescence lifetime: τm = (α₁τ₁ + α₂τ₂) / (α₁ + α₂).
- Generate lifetime maps for each excitation wavelength.
- Compute the redox ratio: R = τm(405 nm) / τm(488 nm). Calibrate R to Eh using DTT (fully reduced) and Diamide (fully oxidized) values.
- Quantification: Report Eh values in mV for regions of interest (e.g., mitochondrial clusters).
Protocol 2: Assessing Protein-Specific Thiol Oxidation via FLIM-FRET
Objective: Measure the oxidation status of a specific protein thiol (e.g., in PTEN or Parkin) in response to oxidative stress. Principle: A fusion protein is created with the protein of interest sandwiched between a donor fluorophore (e.g., mCerulean3) and an acceptor (e.g., cpVenus). Conformational change upon thiol oxidation alters FRET efficiency, detected as a change in donor fluorescence lifetime via FLIM.
Materials:
- Cells expressing the FLIM-FRET biosensor for the target protein (e.g., Cyto-/Mito-Rex).
- FLIM microscope (as in Protocol 1).
- 440 nm pulsed laser for CFP excitation.
- H₂O₂ (100 µM - 1 mM) or disease-relevant stressors (e.g., rotenone for PD models).
Procedure:
- Express the biosensor in relevant cell models (e.g., neuronal lines for neurodegeneration).
- Acquire baseline donor FLIM image (440 nm excitation, 470-500 nm emission).
- Treat cells with stressor (e.g., 500 µM H₂O₂) and acquire time-lapse FLIM images.
- Analyze donor lifetime (τDA) per pixel. A decrease in τDA indicates increased FRET (reduced state of thiol), while an increase indicates decreased FRET (oxidized state).
- Normalize data: % Oxidation = [(τDA, treated - τDA, reduced) / (τDA, oxidized - τDA, reduced)] * 100, where controls are defined using DTT and Diamide.
Pathways and Workflows
Diagram Title: Redox Dysregulation Drives Divergent Disease Pathologies.
Diagram Title: FLIM Protocol for Redox Potential Quantification.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for FLIM Redox Biosensing Research
| Reagent / Material | Function in Redox Research | Example Product / Note |
|---|---|---|
| Genetically Encoded Biosensors (e.g., roGFP, HyPer, Rex) | Target-specific, real-time reporting of redox states in live cells. | Addgene plasmids; AAV for in vivo use. |
| TCSPC FLIM Module | Enables precise measurement of nanosecond fluorescence lifetime decays. | Becker & Hickl SPC-150; PicoQuant PicoHarp. |
| Pulsed Laser Diodes (405, 440, 488 nm) | Provide time-correlated excitation for lifetime measurement. | LDH-D-C-405; PicoQuant lasers. |
| Environmental Chamber | Maintains live cells at 37°C, 5% CO₂ during imaging. | Okolab stage-top incubator. |
| Redox Modulators (DTT, Diamide) | Used for in situ calibration of biosensor response. | High-purity, cell culture grade. Prepare fresh. |
| N-Ethylmaleimide (NEM) | Alkylates free thiols to "snapshot" and fix redox state pre-fixation. | Use at 10-20 mM for 5 min. |
| Mito/ER-Targeted Redox Dyes (e.g., MitoTracker Red CM-H₂XRos) | Complementary tools to validate biosensor data and localize organelles. | Use at low nM concentrations. |
| H₂O₂-Sensitive Probes (e.g., Peroxy Orange 1) | Chemically specific probes to cross-validate ROS levels. | Small-molecule, rationetric alternatives. |
| Image Analysis Software | For FLIM decay fitting, rationetric calculation, and lifetime map generation. | FLIMfit (open-source), SPCImage, SymPhoTime. |
The Need for Dynamic, Non-Invasive Measurement in Live Cells
Within the broader thesis on applying Fluorescence Lifetime Imaging (FLIM) biosensing to thiol-disulfide redox research, this application note addresses the critical need for dynamic, non-invasive measurement in live cells. Static, endpoint assays and invasive lysis methods disrupt the delicate redox equilibrium, failing to capture the spatiotemporal dynamics of redox regulation. This note details the application of genetically encoded FLIM-based biosensors to monitor redox parameters in real-time, providing protocols and data for studying oxidative stress, drug mechanisms, and metabolic signaling in their native cellular context.
Table 1: Comparison of Redox Measurement Methodologies
| Method | Spatial Resolution | Temporal Resolution | Invasiveness | Key Measurand | Compatibility with Live Cells |
|---|---|---|---|---|---|
| Mass Spectrometry (Lysates) | N/A (Population Average) | Low (Endpoint) | High (Cell Lysis) | Global thiol/disulfide status | No |
| Ellman's Assay | N/A (Population Average) | Low (Endpoint) | High (Cell Lysis) | Total glutathione | No |
| Chemical Dyes (e.g., roGFP) | Organelle/Cytosol | Medium (Seconds-Minutes) | Moderate (Dye Loading) | Relative redox potential (EGSH) | Yes |
| FLIM Biosensors (e.g., Grx1-roGFP2) | Subcellular (Organelle) | High (Milliseconds-Seconds) | Low (Genetically Encoded) | Absolute EGSH via τ | Yes |
Table 2: Example FLIM Data for Redox Biosensor Response
| Condition / Treatment | Average FLIM Lifetime (τ) [ns] | Interpreted EGSH [mV] | Cellular Compartment | Key Observation |
|---|---|---|---|---|
| Untreated Control | 2.85 ± 0.05 | -315 ± 5 | Cytosol | Baseline reduced state |
| H2O2 (200 µM, 5 min) | 2.35 ± 0.07 | -265 ± 8 | Cytosol | Significant oxidation |
| DTT (10 mM, 10 min) | 2.95 ± 0.04 | -330 ± 4 | Cytosol | Forced reduction |
| NAC Pre-treatment | 2.82 ± 0.06 | -312 ± 6 | Cytosol | Protection from oxidation |
Detailed Experimental Protocols
Protocol 1: Live-Cell FLIM for Redox Sensing with Grx1-roGFP2
- Objective: To quantify the glutathione redox potential (EGSH) in the cytosol of live cells under oxidative challenge.
- Materials: See "The Scientist's Toolkit" below.
- Procedure:
- Cell Preparation: Seed cells in a 35mm glass-bottom dish. Transfect with a plasmid encoding the Grx1-roGFP2 biosensor targeted to the cytosol. Incubate for 24-48h.
- Microscope Setup: Use a confocal microscope equipped with a TCSPC FLIM module. Set excitation to 920nm (2-photon) or 488nm laser line. Collect emission using a 525/50 nm bandpass filter.
- Calibration (In-situ): For absolute EGSH calculation, perform a two-point calibration on each field of view:
- Acquire a baseline FLIM image.
- Perfuse with Calibration Buffer A (10mM DTT) for 15 min to fully reduce the biosensor. Acquire FLIM image.
- Wash and perfuse with Calibration Buffer B (100µM Diamide) for 15 min to fully oxidize the biosensor. Acquire FLIM image.
- Experimental Imaging: In fresh, untreated cells, acquire a time-series of FLIM images (e.g., 1 image/minute). After establishing a baseline, add the experimental stimulus (e.g., 200µM H2O2) directly to the media. Continue acquisition for the desired duration.
- Data Analysis: Fit the fluorescence decay curve of each pixel using a bi-exponential model. Calculate the amplitude-weighted mean lifetime (τm). Use the calibration lifetimes to convert τm to the degree of oxidation (OxD), and subsequently to EGSH using the Nernst equation.
Protocol 2: Pharmacological Modulation & FLIM Assessment
- Objective: To test the protective effect of antioxidant pre-treatments on EGSH.
- Procedure:
- Prepare two sets of cells expressing the biosensor as in Protocol 1.
- Pre-treat one set with 5mM N-Acetylcysteine (NAC) for 2 hours. The other set serves as an untreated control.
- Mount both dishes on the FLIM microscope and acquire baseline images.
- Challenge both sets with 500µM tert-Butyl hydroperoxide (tBHP).
- Acquire FLIM images every 30 seconds for 20 minutes.
- Analyze the rate and magnitude of EGSH oxidation, comparing the NAC-treated vs. control populations.
Visualizations
Diagram 1: FLIM Redox Biosensing Mechanism (86 chars)
Diagram 2: FLIM Redox Experiment Workflow (62 chars)
The Scientist's Toolkit: Key Research Reagent Solutions
- Genetically Encoded Redox Biosensor (e.g., Grx1-roGFP2): A fusion protein comprising human glutaredoxin-1 and redox-sensitive GFP. Serves as the specific, reversible molecular probe for EGSH.
- TCSPC FLIM Module: Time-Correlated Single Photon Counting hardware and software. Essential for precisely measuring the nanosecond fluorescence lifetime decay, which is independent of biosensor concentration and excitation intensity.
- Two-Photon or Confocal Laser Microscope: Provides the optical sectioning and precise excitation needed for high-resolution, live-cell imaging without excessive photobleaching.
- In-situ Calibration Reagents:
- DTT (Dithiothreitol, 10mM): Strong reducing agent to define the fully reduced lifetime (τred) of the biosensor.
- Diamide (100-500µM): Thiol-specific oxidant to define the fully oxidized lifetime (τox).
- Oxidative Stressors:
- tBHP (tert-Butyl hydroperoxide): Organic peroxide; a more stable and membrane-permeable oxidant than H2O2.
- Menadione: Redox-cycling quinone that generates superoxide, testing specific pathway responses.
- Antioxidant Compounds (for Modulation):
- N-Acetylcysteine (NAC, 1-5mM): Precursor for glutathione synthesis, used to bolster cellular antioxidant capacity.
- Mito-TEMPO: Mitochondria-targeted superoxide scavenger for compartment-specific studies.
Within the broader thesis on FLIM biosensing for thiol-disulfide redox research, this application note establishes why Fluorescence Lifetime Imaging Microscopy (FLIM) is the superior modality for quantifying cellular redox state. Unlike intensity-based fluorescence measurements, fluorescence lifetime (τ) is an intrinsic property of a fluorophore that is independent of probe concentration, excitation light intensity, and photobleaching. This makes it uniquely robust for sensing the molecular microenvironment.
The principle central to redox sensing is that the fluorescence lifetime of certain genetically encoded or chemical probes is exquisitely sensitive to the presence of specific reactive species or to changes in local physicochemical conditions (e.g., pH, viscosity) that correlate with redox metabolism. For thiol-disulfide redox research, probes sensitive to glutathione (GSH)/glutathione disulfide (GSSG) balance, NADH/NAD⁺ ratio, or reactive oxygen species (ROS) like H₂O₂ are key. FLIM detects the shift in lifetime that occurs when the probe is oxidized or reduced, or when it binds to a target molecule, providing a quantitative map of redox potential with subcellular resolution.
Key Advantages of FLIM for Redox Sensing
The following table summarizes the critical advantages of FLIM over intensity-based methods for redox biology applications.
Table 1: Advantages of FLIM vs. Intensity-Based Imaging for Redox Sensing
| Parameter | Intensity-Based Fluorescence | FLIM-Based Sensing | Implication for Redox Research |
|---|---|---|---|
| Quantitative Accuracy | Highly susceptible to variations in probe concentration, expression level, and tissue turbidity. | Independent of fluorophore concentration; reports on molecular environment. | Enables accurate comparison between different cells, treatments, and time points. |
| Photobleaching Resistance | Signal loss directly compromises quantification. | Lifetime is largely unaffected by moderate photobleaching. | Allows for longer-term live-cell imaging of dynamic redox processes. |
| Artifact Rejection | Difficult to distinguish true signal from autofluorescence or background. | Can separate probe fluorescence from background based on lifetime signature. | Improves specificity in tissues with high autofluorescence. |
| Multiplexing Capacity | Limited by spectral overlap of emission spectra. | Enables multiplexing of probes with similar emission spectra but different lifetimes. | Allows simultaneous monitoring of multiple redox couples (e.g., GSH/GSSG and NADH). |
| Microenvironment Sensing | Indirect, often requires ratiometric probes. | Directly sensitive to quenching, FRET, and molecular interactions. | Directly reports on molecular binding/oxidation events related to redox state. |
Experimental Protocols
Protocol 1: FLIM of roGFP for Glutathione Redox Potential (EGSSG/2GSH)
Objective: To quantify the thiol-disulfide redox state in live cells using the redox-sensitive green fluorescent protein (roGFP) coupled to Grx1 (glutaredoxin-1).
Principle: roGFP has two cysteine residues that form a disulfide bond upon oxidation, altering the chromophore's excitation spectrum. Rationetric intensity measurements are common, but FLIM of roGFP provides a concentration-independent alternative. Oxidation typically results in a measurable decrease in fluorescence lifetime.
Materials:
- Cells expressing roGFP1-Grx1 (e.g., HeLa, HEK293).
- Confocal or multiphoton microscope with time-correlated single photon counting (TCSPC) FLIM module.
- Pulsed laser: 485 nm or two-photon excitation at 950 nm.
- Imaging chamber with temperature and CO₂ control.
- Control Reagents: 10 mM Dithiothreitol (DTT, reducing agent), 500 µM Diamide (oxidizing agent).
Procedure:
- Cell Preparation: Plate cells expressing roGFP1-Grx1 on glass-bottom dishes 24-48 hours before imaging.
- System Calibration: Perform daily calibration of the FLIM system using a reference fluorophore with a known, stable lifetime (e.g., fluorescein at pH 10, τ ~4.0 ns).
- Image Acquisition:
- Maintain cells in imaging medium at 37°C, 5% CO₂.
- Set excitation to 485 nm (pulsed) and collect emission through a 500-550 nm bandpass filter.
- Acquire FLIM data until sufficient photons are collected for robust fitting (>1000 photons at the peak pixel is a typical minimum).
- For validation, acquire images before and after treatment with 10 mM DTT (full reduction, 5 min) and 500 µM Diamide (full oxidation, 5 min).
- Data Analysis:
- Fit fluorescence decay curves per pixel using a bi-exponential model:
I(t) = α₁ exp(-t/τ₁) + α₂ exp(-t/τ₂) + C. - Calculate the amplitude-weighted mean lifetime:
τ<sub>m</sub> = (α₁τ₁ + α₂τ₂) / (α₁ + α₂). - Generate pseudocolor lifetime maps (τm). A shift to shorter lifetime indicates oxidation.
- The redox state can be expressed as the fraction of oxidized probe, which correlates linearly with τm.
- Fit fluorescence decay curves per pixel using a bi-exponential model:
Protocol 2: FLIM of Endogenous NAD(P)H for Metabolic Redox Imaging
Objective: To assess the metabolic redox state via the fluorescence lifetime of endogenous coenzyme NAD(P)H.
Principle: NADH and NADPH are fluorescent, while their oxidized forms (NAD⁺, NADP⁺) are not. The fluorescence lifetime of NAD(P)H is sensitive to its protein-binding status: free NAD(P)H has a short lifetime (~0.4 ns), while enzyme-bound NAD(P)H has a long lifetime (~2-4 ns). The ratio of bound-to-free (α₂%/α₁%) or the mean lifetime serves as a sensitive indicator of the cellular metabolic poise (e.g., glycolysis vs. oxidative phosphorylation).
Materials:
- Untreated live cells (no transfection required).
- Multiphoton microscope with TCSPC FLIM module is ideal to minimize cellular damage and improve optical sectioning.
- Pulsed Ti:Sapphire laser tuned to 740-750 nm for two-photon excitation of NAD(P)H.
- Emission filter: 400-480 nm (bandpass for NAD(P)H) or 460/80 nm.
- Control Reagents: 10 µM Rotenone/Antimycin A (inhibits oxidative phosphorylation), 50 mM 2-Deoxy-D-glucose (2-DG, inhibits glycolysis).
Procedure:
- Sample Preparation: Plate cells on glass-bottom dishes in standard growth medium. For imaging, replace with a phenol-red free, buffered imaging medium.
- FLIM Acquisition:
- Use two-photon excitation at 740 nm. Keep laser power as low as possible to avoid photodamage and metabolic perturbation.
- Collect emitted photons through a 460/80 nm bandpass filter.
- Acquire images (256x256 pixels) with a pixel dwell time sufficient to collect >500 photons at the peak.
- Pharmacological Perturbation (Optional):
- Acquire a baseline FLIM image.
- Treat cells with 10 µM Rotenone/Antimycin A and image after 30 minutes to shift metabolism toward glycolysis (expected increase in free NADH, shorter mean lifetime).
- Alternatively, treat with 50 mM 2-DG and image after 60 minutes to inhibit glycolysis (expected change in bound/free ratio).
- Data Analysis:
- Fit decays with a bi-exponential model:
I(t) = α₁ exp(-t/τ₁) + α₂ exp(-t/τ₂). - Report the mean lifetime (τm) and the fractional contributions (α₁%, α₂%). The long-lifetime component (τ₂, α₂%) is associated with protein-bound NAD(P)H and is often used as a metabolic index.
- Fit decays with a bi-exponential model:
Table 2: Representative FLIM Data for Redox Probes
| Probe/Target | Reduced/Free State Lifetime (τ) | Oxidized/Bound State Lifetime (τ) | Lifetime Shift Direction upon Oxidation/Binding | Typical Biological Context |
|---|---|---|---|---|
| roGFP1-Grx1 | ~2.9 ns (fully reduced) | ~2.4 ns (fully oxidized) | Decrease | Cytosolic glutathione redox potential. |
| NAD(P)H (Endogenous) | Free: τ₁ ~0.4 ns (α₁%) | Bound: τ₂ ~2.0-4.0 ns (α₂%) | Increase in τm and α₂% upon binding | Metabolic shift to oxidative phosphorylation. |
| HyPer (H₂O₂) | ~3.1 ns (basal) | ~2.7 ns (H₂O₂ oxidized) | Decrease | Hydrogen peroxide dynamics. |
| FLIM-FRET Sensor | Donor alone: τ ~3.0 ns | FRET with acceptor: τ < 3.0 ns | Decrease | Conformational changes in redox-sensitive proteins. |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for FLIM-based Redox Sensing
| Reagent/Material | Function | Example Product/Catalog # |
|---|---|---|
| Genetically Encoded Sensor Plasmid | Enables specific, subcellularly targeted redox sensing. | pCAGGS-roGFP1-Grx1 (Addgene #64995) |
| Transfection Reagent | For delivery of plasmid DNA into mammalian cells. | Lipofectamine 3000 (Thermo Fisher L3000015) |
| Live-Cell Imaging Medium | Phenol-red free, buffered medium for maintaining cell health during imaging. | FluoroBrite DMEM (Gibco A1896701) |
| Redox Control Reagents | For defining the dynamic range (fully reduced/oxidized states) of the sensor. | DTT (Sigma 43815), Diamide (Sigma D3648) |
| Metabolic Inhibitors | For perturbing and validating metabolic redox readouts (e.g., NAD(P)H FLIM). | Rotenone (Sigma R8875), 2-Deoxy-D-glucose (Sigma D8375) |
| FLIM Calibration Standard | A fluorophore with known, stable lifetime for daily system calibration. | Fluorescein (0.1M NaOH, τ ~4.0 ns) (Sigma 46955) |
| Mounting Medium (for fixed samples) | For preserving fluorescence lifetime properties post-fixation (if required). | ProLong Glass (Thermo Fisher P36980) |
Visualizations
FLIM Redox Sensing Principle
roGFP FLIM Experimental Workflow
NAD(P)H FLIM Reports Metabolic State
A Practical Guide to FLIM-Based Redox Biosensors: From Design to Live-Cell Imaging
This application note details the use of genetically encoded biosensors based on redox-sensitive green fluorescent proteins (roGFPs) within the broader thesis of Fluorescence Lifetime Imaging Microscopy (FLIM) for thiol-disulfide redox research. FLIM biosensing offers a powerful, quantitative, and rationetric-independent method to monitor cellular redox dynamics, providing insights into oxidative stress, signaling, and disease mechanisms critical for researchers and drug development professionals.
Biosensor Principles & Key Variants
roGFPs are engineered GFPs containing two surface-exposed cysteine residues that form a disulfide bond upon oxidation, causing a conformational shift that alters the excitation spectrum. Grx1-roGFP fuses human glutaredoxin-1 to roGFP, enabling rapid and specific equilibration with the glutathione redox couple (GSH/GSSG). FLIM measures the fluorescence decay rate (lifetime), which is sensitive to the roGFP's conformational state, offering advantages over intensity-based rationetry, including reduced artifacts from sensor concentration, excitation light path, or photobleaching.
Key Variants Table:
| Biosensor Name | Redox Couple Specificity | Dynamic Range (Lifetime Change)* | Typical τ (ns) Reduced* | Primary Application |
|---|---|---|---|---|
| roGFP1 | General Thiol Disulfide | ~0.3-0.5 ns | ~2.9 ns | Cytosol, ER Lumen |
| roGFP2 | General Thiol Disulfide | ~0.4-0.6 ns | ~2.8 ns | Mitochondrial Matrix, Cytosol |
| Grx1-roGFP2 | GSH/GSSG (2GSH/GSSG) | ~0.5-0.7 ns | ~2.8 ns | Cytosolic GSH Redox Potential |
| rxYFP (Grx1-rxYFP) | GSH/GSSG | ~0.2-0.3 ns | ~2.2 ns | Alternative to roGFP |
| roGFP1-R12 | General Thiol Disulfide | ~0.3-0.5 ns | ~2.9 ns | Nucleus |
| roGFP2-Orp1 | H₂O₂ (via Orp1) | ~0.5-0.7 ns | ~2.8 ns | Peroxides |
Note: Fluorescence lifetime values (τ) are approximate and depend on instrumentation, pH, and cellular environment. The dynamic range refers to the change in lifetime between fully reduced and oxidized states.
Detailed Experimental Protocols
Protocol 1: FLIM Measurement of roGFP2 Redox State in Live Mammalian Cells
Objective: Quantify the cytosolic glutathione redox potential using Grx1-roGFP2 via FLIM.
Materials & Reagents:
- Cell Line: HeLa or HEK293 cells expressing Grx1-roGFP2 (transient or stable).
- Imaging Buffer: Hanks' Balanced Salt Solution (HBSS) with 10mM HEPES, pH 7.4.
- Control Reagents:
- 10mM Dithiothreitol (DTT) in water (reducing agent).
- 2mM Diamide in water (thiol-oxidizing agent).
- 1mM H₂O₂ in imaging buffer (oxidative stress inducer).
- FLIM System: Confocal or multiphoton microscope with time-correlated single photon counting (TCSPC) module, pulsed laser at 488 nm.
Procedure:
- Cell Preparation: Seed cells on glass-bottom dishes. Transfect/transduce with Grx1-roGFP2 plasmid/virus 24-48h before imaging.
- System Calibration: Perform daily calibration using a fluorescent dye with a known, single-exponential lifetime (e.g., Fluorescein, τ ~4.0 ns in 0.1M NaOH).
- Image Acquisition:
- Maintain cells at 37°C/5% CO₂.
- Use a 40x or 60x oil-immersion objective.
- Excite at 488 nm, collect emission at 500-550 nm.
- Acquire FLIM data until 1000-2000 photons are collected in the brightest pixel for sufficient decay curve fitting.
- Lifetime Analysis:
- Fit fluorescence decay curves per pixel to a bi-exponential model:
I(t) = α₁ exp(-t/τ₁) + α₂ exp(-t/τ₂) + C. - Calculate the amplitude-weighted mean fluorescence lifetime:
τₘ = (α₁τ₁ + α₂τ₂) / (α₁ + α₂). - Generate lifetime maps (τₘ).
- Fit fluorescence decay curves per pixel to a bi-exponential model:
- In-Situ Calibration (for each experiment):
- Acquire a baseline FLIM image.
- Treat cells with 10mM DTT for 10 min (fully reduced state, τred), acquire image.
- Wash 3x with buffer.
- Treat cells with 2mM Diamide for 10 min (fully oxidized state, τox), acquire image.
- Data Normalization: Calculate the normalized redox index (RI) from lifetime:
RI_lifetime = (τₘ - τ_red) / (τ_ox - τ_red). An RI of 0 = fully reduced, 1 = fully oxidized.
Protocol 2: Rationetric Calibration (for Correlation with FLIM)
Objective: To correlate FLIM data with established rationetric measurements.
- Acquire excitation rationetric images (ex: 405/488 nm, em: 500-550 nm) of the same cells pre- and post-treatments (DTT/Diamide).
- Calculate ratio (R) = Intensity(405 nm)/Intensity(488 nm).
- Determine Rox and Rred from diamide and DTT treatments.
- Compute rationetric redox index:
RI_ratio = (R - R_red)/(R_ox - R_red). - Correlate
RI_lifetimewithRI_ratioto validate FLIM measurements.
The Scientist's Toolkit: Essential Research Reagent Solutions
| Item Name | Function/Benefit | Example Supplier/Cat. No. (for information) |
|---|---|---|
| Grx1-roGFP2 Plasmid | Encodes the specific GSH/GSSG biosensor. | Addgene, #64995 |
| Lipofectamine 3000 | High-efficiency transfection reagent for plasmid delivery. | Thermo Fisher, L3000015 |
| Dithiothreitol (DTT) | Strong reducing agent for in-situ calibration (fully reduced state). | Sigma-Aldrich, D0632 |
| Diamide | Thiol-oxidizing agent for in-situ calibration (fully oxidized state). | Sigma-Aldrich, D3648 |
| CellMask Deep Red | Plasma membrane stain for cell segmentation in FLIM analysis. | Thermo Fisher, C10046 |
| Fluorescein (0.1M NaOH) | FLIM system calibration standard with known single-exponential decay. | Sigma-Aldrich, 46955 |
| Poly-D-Lysine | Coats glass-bottom dishes for improved cell adhesion. | Sigma-Aldrich, P7280 |
| H₂O₂ (Hydrogen Peroxide) | Inducer of physiological oxidative stress for challenge experiments. | Sigma-Aldrich, H1009 |
| Butylated Hydroxytoluene (BHT) | Antioxidant for imaging media to prevent auto-oxidation during long experiments. | Sigma-Aldrich, B1378 |
Signaling Pathways & Experimental Workflows
Title: roGFP Redox Sensing Pathway for FLIM
Title: FLIM-roGFP Experimental Workflow
Fluorescence Lifetime Imaging Microscopy (FLIM) is a powerful quantitative technique for studying cellular processes, including thiol-disulfide redox dynamics. Unlike intensity-based measurements, FLIM reports on the average time a fluorophore spends in its excited state, a parameter that is highly sensitive to the local molecular environment but is independent of probe concentration, excitation light intensity, and photobleaching. This makes it ideal for studying subtle changes in cellular microenvironments. Within the context of a thesis on FLIM biosensing for redox research, this document focuses on the principles, selectivity, and application of small-molecule fluorescent probes designed for FLIM-based detection of key biological thiols, with a primary example being glutathione (GSH).
The core principle involves designing a probe whose fluorescence lifetime changes upon a specific biochemical reaction with the target analyte. For selective GSH detection, probes often exploit the unique nucleophilicity and concentration disparity of GSH over other biothiols like cysteine (Cys) and homocysteine (Hcy). GSH is typically present at millimolar concentrations (1-10 mM), while Cys and Hcy are at much lower micromolar levels.
Probe Design Principles and Selectivity Mechanisms
Selective FLIM probes for GSH operate on several design principles:
- Differential Reaction Kinetics (Nucleophilic Aromatic Substitution): A common strategy uses an electron-deficient aromatic ring (e.g., 2,4-dinitrobenzenesulfonyl) as a quenching and reactive group. GSH, being a strong nucleophile (thiolate anion at physiological pH), undergoes a faster nucleophilic aromatic substitution reaction with this group compared to Cys/Hcy, leading to cleavage and fluorescence dequenching. The change in the local electrostatic and hydrophobic environment upon cleavage alters the fluorophore's lifetime.
- GSH-S-Transferase (GST)-Mediated Reaction: Some probes are designed as substrates for GST, an enzyme that catalyzes the conjugation of GSH to electrophilic centers, enhancing selectivity.
- Disulfide Exchange: Probes containing a disulfide bond can undergo thiol-disulfide exchange. Selectivity for GSH can be tuned by the steric and electronic properties of the disulfide, favoring reaction with the abundant GSH to form a mixed disulfide adduct, which has a distinct fluorescence lifetime.
The selectivity is quantified by parameters like reaction rate constants (k~GSH~ >> k~Cys~) and detection limits. The FLIM readout directly visualizes the lifetime shift (τ), which correlates with local GSH concentration or redox potential.
Table 1: Characteristics of Exemplary Small-Molecule FLIM Probes for GSH
| Probe Name | Core Fluorophore | Reactive/Quenching Group | Lifetime Change (τ~off~ → τ~on~) | Selectivity (GSH vs. Cys/Hcy) | App. K~d~ or LOD for GSH | Primary Application (Cell Type) |
|---|---|---|---|---|---|---|
| NBD-Based Probe | 7-Nitrobenz-2-oxa-1,3-diazole (NBD) | 2,4-Dinitrobenzenesulfonyl (DNBS) | ~1.0 ns → ~8.5 ns | >50-fold | LOD: ~0.5 µM in vitro | Cytosolic GSH imaging (HeLa, MCF-7) |
| Rhodol-Based Probe | Rhodol | 2,4-Dinitrobenzenesulfonyl (DNBS) | ~0.8 ns → ~3.9 ns | >100-fold (kinetic) | -- | Redox stress monitoring (HepG2) |
| Cy3/Cy5-Based | Cyanines | Disulfide (S-S) | Cy3: ~0.3 ns → ~1.6 ns | ~20-fold (by concentration) | -- | Ratiometric FLIM, in vivo models |
| GST-Activated Probe | Silicon Rhodamine (SiR) | Chlorine (GST substrate) | ~1.2 ns → ~2.8 ns | Enzyme-dependent | -- | GST activity/GSH in tumors |
Table 2: Typical FLIM Acquisition Parameters for GSH Probes
| Parameter | Typical Setting | Notes |
|---|---|---|
| Excitation Source | Pulsed Diode Laser (470-510 nm) or Ti:Sapphire Laser | Depends on probe absorbance. |
| Pulse Repetition Rate | 20-40 MHz | Must be >1/τ to avoid pulse pile-up. |
| Detection | TCSPC or gated CCD | TCSPC is gold standard for accuracy. |
| Acquisition Time | 30-180 seconds | Balances S/N ratio and live-cell viability. |
| Lifetime Fit Model | Bi-exponential or phasor approach | Probes often exhibit two distinct lifetimes. |
| Reference Standard | Fluorescein (τ ≈ 4.1 ns in 0.1M NaOH) | For instrument calibration. |
Experimental Protocols
Protocol 1:In VitroCalibration and Selectivity Assay
Objective: To determine the fluorescence lifetime response and selectivity of a GSH probe. Materials: Probe stock solution (in DMSO), GSH, Cys, Hcy stock solutions (in PBS, pH 7.4), PBS buffer (10 mM, pH 7.4 with 1% DMSO), 96-well plate or quartz cuvette, FLIM microscope or time-resolved fluorometer. Procedure:
- Prepare 1 mL solutions of the probe (e.g., 1 µM) in PBS buffer.
- Kinetic/Selectivity Test: Add aliquots of each thiol (GSH, Cys, Hcy) to final concentrations of 0, 10, 50, 100, 500 µM, and 1, 5 mM. Incubate at 37°C.
- FLIM Measurement: At defined time points (e.g., 0, 5, 15, 30, 60 min), place a sample drop on a slide or in a cuvette. Acquire FLIM data using standard settings (e.g., 470 nm excitation, 500-550 nm emission filter, 30s acquisition).
- Data Analysis: Fit decay curves to a bi-exponential model. Plot the average lifetime (τ~avg~) or amplitude-weighted lifetime against thiol concentration and time. The probe selective for GSH will show a significant, rapid lifetime shift only with GSH, even at lower mM concentrations.
Protocol 2: Live-Cell FLIM Imaging of Glutathione
Objective: To image basal and perturbed GSH levels in live cells. Materials: Adherent cells (e.g., HeLa), culture medium, probe stock solution (in DMSO), N-ethylmaleimide (NEM, thiol blocker), Buthionine sulfoximine (BSO, GSH synthesis inhibitor), H~2~O~2~ (oxidative stress inducer), glass-bottom dish, FLIM microscope with environmental chamber (37°C, 5% CO~2~). Procedure:
- Cell Preparation: Seed cells in a glass-bottom dish and culture until 70-80% confluent.
- Probe Loading: Dilute the probe in serum-free medium to a final working concentration (typically 1-5 µM). Incubate cells for 20-30 min at 37°C.
- Washing: Rinse cells 3x with warm PBS or FluoroBrite/DMEM imaging medium.
- Control Measurement: Acquire FLIM images of untreated, probe-loaded cells to establish the basal lifetime map (τ~basal~).
- Perturbation Experiments:
- Depletion: Treat cells with BSO (100 µM, 4-24 hr pre-incubation) or NEM (100 µM, 15 min co-incubation with probe).
- Oxidation: Treat cells with H~2~O~2~ (100-500 µM, 15-30 min).
- FLIM Acquisition: For each condition, acquire FLIM images using identical parameters (laser power, gain, acquisition time). Use a 40x or 60x oil immersion objective.
- Analysis: Generate lifetime histograms and pseudocolor lifetime maps. Calculate the mean population lifetime for each condition. A decrease in lifetime (toward the "off" state) upon BSO/NEM treatment indicates successful GSH depletion.
Visualization Diagrams
Diagram 1: GSH Probe Activation via S_NAr
Diagram 2: Live-Cell FLIM Workflow for Redox
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for FLIM-based GSH Sensing Experiments
| Item | Function/Benefit | Example Product/Catalog # (Illustrative) |
|---|---|---|
| GSH-Selective FLIM Probe | Small molecule whose fluorescence lifetime increases upon specific reaction with glutathione. | e.g., Mito or Cyto-targeted DNBS-based probes (commercially available or synthesized). |
| TCSPC FLIM System | Gold-standard system for precise lifetime measurement at each pixel; provides high accuracy. | Becker & Hickl SPC-150; PicoQuant SymPhoTime; Leica STELLARIS 8 FALCON. |
| Pulsed Laser Diode | Provides repetitive, short-pulse excitation at wavelengths matching probe absorption. | LDHP-C-470 (470 nm) or LDH-D-C-510 (510 nm) with PDL 800-D driver. |
| Glass-Bottom Dishes | High-quality #1.5 coverslip bottom for optimal optical resolution and live-cell imaging. | MatTek P35G-1.5-14-C or ibidi µ-Dish 35 mm, high. |
| Thiol Modulators | Pharmacological tools to perturb cellular GSH levels for validation experiments. | BSO (Buthionine sulfoximine, GSH synthesis inhibitor). NEM (N-ethylmaleimide, thiol alkylator). |
| Fluorescence Lifetime Reference Standard | Dye with known, stable lifetime for daily instrument calibration and verification. | Fluorescein in 0.1 M NaOH (τ ≈ 4.1 ns). Rose Bengal in water (τ ≈ 0.16 ns). |
| Phasor Analysis Software | Simplifies lifetime data analysis, allowing for graphical representation and unmixing of components without fitting. | SimFCS (LFD, UC Irvine) or vendor-specific implementations (e.g., Leica LAS X). |
| HPLC-Grade DMSO | High-purity solvent for preparing probe stock solutions, minimizing background fluorescence. | Sigma-Aldrich D8418 or equivalent, anhydrous, ≥99.9%. |
| Physiological Buffer for Imaging | Phenol-red free medium with stable pH under atmospheric conditions for live-cell imaging. | Gibco FluoroBrite DMEM or HBSS with 10-20 mM HEPES. |
Within the context of FLIM biosensing for thiol-disulfide redox research, the selection of an appropriate lifetime measurement technique is critical. The redox state of cellular thiols, primarily glutathione (GSH), and disulfide bonds is a fundamental regulator of cellular homeostasis, signaling, and disease progression. FLIM provides a robust, quantitative method for monitoring redox-sensitive fluorescent biosensors, as fluorescence lifetime is independent of probe concentration and laser intensity, but exquisitely sensitive to the local molecular environment. This application note compares three principal FLIM methodologies—Two-Photon Excitation, Time-Correlated Single Photon Counting (TCSPC), and Frequency Domain (FD)—detailing their principles, protocols, and applications in redox biosensing.
Principle & Application to Redox Sensing
Two-photon (2P) excitation utilizes near-infrared (NIR) photons to excite fluorophores via the quasi-simultaneous absorption of two lower-energy photons. This is particularly advantageous for redox research in live cells and tissues:
- Deep Tissue Imaging: NIR light scatters less and penetrates deeper (up to ~1 mm) into biological samples, enabling studies in tissue explants or 3D organoids.
- Reduced Phototoxicity & Autofluorescence: Confined excitation volume minimizes out-of-focus photodamage and lowers cellular autofluorescence background, crucial for long-term imaging of dynamic redox processes.
- Compatibility: Typically coupled with TCSPC for lifetime acquisition (2P-TCSPC FLIM).
Experimental Protocol: 2P-TCSPC FLIM for roGFP Biosensors
Objective: Quantify the redox state in live HeLa cells expressing the redox-sensitive biosensor roGFP2-Orp1 using 2P-TCSPC FLIM.
Key Reagents & Materials:
- roGFP2-Orp1 expressing cells: Genetically encoded sensor where roGFP2 fluorescence lifetime is modulated by H₂O₂ via Orp1.
- Imaging Buffer: Hanks' Balanced Salt Solution (HBSS) with 20 mM HEPES, pH 7.4.
- Redox Modulators: 1-5 mM Dithiothreitol (DTT, reducing agent) or 1-2 mM H₂O₂ (oxidizing agent) for calibration.
- Two-Photon Microscope: equipped with a mode-locked Ti:Sapphire laser (~900 nm for roGFP), high-sensitivity detectors (e.g., PMT or GaAsP), and TCSPC electronics.
Procedure:
- Sample Preparation: Seed cells on glass-bottom dishes. Transfect with roGFP2-Orp1 plasmid 24-48 hours prior to imaging.
- System Calibration: Measure the instrument response function (IRF) using a scattering sample (e.g., saturated potassium iodide solution or fluorescent dye with sub-nanosecond lifetime).
- Acquisition Settings: Set laser power to the minimum required for a sufficient photon count rate (<1-5% of laser repetition rate to avoid pile-up). Typical pixel dwell time: 10-50 µs. Acquire images until 100-1000 photons are collected at the peak pixel for a robust fit.
- Lifetime Calculation: Fit decay curves per pixel to a double-exponential model:
I(t) = α₁ exp(-t/τ₁) + α₂ exp(-t/τ₂) + C. Calculate the amplitude-weighted mean lifetime:τ_mean = (α₁τ₁ + α₂τ₂) / (α₁ + α₂). - Redox Calibration: Acquire lifetimes after sequential treatment with DTT (fully reduced state, τred) and H₂O₂ (fully oxidized state, τox). The redox ratio can be expressed as:
Oxidation Degree = (τ_sample - τ_red) / (τ_ox - τ_red).
Time-Correlated Single Photon Counting (TCSPC) FLIM
Principle & Application to Redox Sensing
TCSPC is a digital timing technique that builds a histogram of photon arrival times relative to the excitation pulse. It is the gold standard for time-domain FLIM due to its high precision and suitability for fast, low-light measurements.
- High Accuracy: Ideal for resolving multi-exponential decays common in biosensors interacting with multiple molecular species.
- Quantitative Precision: Excellent signal-to-noise ratio at low photon counts, perfect for monitoring subtle redox shifts.
- Versatility: Can be coupled with both single-photon (confocal) and multi-photon excitation.
Experimental Protocol: Confocal TCSPC FLIM with a Small-Molecule Redox Probe
Objective: Measure mitochondrial redox potential in live cells using the small-molecule probe Mito-roGFP (or equivalent) via confocal TCSPC FLIM.
Key Reagents & Materials:
- Mito-roGFP or similar probe: e.g., CellROX Deep Red (lifetime sensitive to oxidative stress).
- Live cells with stained mitochondria.
- Confocal Microscope with TCSPC: Pulsed laser (e.g., 485 nm LED or 510 nm laser diode), high-speed detector, TCSPC module (e.g., SPC-150, HydraHarp).
Procedure:
- Staining: Incubate cells with 1-5 µM Mito-roGFP or CellROX Deep Red in imaging buffer for 30 min at 37°C. Wash thoroughly.
- IRF Measurement: Use a reference dye with a known single-exponential decay (e.g., fluorescein at pH high, τ ~4 ns).
- Acquisition: Use low excitation power. Set the TCSPC module for optimal time resolution (e.g., 256 time bins per decay). Collect data until the peak channel of the histogram reaches 10,000 counts for a good fit.
- Data Analysis: Perform a bi-exponential tail fit of the decay histogram, excluding the IRF region. Analyze the short lifetime component (τ₁), which is often more sensitive to redox changes. Create lifetime maps (τ_mean or τ₁) for visualization.
Frequency Domain (FD) FLIM
Principle & Application to Redox Sensing
FD-FLIM modulates the intensity of the excitation light at high frequencies (10s-100s MHz) and measures the phase shift (τφ) and demodulation (τm) of the emitted fluorescence.
- High-Speed Acquisition: Can be faster than TCSPC for single-lifetime measurements, suitable for high-throughput screening of redox-active drug candidates.
- Real-Time Imaging: Enables video-rate FLIM for monitoring rapid redox fluctuations.
- Instrumentation: Can be implemented with gain-modulated detectors (e.g., PMTs) or using specialized CMOS cameras.
Experimental Protocol: Wide-field FD-FLIM for High-Throughput Redox Screening
Objective: Screen a compound library for redox-modulating effects in cells expressing a redox biosensor using wide-field FD-FLIM.
Key Reagents & Materials:
- Cells in a 96- or 384-well plate expressing a homogeneous lifetime redox sensor (e.g., Grx1-roGFP2).
- FD-FLIM System: Wide-field epifluorescence microscope with an intensity-modulated light source (e.g., LED or laser diode) and a modulated image intensifier coupled to a CCD/CMOS camera.
- Liquid handling robot for compound addition.
Procedure:
- Plate Preparation: Seed and transfect cells in optical-bottom microplates. Pre-incubate with compounds or controls.
- System Calibration: Measure the system's frequency response using a reference dye of known lifetime.
- Rapid Acquisition: For each well, acquire a series of phase-sensitive images (typically at 3-12 different phase offsets). Modulation frequency is typically set near the inverse of the expected lifetime (e.g., ~80 MHz for a ~2 ns lifetime).
- Lifetime Calculation: Compute the phase (τφ) and modulation (τm) lifetimes per pixel from the phase stack. For a single-exponential decay, τφ and τm are equal. The mean lifetime can be calculated as
τ_mean = sqrt(τ_φ * τ_m). - Analysis: Average the τ_mean across all cells in a well. Normalize to positive (oxidizing) and negative (reducing) controls to generate a Z'-factor for assay quality and identify hit compounds.
Table 1: Quantitative Comparison of FLIM Techniques for Redox Biosensing
| Feature | Two-Photon (with TCSPC) | Time-Domain TCSPC (Confocal) | Frequency Domain (Wide-field) |
|---|---|---|---|
| Typical Lifetime Resolution | < 10 ps | < 5 ps | ~50 ps |
| Temporal Dynamic Range | Very High | Very High | Moderate |
| Acquisition Speed (per frame) | Slow (0.5 - 5 s) | Slow to Moderate (0.1 - 2 s) | Fast (≤ 0.1 s) |
| Excitation Photon Energy | Low (NIR) | High (Visible) | High (Visible) |
| Optical Sectioning | Excellent (Intrinsic) | Good (Confocal Pinhole) | Poor (Requires deconvolution) |
| Photodamage Risk | Low | Moderate | Moderate-High |
| Best for Sample Type | Deep tissue, live animals, 3D cultures | Live cells, fixed tissues | High-throughput, rapid kinetics |
| Primary Redox Application | Redox imaging in thick specimens & long-term dynamics | Precise quantification of multi-exponential decays in subcellular compartments | Screening redox-modulating drugs & fast cellular responses |
Visualization of Workflows
Title: Workflow for Two-Photon TCSPC FLIM Imaging
Title: Principle of Frequency Domain FLIM Measurement
Title: Decision Tree for Selecting FLIM Techniques
The Scientist's Toolkit: Key Research Reagents & Materials
Table 2: Essential Reagents for FLIM-based Thiol-Disulfide Redox Research
| Item | Function in Redox FLIM Experiments |
|---|---|
| roGFP2 (or rxYFP) based biosensors (e.g., roGFP2-Orp1, Grx1-roGFP2) | Genetically encoded, rationetric (intensity) or lifetime-sensitive probes for specific redox couples (e.g., GSH/GSSG, H₂O₂). |
| Small-molecule redox probes (e.g., CellROX Deep Red, MitoTracker Red CM-H2XRos) | Chemical dyes whose fluorescence lifetime changes with oxidative stress; used for specific organelles like mitochondria. |
| Dithiothreitol (DTT) | Strong reducing agent used to fully reduce thiol groups in biosensors for calibration (defines τ_red). |
| Hydrogen Peroxide (H₂O₂) | Common oxidizing agent used to fully oxidize biosensors for calibration (defines τ_ox). |
| N-Acetylcysteine (NAC) | Cell-permeable antioxidant and precursor to GSH; used as a positive control for reductive stress. |
| Menadione or Antimycin A | Pharmacological agents to induce mitochondrial ROS production (oxidative stress positive control). |
| Hanks' Balanced Salt Solution (HBSS) with HEPES | Physiological imaging buffer that maintains pH without CO₂ control during microscopy. |
| Glass-bottom culture dishes (No. 1.5 coverslip) | Optimal for high-resolution microscopy, providing the correct refractive index and thickness for oil-immersion objectives. |
| Reference Fluorophores (e.g., Fluorescein, Rose Bengal) | Dyes with stable, known single-exponential lifetimes for daily system calibration and IRF verification. |
Step-by-Step Protocol for Live-Cell Redox FLIM Imaging
1. Introduction Within the broader thesis on FLIM biosensing for thiol-disulfide redox research, this protocol details the application of Fluorescence Lifetime Imaging Microscopy (FLIM) to quantitatively assess cellular redox states in live cells. FLIM of genetically encoded biosensors, such as roGFP or rxYFP, provides a ratiometric, quantitative, and environment-insensitive measure of thiol oxidation, crucial for studying redox signaling in physiology, disease models, and drug development.
2. Key Research Reagent Solutions
| Reagent / Material | Function in Experiment |
|---|---|
| Genetically Encoded Redox Sensor (e.g., roGFP2, Grx1-roGFP2, rxYFP) | Engineered fluorescent protein whose fluorescence lifetime (and intensity) changes reversibly upon cysteine thiol oxidation/reduction. The primary biosensor. |
| Lipid-Based Transfection Reagent (e.g., Lipofectamine 3000) | For delivery of plasmid DNA encoding the redox biosensor into mammalian cell lines. |
| Phenol Red-Free Imaging Medium | Culture medium without phenol red to minimize background fluorescence during sensitive FLIM measurements. |
| Redox Modulators: Dithiothreitol (DTT) | Strong reducing agent (positive control) to fully reduce sensor thiols. |
| Redox Modulators: Hydrogen Peroxide (H₂O₂) | Oxidizing agent (positive control) to fully oxidize sensor thiols. |
| Pharmacologic Agents (e.g., Auranofin, Paraquat) | Tool compounds to perturb specific redox pathways (thioredoxin inhibition, ROS induction) in experimental settings. |
| Mounting Chamber with Gas Permeability (e.g., Lab-Tek) | For maintaining cells under controlled temperature and CO₂ during extended live-cell imaging. |
3. Detailed Experimental Protocol
3.1. Cell Preparation and Biosensor Expression
- Day 1: Seed appropriate cell line (e.g., HeLa, HEK293) into a 35-mm glass-bottom imaging dish at ~50% confluence.
- Day 2: Transfert cells with plasmid encoding the redox biosensor (e.g., pLVX-Grx1-roGFP2) using recommended transfection protocol. Use a transfection reagent-to-DNA ratio optimized for your cell line (e.g., 3 µL Lipofectamine 3000: 1 µg DNA).
- Day 3-4: Allow 24-48 hours for robust biosensor expression. For stable expression, generate lentiviral particles and transduce cells, followed by antibiotic selection.
3.2. FLIM System Calibration & Setup
- Turn on multiphoton or confocal FLIM system (e.g., Ti:Sapphire laser coupled to time-correlated single photon counting (TCSPC) module) at least 1 hour prior.
- Calibrate the system using a standard fluorescent dye with a known, single-exponential lifetime (e.g., Fluorescein, τ ~4.0 ns in 0.1 M NaOH) to check instrument response function (IRF).
- Set imaging parameters: Excitation: 960 nm (for two-photon) or 488 nm (for confocal); Emission Filter: 500-550 nm bandpass; Laser Power: Minimize to avoid phototoxicity (typically 0.1-2% for multiphoton); Pixel Dwell Time: 10-50 µs; Accumulate 300-1000 photons per pixel for robust lifetime fitting.
3.3. Live-Cell Imaging and Treatment Workflow
- Replace culture medium with pre-warmed, phenol-red free imaging medium.
- Mount dish on the microscope stage equilibrated to 37°C and 5% CO₂.
- Locate a field of view with healthy, well-expressing cells.
- Acquire a pre-treatment FLIM dataset: Collect a minimum of 3-5 image stacks for statistical analysis.
- Administer treatment: Carefully add redox modulator or drug candidate directly to the dish (e.g., 1 mM H₂O₂ final concentration, 5 mM DTT). Gently swirl to mix.
- Acquire time-series FLIM data: Initiate sequential FLIM acquisitions every 30-60 seconds for 15-30 minutes to capture dynamics.
3.4. Data Analysis & Quantification
- Fit fluorescence decay curves per pixel using a bi-exponential model:
I(t) = α₁exp(-t/τ₁) + α₂exp(-t/τ₂) + C, where τ are lifetimes and α are fractional amplitudes. - Calculate the amplitude-weighted mean lifetime:
τ_mean = (α₁τ₁ + α₂τ₂) / (α₁ + α₂). - Generate pseudocolor τ_mean maps for visualization.
- Quantify data by defining regions of interest (ROIs) for the cytosol, nucleus, or mitochondria. Report τ_mean as mean ± SEM from n cells across N independent experiments.
- Optional: Calculate the redox ratio from intensity-based images if using a ratiometric sensor:
R = I(510 nm, Ex405 nm) / I(510 nm, Ex488 nm).
4. Representative Quantitative Data & Interpretation Table 1: Example FLIM Data for roGFP2 in HeLa Cells under Redox Challenges
| Condition | Amplitude-Weighted Mean Lifetime, τ_mean (ns) * | Redox State Interpretation |
|---|---|---|
| Fully Reduced (5 mM DTT) | 2.95 ± 0.05 | Baseline, reduced state of sensor cysteines. |
| Untreated Control | 2.75 ± 0.07 | Physiological resting redox potential. |
| Oxidized (1 mM H₂O₂) | 2.55 ± 0.04 | Fully oxidized state of sensor cysteines. |
| Auranofin (1 µM, 30 min) | 2.60 ± 0.06 | Thioredoxin system inhibition, shift toward oxidation. |
| N-Acetylcysteine (5 mM, 1 hr pre-treat) | 2.85 ± 0.06 | Antioxidant treatment, shift toward reduction. |
Note: *Example lifetime values are instrument-dependent. Absolute values vary; relative changes are key.
5. Diagrams of Workflows and Pathways
Title: Live-Cell Redox FLIM Experimental Workflow
Title: Redox Signaling to FLIM Readout Pathway
The development of therapeutics that modulate cellular thiol-disulfide redox states is a critical frontier in treating diseases involving oxidative stress, including cancer, neurodegeneration, and inflammatory disorders. Fluorescence Lifetime Imaging Microscopy (FLIM) biosensing provides a quantitative, non-invasive method to monitor dynamic redox changes in live cells and tissues. This application note details protocols for employing genetically encoded FLIM biosensors, such as roGFP2 and rxYFP, to screen and characterize drug candidates that alter the glutathione redox potential (EGSSG/2GSH).
Key Research Reagent Solutions & Materials
Table 1: Essential Toolkit for FLIM-based Redox Drug Screening
| Item | Function/Description | Example Supplier/Catalog |
|---|---|---|
| Genetically Encoded Redox Biosensor | FRET-based or single-FP sensor whose fluorescence lifetime changes with thiol redox state. | roGFP2-Orp1 (Addgene #64985); Grx1-roGFP2 (Addgene #64965) |
| FLIM-Compatible Live-Cell Imaging Medium | Phenol-red free medium with stable pH and minimal autofluorescence. | FluoroBrite DMEM (Thermo Fisher, A1896701) |
| Redox-Modulating Controls | Pharmacological agents to validate assay response (oxidants/reductants). | Dithiothreitol (DTT, reducing agent); Diamide (oxidizing agent) |
| FLIM Dye for Calibration (Optional) | Reference standard with known single-exponential lifetime. | Fluorescein (0.1M NaOH, τ ~4.0 ns) |
| 96/384-well Glass-bottom Plates | Plates compatible with high-resolution microscopy and compound addition. | CellVis (P96-1.5H-N) or Corning (3841) |
| Automated Liquid Handling System | For precise, high-throughput compound library addition. | Integra ViaFlo, or similar |
| FLIM Analysis Software | For rapid fitting of fluorescence decay curves and lifetime mapping. | SPCImage NG (Becker & Hickl), FLIMfit (Imperial College) |
Core Experimental Protocols
Protocol: High-Throughput FLIM Screening for Redox-Modulating Compounds
Objective: To identify hits from a compound library that alter cytosolic glutathione redox potential in live cells.
Materials:
- HEK293T or relevant disease model cell line stably expressing Grx1-roGFP2.
- FLIM-optimized 384-well plate.
- Compound library (e.g., 10µM final concentration per well).
- FLIM microscope with time-correlated single photon counting (TCSPC) and automated stage.
Procedure:
- Cell Seeding & Preparation: Seed cells expressing the biosensor at 10,000 cells/well in 384-well plates. Culture for 24-48 hours to reach ~80% confluence.
- Compound Treatment: Using an automated liquid handler, add compounds from the library. Include control wells: DTT (10mM, full reduction), Diamide (2mM, full oxidation), and DMSO vehicle (0.1%).
- FLIM Image Acquisition: 30 minutes post-treatment, acquire FLIM data.
- Excitation: 920 nm (two-photon) or 488 nm laser line.
- Emission Filter: 525/50 nm bandpass.
- Acquisition Time: Aim for >1000 photons per pixel in the brightest region; limit to 3 minutes per well for throughput.
- Maintain environmental control at 37°C, 5% CO2.
- Data Processing:
- Fit fluorescence decay curves per pixel to a double-exponential model:
I(t) = α₁exp(-t/τ₁) + α₂exp(-t/τ₂). - Calculate the amplitude-weighted mean fluorescence lifetime:
τ<sub>m</sub> = (α₁τ₁ + α₂τ₂) / (α₁ + α₂). - Generate false-color lifetime maps per well.
- Fit fluorescence decay curves per pixel to a double-exponential model:
- Hit Identification: Compounds inducing a statistically significant (e.g., >2 SD from DMSO mean) shift in τm are considered primary hits.
Table 2: Example FLIM Data from a Pilot Screen (Hypothetical Data)
| Condition | Mean Lifetime, τm (ns) | SD (ns) | % Change vs. DMSO | p-value (vs. DMSO) |
|---|---|---|---|---|
| DMSO (Vehicle) | 2.65 | 0.08 | -- | -- |
| DTT (Reducing Control) | 2.92 | 0.07 | +10.2% | <0.001 |
| Diamide (Oxidizing Control) | 2.41 | 0.09 | -9.1% | <0.001 |
| Compound A | 2.81 | 0.10 | +6.0% | 0.003 |
| Compound B | 2.48 | 0.08 | -6.4% | 0.002 |
Protocol: Dose-Response & IC/EC50Determination via FLIM
Objective: To characterize the potency of primary hit compounds. Procedure:
- Prepare a 10-point, 1:3 serial dilution of the hit compound (e.g., 100 µM to 0.5 nM).
- Treat biosensor-expressing cells in triplicate for each concentration.
- Acquire and process FLIM data as in Protocol 3.1.
- Plot τm (normalized to DTT/Diamide controls) vs. log[compound].
- Fit data to a four-parameter logistic curve to determine EC50 (for reductants) or IC50 (for oxidants).
Protocol: Target Pathway Validation via FLIM
Objective: To elucidate if a hit compound acts through a specific pathway (e.g., NRF2, Thioredoxin). Procedure:
- Use siRNA or CRISPR-Cas9 to knock down a candidate target gene (e.g., KEAP1, TXNRD1) in the biosensor cell line.
- Treat knockdown and control cells with the hit compound at its EC50 concentration.
- Acquire FLIM data. A significantly attenuated lifetime shift in the knockdown cells indicates the compound acts through that pathway.
Data Visualization & Workflow Diagrams
Diagram Title: FLIM-based redox drug discovery workflow.
Diagram Title: NRF2 pathway activation by redox-modulating compounds.
Within the broader thesis on FLIM (Fluorescence Lifetime Imaging) biosensing for thiol-disulfide redox research, this document provides detailed application notes and protocols for quantifying redox dynamics in specific cellular compartments. Compartmentalized redox regulation is critical for cellular function, signaling, and disease pathogenesis. FLIM-based biosensors offer a powerful, quantitative, and non-invasive method to measure redox potentials with high spatiotemporal resolution in living cells.
Research Reagent Solutions Toolkit
The following table details essential reagents and materials for performing compartment-specific redox imaging using FLIM.
| Reagent/Material | Function/Brief Explanation |
|---|---|
| roGFP2-Orp1 (e.g., mito-roGFP2-Orp1) | Genetically encoded biosensor targeted to mitochondria. roGFP2 is redox-sensitive; Orp1 specifically reacts with H₂O₂, enabling ratiometric, quantitative measurement of mitochondrial H₂O₂ dynamics. |
| ER-roGFP-iE | ER-targeted redox-sensitive GFP variant. The iE tag (improved Ero1) optimizes equilibration with the local ER redox pool, allowing accurate measurement of the more oxidative ER glutathione redox potential (EGSSG/2GSH). |
| NLS-roGFP2 | Redox-sensitive GFP fused to a Nuclear Localization Signal (NLS). Enables specific measurement of the nuclear glutathione redox state, which is crucial for transcription factor regulation and DNA repair. |
| Fluorescence Lifetime Microscope | System equipped with time-correlated single photon counting (TCSPC) capabilities, pulsed laser (e.g., 405 nm, 488 nm), and high-sensitivity detectors. FLIM measures the decay rate of the biosensor's fluorescence, which is sensitive to its redox state independent of concentration. |
| Dithiothreitol (DTT) | Strong reducing agent. Used as a control to fully reduce biosensor, defining the minimum fluorescence lifetime or 405/488 nm excitation ratio. |
| Hydrogen Peroxide (H₂O₂) | Oxidizing agent. Used as a control to fully oxidize the biosensor (often with aid of Orp1), defining the maximum fluorescence lifetime or ratio. |
| Antimycin A | Mitochondrial complex III inhibitor. Induces superoxide production from the electron transport chain, used to perturb mitochondrial redox state. |
| Tunicamycin | Induces ER stress by inhibiting N-linked glycosylation, leading to protein misfolding and altered ER redox balance. |
| Transfection Reagents (e.g., Lipofectamine 3000) | For delivery of plasmid DNA encoding compartment-targeted roGFP biosensors into mammalian cell lines. |
Table 1: Typical Compartment-Specific Redox Potentials (EGSSG/2GSH) in Mammalian Cells under Homeostasis. Data derived from published studies using roGFP-based imaging.
| Cellular Compartment | Approximate Redox Potential (mV) | Key Characteristics & Notes |
|---|---|---|
| Cytoplasm | -280 to -300 | Reducing environment, primary site of NADPH production. |
| Mitochondria | -290 to -320 | Matrix is reducing, but site of ROS production. Can become more oxidized during stress. |
| Endoplasmic Reticulum (ER) | -190 to -210 | More oxidative environment necessary for disulfide bond formation in proteins. |
| Nucleus | -270 to -290 | Generally similar to cytoplasm but can be regulated independently. |
| Peroxisomes | ~ -250 | Oxidative due to H₂O₂-producing enzymes. |
Table 2: FLIM Parameters for roGFP2 in Different Redox States. Example data for a generic roGFP2 biosensor excited at 920 nm (two-photon) with emission collected at 500-550 nm.
| roGFP2 Redox State | Average Fluorescence Lifetime (τ, nanoseconds) | Donor-to-Acceptor FRET Efficiency |
|---|---|---|
| Fully Reduced | ~2.9 ns | Low (Acceptor not fluorescent) |
| Fully Oxidized | ~1.7 ns | High (FRET to Acceptor active) |
| 50% Oxidized | ~2.3 ns | Intermediate |
Detailed Experimental Protocols
Protocol 1: Measuring Mitochondrial H₂O₂ Dynamics using Mito-roGFP2-Orp1 with FLIM
Objective: To quantitatively image hydrogen peroxide fluctuations in the mitochondrial matrix of live cells using FLIM.
Materials:
- Cells (e.g., HeLa, MEFs) cultured on 35mm glass-bottom dishes.
- Plasmid: pMito-roGFP2-Orp1.
- Transfection reagent.
- Live-cell imaging medium (e.g., FluoroBrite DMEM, without phenol red).
- 10 mM DTT stock (in water).
- 100 mM H₂O₂ stock (in water, prepare fresh).
- 10 mM Antimycin A stock (in DMSO).
- FLIM microscope system.
Procedure:
- Transfection: Transfect cells with pMito-roGFP2-Orp1 using standard protocols 24-48 hours prior to imaging to ensure robust expression.
- Microscope Setup: Configure the FLIM system for GFP/roGFP imaging. Use a pulsed 405 nm or 920 nm (two-photon) laser for excitation. Set emission filters to 500-550 nm. Calibrate TCSPC electronics.
- Sample Preparation: On imaging day, replace culture medium with pre-warmed live-cell imaging medium.
- Define Controls:
- Full Reduction: Acquire a FLIM image. Then, add DTT to a final concentration of 5-10 mM directly to the dish. Incubate for 5-10 minutes and acquire a second FLIM image.
- Full Oxidation: Carefully wash cells 2x with imaging medium to remove DTT. Add H₂O₂ to a final concentration of 1-5 mM. Incubate for 5-10 minutes and acquire a third FLIM image.
- Note: The order can be reversed (oxidize first, then reduce).
- Experimental Perturbation: Using a new dish of transfected cells, acquire a baseline FLIM image. Add Antimycin A (final conc. 1-10 µM) to the medium. Acquire sequential FLIM images every 1-5 minutes for 30-60 minutes.
- Data Analysis:
- Process FLIM data using specialized software (e.g., SPCImage, SymPhoTime, FLIMfit).
- Fit fluorescence decay curves per pixel to a double- or triple-exponential model. The average fluorescence lifetime (τavg) is the primary readout.
- Generate pseudocolored lifetime maps. The lifetime (τ) is inversely correlated with oxidation (shorter τ = more oxidized).
- Quantify τavg within regions of interest (ROIs) drawn on mitochondria.
Protocol 2: Quantifying ER Glutathione Redox Potential using ER-roGFP-iE
Objective: To determine the glutathione redox potential (EGSSG/2GSH) within the endoplasmic reticulum using ratiometric imaging (complementary to FLIM).
Materials:
- Cells cultured on glass-bottom dishes.
- Plasmid: pER-roGFP-iE.
- Live-cell imaging medium.
- DTT and H₂O₂ stocks.
- 10 mg/mL Tunicamycin stock (in DMSO).
- Fluorescence microscope with capability for rapid excitation switching between 405 nm and 488 nm, and emission collection at 500-550 nm.
Procedure:
- Transfection & Preparation: Transfect cells with pER-roGFP-iE 24-48h before imaging. Use live-cell imaging medium during the experiment.
- Ratiometric Imaging Setup: Set up sequential or rapid-alternating excitation at 405 nm and 488 nm. Use a single emission channel (500-550 nm).
- Calibration (in situ): For each cell/dish, perform a full reduction (5-10 mM DTT) and full oxidation (1-5 mM H₂O₂) sequence as in Protocol 1, capturing paired 405/488 nm excited images at each step.
- Experimental Treatment: Image a separate dish to establish a baseline ratio (405/488 nm emission). Add Tunicamycin (final conc. 2-5 µg/mL) to induce ER stress. Acquire image pairs at regular intervals (e.g., every 10 minutes for 2-4 hours).
- Data Analysis:
- Calculate the 405 nm/488 nm emission intensity ratio (R) for each pixel from the paired images.
- Determine the minimum (Rred) and maximum (Rox) ratios from the DTT and H₂O₂ calibration images.
- Calculate the oxidation degree: OxD = (R - Rred) / (Rox - Rred).
- Convert OxD to redox potential using the Nernst equation: E = E0 - (RT/nF) ln([GSH]²/[GSSG]), where the biosensor-specific E0 for roGFP2 is approximately -280 mV at pH 7.0. Use the relationship: [GSSG]/[GSH]² = OxD / (1 - OxD).
Visualizations
Diagram 1 Title: FLIM-Based Redox Biosensing Workflow
Diagram 2 Title: Key Redox Pathways & Perturbations by Compartment
Overcoming Challenges: Optimizing FLIM Redox Experiments for Reliable Data
Application Notes
In FLIM (Fluorescence Lifetime Imaging) biosensing for thiol-disulfide redox research, the quantification of cellular redox potential (E~h~) requires precise calibration of the biosensor within its native cellular environment. In situ titration using the reducing agent DTT (dithiothreitol) and the oxidizing agent diamide provides a critical method for establishing the sensor's dynamic range and response curve within live cells. This protocol details the application of this titration to ratiometric or FLIM-based redox biosensors, such as roGFP or rxYFP, enabling accurate conversion of sensor readouts into absolute E~h~ values.
This calibration is fundamental for studies investigating redox dysregulation in diseases like cancer or neurodegeneration, and for assessing the efficacy of redox-modulating drug candidates. The following tables summarize key quantitative data and reagents.
Table 1: Key Redox Reagents for In Situ Titration
| Reagent | Typical Working Concentration Range | Primary Function | Key Consideration |
|---|---|---|---|
| DTT (Dithiothreitol) | 0.1 - 10 mM | Reducing agent; fully reduces sensor to establish R~min~ or τ~min~. | Cell-permeable but can alter global redox balance; use at minimum effective dose. |
| Diamide (Azodicarboxylic acid bis(Dimethylamide)) | 0.5 - 5 mM | Thiol-specific oxidant; fully oxidizes sensor to establish R~max~ or τ~max~. | Acts rapidly; concentration must be optimized to avoid nonspecific stress. |
| Buffer/Solution | |||
| Live Cell Imaging Medium | N/A | Phenol-red free medium for imaging. | Must contain necessary supplements (e.g., glutamine) but no thiols (e.g., cysteine). |
Table 2: Example Calibration Data for a Hypothetical roGFP2 Biosensor
| Condition | Mean Ratio (R) (Ex405/Ex488) | Fractional Oxidation (OxD) | Calculated E~h~ (mV) at pH 7.2* |
|---|---|---|---|
| Fully Reduced (10 mM DTT) | 0.2 (R~min~) | 0.0 | N/A |
| Fully Oxidized (2 mM Diamide) | 1.8 (R~max~) | 1.0 | N/A |
| Untreated Control | 0.65 | 0.36 | -308 |
| Post-Drug Treatment | 1.25 | 0.75 | -265 |
*E~h~ = E~0~ - (59/n) * log((1-OxD)/OxD) at 30°C, where n=2 for roGFP, E~0~ ≈ -280 mV.
Experimental Protocols
Protocol 1:In SituTitration for Ratiometric (roGFP) Calibration
Objective: To generate a calibration curve by titrating the biosensor within cells to defined redox states.
Materials:
- Cells expressing the redox biosensor (e.g., roGFP2, rxYFP).
- Phenol-red free imaging medium.
- DTT stock solution (1 M in water, aliquoted, stored at -20°C).
- Diamide stock solution (500 mM in DMSO, aliquoted, stored at -20°C).
- Confocal or widefield fluorescence microscope with capable ratiometric imaging.
Method:
- Preparation: Seed cells on an imaging-appropriate dish. Allow to adhere and express biosensor for 24-48 hours.
- Baseline Imaging: Replace medium with fresh imaging medium. Acquire baseline ratiometric images (typically using 405 nm and 488 nm excitation, collecting emission >500 nm).
- Full Reduction: Add DTT from stock directly to the imaging medium to a final concentration of 5-10 mM. Incubate for 5-10 minutes at 37°C to fully reduce the biosensor. Acquire a second set of images (R~min~).
- Wash: Gently wash cells 3 times with pre-warmed, reagent-free imaging medium.
- Oxidation Titration: Perform a stepwise oxidation. Start by adding a low concentration of diamide (e.g., 10 µM). Incubate for 5 min, then image. Sequentially increase diamide concentration (e.g., 50 µM, 100 µM, 500 µM, 1 mM, 2 mM), imaging after each equilibration period.
- Full Oxidation: A final treatment with 2 mM diamide for 5 min establishes R~max~.
- Data Analysis: For each cell/pixel, calculate the oxidation degree: OxD = (R - R~min~) / (R~max~ - R~min~). Plot OxD against the known E~h~ of the DTT/diamide redox buffer at each step (can be calculated using the Nernst equation) to generate a sensor-specific calibration curve.
Protocol 2: FLIM-based Calibration for Lifetime Sensors
Objective: To correlate fluorescence lifetime (τ) changes with redox state.
Materials:
- Cells expressing a lifetime-based redox biosensor.
- Materials listed in Protocol 1.
- FLIM-capable multiphoton or confocal microscope.
Method:
- Baseline FLIM: Acquire fluorescence lifetime decay curves for the biosensor in untreated cells.
- Full Reduction & Oxidation: Treat cells sequentially with 10 mM DTT and 2 mM diamide as in Protocol 1, steps 3-6, acquiring a FLIM dataset after each treatment to establish τ~reduced~ and τ~oxidized~.
- Lifetime Analysis: Fit decay curves to a bi-exponential or phasor model. The mean lifetime or the fraction of molecules in the oxidized conformation is used as the readout.
- Calibration: The fractional oxidation can be derived from: OxD = (τ - τ~reduced~) / (τ~oxidized~ - τ~reduced~). This OxD can then be converted to E~h~ using the sensor's known midpoint potential (E~0~) via the Nernst equation.
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Experiment |
|---|---|
| roGFP2 or rxYFP Plasmid | Genetically-encoded biosensor whose cysteine thiols undergo reversible disulfide formation, altering fluorescence properties. |
| DTT (1M Stock) | Strong reducing agent used to fully reduce the biosensor, defining the "0% oxidized" state for calibration. |
| Diamide (500mM Stock) | Thiol-specific oxidant used to fully oxidize the biosensor, defining the "100% oxidized" state. |
| Phenol-Red Free Imaging Medium | Maintains cell health during imaging without introducing autofluorescence that interferes with ratio or lifetime measurements. |
| Hank's Balanced Salt Solution (HBSS) | A common, defined buffer for imaging experiments, ensuring ionic and pH stability. |
| Carbonyl Cyanide 3-Chlorophenylhydrazone (CCCP) | Mitochondrial uncoupler; optional control to collapse mitochondrial membrane potential when calibrating organelle-targeted sensors. |
Title: Redox Sensor Calibration Workflow
Title: Thiol-Disulfide Pathways in Research
In FLIM (Fluorescence Lifetime Imaging) biosensing for thiol-disulfide redox research, accurate quantification is paramount. The fluorescence lifetime of genetically encoded biosensors, such as roGFP or rxYFP, is sensitive not only to redox potential but also to confounding environmental factors. Changes in intracellular pH, dissolved oxygen concentration, and cellular autofluorescence can introduce significant artifacts, leading to misinterpretation of redox states. This application note details protocols to identify, quantify, and correct for these artifacts, ensuring robust and reliable redox measurements.
Impact of Environmental Factors on FLIM-Based Redox Sensing
pH Artifacts
Many fluorescent proteins used in redox biosensors exhibit pH-dependent protonation states, altering their fluorescence lifetime independently of redox state. The pKa of the chromophore can shift upon oxidation/reduction, creating a complex interplay.
Oxygen Concentration
Molecular oxygen is a primary quencher of fluorescent protein excited states via collisional quenching. Fluctuations in local oxygen concentration, especially in hypoxic tissues or 3D culture models, can directly modulate measured fluorescence lifetimes, mimicking or masking redox changes.
Native Autofluorescence
Cellular metabolites (e.g., NAD(P)H, FAD, lipofuscin) exhibit intrinsic fluorescence with lifetimes that can overlap with those of common biosensors. This autofluorescence contaminates the signal, skewing lifetime decay fits and calculated redox ratios.
Table 1: Quantitative Impact of Environmental Factors on Common Redox Biosensors
| Biosensor | Redox-Sensitive Lifetime Change (τ ox - τ red) | pH Sensitivity (Δτ/ΔpH) | O2 Quenching Constant (kq) | Primary Autofluorescence Overlap |
|---|---|---|---|---|
| roGFP2 | ~0.4 ns | High (~0.15 ns/pH unit near pKa) | Low | Moderate (NADPH) |
| rxYFP | ~0.8 ns | Moderate (~0.08 ns/pH unit) | Moderate | Low |
| Grx1-roGFP2 | ~0.4 ns | High (Inherited from roGFP2) | Low | Moderate (NADPH) |
| mrCPY1 | ~1.2 ns | Low (<0.03 ns/pH unit) | High | High (FAD) |
Protocols for Artifact Correction
Protocol 3.1: Simultaneous pH Monitoring and Correction
Objective: To measure and correct for pH-induced lifetime changes during redox FLIM experiments. Materials:
- FLIM microscope (TCSPC or phase-modulation system)
- Cells expressing both redox biosensor and a pH-insensitive reference fluorophore (e.g., mCherry) or a rationetric pH biosensor (e.g., pHluorin).
- Calibration buffers (pH 6.0 to 8.0).
- Ionophores (e.g., nigericin) for in situ calibration. Procedure:
- Dual-Channel FLIM Acquisition: Acquire lifetime images in two channels: one for the redox biosensor (e.g., 515/30 nm emission) and one for the pH reference/sensor (e.g., 610/60 nm for mCherry).
- In Situ pH Calibration: Perfuse cells with high-K+ calibration buffers of known pH (pH 6.0, 6.5, 7.0, 7.5, 8.0) containing 10 µM nigericin to equilibrate intra- and extracellular pH. Acquire FLIM data for both biosensors at each pH.
- Generate Correction Curve: Plot the redox biosensor's lifetime vs. the reference lifetime (or pH sensor ratio). Fit with a polynomial function to derive a correction algorithm.
- Apply Correction: During redox experiments, apply this function pixel-by-pixel to calculate the pH-corrected lifetime of the redox biosensor.
Protocol 3.2: Accounting for Oxygen Quenching
Objective: To quantify and normalize for oxygen-dependent quenching effects. Materials:
- Hypoxia chamber or microfluidic gas control system for the microscope stage.
- Oxygen-sensitive probe (e.g., Ru(Phen)3 or PtPFPP) for independent O2 mapping.
- FLIM system. Procedure:
- Characterize Sensor Sensitivity: Under controlled atmospheres (21%, 10%, 5%, 1% O2), acquire FLIM images of cells expressing only the redox biosensor. Plot lifetime (τ) vs. [O2] to determine quenching constant (kq) via Stern-Volmer analysis.
- Dual Sensing Experiment: Co-load cells with the redox biosensor and a referenced O2-sensitive probe (e.g., PtPFPP, lifetime-based). Acquire simultaneous dual-lifetime images.
- Lifetime Normalization: Using the predetermined kq, normalize the redox biosensor's lifetime in each pixel to the value expected at a standard reference oxygen level (e.g., 21%).
Protocol 3.3: Spectral Lifetime Unmixing of Autofluorescence
Objective: To isolate the biosensor's lifetime signal from contaminating autofluorescence. Materials:
- Multispectral or Hyperspectral FLIM system.
- Untransfected/unlabeled control samples. Procedure:
- Characterize Autofluorescence: Acquire spectral-FLIM data (λ emission, τ) from control cells. Fit decay curves across emission bands to establish the characteristic spectral-lifetime signature of autofluorescence.
- Acquire Sample Data: Collect full spectral-FLIM data from biosensor-expressing cells under experimental conditions.
- Linear Unmixing Analysis: Use a model (e.g., phasor-based or least-squares fitting) to unmix the total signal into contributions from the biosensor and autofluorescence components based on their distinct spectral-lifetime fingerprints. Report the unmixed biosensor lifetime.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Artifact Correction in Redox FLIM
| Item | Function & Rationale |
|---|---|
| Nigericin (K+ ionophore) | Clamps intracellular pH to extracellular buffer pH for in situ biosensor calibration. |
| High-K+ Calibration Buffers (pH 6.0-8.0) | Provide a range of known pH values without altering membrane potential during nigericin use. |
| Platinum(II)-5,10,15,20-tetrakis-(2,3,4,5,6-pentafluorphenyl)-porphyrin (PtPFPP) | Phosphorescent, lifetime-based O2 probe; inert and suitable for simultaneous imaging with GFP-based biosensors. |
| Ruthenium phenanthroline complex | Alternative oxygen-sensitive lifetime probe. |
| Sodium Dithionite (Na2S2O4) | Chemical reductant for defining fully reduced (τ_red) biosensor state during calibration. |
| Hydrogen Peroxide (H2O2) / Diamide | Chemical oxidants for defining fully oxidized (τ_ox) biosensor state during calibration. |
| Carbonyl Cyanide 3-Chlorophenylhydrazone (CCCP) | Mitochondrial uncoupler; alters metabolic state and autofluorescence, used as a control. |
| Rotterdam / SP8 FALCON systems | Commercial hyperspectral FLIM platforms enabling native multispectral unmixing. |
Integrated Workflow for Artifact-Corrected Redox FLIM
Integrated Workflow for Artifact Correction in FLIM
Data Processing and Validation Protocol
Protocol 6.1: Pixel-Wise Artifact Correction Calculation
Objective: To implement a consolidated correction formula for all artifacts. Input Data: For each pixel: measured biosensor lifetime (τmeas), reference pH lifetime (τpHRef), measured O2 probe lifetime (τO2), unmixing coefficient for autofluorescence (αauto). Procedure:
- Calculate pH Correction: Apply calibration function: τpHCorr = τmeas + f(τ_pHRef).
- Apply O2 Normalization: Use Stern-Volmer: τO2Corr = τpHCorr * (1 + kq[O2_ref]) / (1 + kq[O2meas]). [O2] is derived from τO2.
- Subtract Autofluorescence: τcorr = (τO2Corr - αauto * τauto) / (1 - αauto), where τauto is the predefined autofluorescence lifetime.
- Validate by treating cells with redox modulators (e.g., DTT, H2O2) and environmental perturbants (e.g., CO2 shift, hypoxia). The corrected lifetime (τ_corr) should respond only to redox modulators.
Pixel-Wise Computational Correction Pipeline
Conclusion: For robust thiol-disulfide redox research using FLIM biosensing, a systematic approach to correct for pH, oxygen, and autofluorescence is non-negotiable. The protocols outlined here provide a framework to isolate the true redox signal, enabling reliable discovery and drug development in oxidative stress-related diseases.
Managing Phototoxicity and Photobleaching in Long-Term Imaging
Within the broader thesis on Fluorescence Lifetime Imaging (FLIM) biosensing for thiol-disulfide redox research, managing photodamage is not merely a technical concern but a fundamental prerequisite for accurate biological measurement. Long-term FLIM enables dynamic quantification of cellular redox states, as the fluorescence lifetime of specific genetically encoded biosensors (e.g., roGFP, rxYFP) is sensitive to the local redox environment. However, prolonged or repeated laser excitation induces phototoxicity (cellular damage) and photobleaching (fluorophore destruction), which corrupts the very physiological processes under study—particularly redox homeostasis. These artifacts introduce significant errors in lifetime-based quantification, leading to false conclusions about oxidative stress or reductive signaling. Therefore, implementing robust strategies to minimize photodamage is critical for generating reliable, reproducible data in redox biology and subsequent drug discovery efforts targeting redox pathways.
Quantitative Data on Photodamage Factors and Mitigation Strategies
Table 1: Key Parameters Influencing Phototoxicity and Photobleaching in FLIM
| Parameter | Effect on Photobleaching | Effect on Phototoxicity | Recommended Optimization for FLIM-Redox Imaging |
|---|---|---|---|
| Excitation Intensity | Quadratic increase with intensity. | Linear to quadratic increase; disrupts redox homeostasis. | Use lowest intensity yielding sufficient SNR (often 1-10 µW at sample). |
| Exposure Time / Dwell Time | Linear increase. | Linear increase; cumulative radical generation. | Minimize; use fastest acceptable pixel dwell time (e.g., 1-10 µs). |
| Excitation Wavelength | Higher energy (shorter λ) increases risk. | Shorter λ (e.g., 405nm) more damaging than longer λ (e.g., 488nm). | Use longest λ compatible with biosensor excitation (e.g., 488nm for roGFP). |
| Repetition Rate (Pulsed Lasers) | High rate increases total dose. | High rate increases thermal stress. | Match to fluorophore lifetime; use ≤ 20-40 MHz for common FPs. |
| Numerical Aperture (NA) | Higher NA concentrates energy. | Higher NA concentrates energy, increasing local damage. | Use lowest NA objective that provides required resolution (e.g., 1.2 vs 1.49). |
| Molecular Oxygen Concentration | Essential for Type II photobleaching. | Major source of reactive oxygen species (ROS). | Consider anoxic imaging chambers for extreme long-term work (caveat: alters physiology). |
| Antioxidant / Scavenger Use | Can reduce bleaching. | Significantly reduces ROS-mediated damage. | Include in imaging medium (e.g., 1-5 mM Trolox, Ascorbic Acid). |
Table 2: Comparison of FLIM Modalities for Long-Term Redox Imaging
| FLIM Modality | Typical Excitation Source | Key Advantage for Reducing Damage | Key Limitation for Long-Term Redox Imaging |
|---|---|---|---|
| Time-Correlated Single Photon Counting (TCSPC) | Pulsed laser (e.g., 485 nm diode). | Extremely sensitive; can use very low photon flux. | Slow acquisition; risk of spatial artifacts during dynamic changes. |
| Frequency-Domain (FD-FLIM) | Intensity-modulated laser or LED. | Faster acquisition, reducing total exposure time. | Generally lower lifetime resolution than TCSPC. |
| Widefield gated/Phasor FLIM | Pulsed LED or laser illumination. | Very fast, low peak power, widefield illumination. | Lower spatial resolution; may require higher total dose. |
Experimental Protocols for Minimizing Photodamage in FLIM-Redox Experiments
Protocol 3.1: Optimizing FLIM Acquisition Parameters for Redox Biosensor Imaging (e.g., roGFP-RL12)
Objective: To establish acquisition parameters that minimize photobleaching while maintaining sufficient signal-to-noise ratio (SNR) for accurate lifetime determination of a redox biosensor. Materials: Cells expressing roGFP-RL12, FLIM system (TCSPC or FD), live-cell imaging chamber. Procedure:
- Initial Setup: Maintain cells at 37°C, 5% CO₂ in phenol-red-free medium, optionally supplemented with 1 mM Trolox.
- Find Minimal Laser Power: a. Set the pixel dwell time to a standard value (e.g., 10 µs). b. Start with the lowest possible laser power (e.g., 0.1 µW at back focal plane). c. Acquire a single FLIM image and check the photon count in the brightest cell (aim for >1000 photons per pixel for a biexponential fit). d. Increase laser power in 10-20% increments until the minimum photon count threshold is reached. Record this power.
- Optimize Dwell Time/Scan Speed: a. Using the laser power from step 2d, gradually decrease the dwell time. b. Acquire images and monitor the decay curve quality and the standard error of the fitted lifetime. c. Select the shortest dwell time that does not significantly increase the lifetime error (e.g., <5% change).
- Validate Physiological Integrity: a. Acquire a time-lapse FLIM series using the optimized parameters (e.g., 1 image every 2 minutes for 60 minutes). b. Include a positive control: After 20 minutes, add 100 µM H₂O₂ to the medium. A robust lifetime shift should be observable. c. Include a negative control: Image without any treatment. The baseline lifetime should remain stable (<5% drift). A consistent increase in lifetime indicates probe bleaching or stress.
- Finalize Parameters: Use the validated power, dwell time, and acquisition interval for all experimental sessions.
Protocol 3.2: System Calibration and Validation for Low-Dose FLIM
Objective: To ensure the FLIM system is calibrated for sensitive, low-light detection to avoid unnecessary excitation. Procedure:
- Instrument Response Function (IRF) Measurement: Daily, acquire the IRF using a scattering solution (e.g., colloidal silica) or a instantaneously decaying fluorophore under identical optical conditions used for cell imaging.
- Detector Linearity Check: Monthly, perform a count rate linearity test. Acquire FLIM data from a stable fluorescent slide at increasing laser powers. Plot detected counts vs. power. Confirm linear response in the used count rate regime (<1-2% of laser repetition rate for TCSPC).
- Background Assessment: Before each experiment, acquire an image from a non-fluorescent region of the sample. Quantify the average background counts. This value should be subtracted during data analysis.
Signaling Pathways and Workflow Diagrams
Diagram Title: Photophysics of FLIM-Redox Imaging and Photodamage Pathways
Diagram Title: Workflow for Low-Photodamage FLIM-Redox Experiments
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Managing Photodamage in Long-Term FLIM
| Item / Reagent | Function & Rationale | Example Product / Specification |
|---|---|---|
| Genetically Encoded Redox Biosensor | FLIM-compatible probe whose lifetime shifts with thiol-disulfide status. Enables measurement without ratiometric intensity artifacts prone to bleaching. | roGFP-RL12, rxYFP-FLIM. Requires stable cell line generation. |
| Phenol-Red Free Imaging Medium | Eliminates background fluorescence and phototoxicity from phenol red upon illumination. | Gibco FluoroBrite DMEM, or custom Hanks' Balanced Salt Solution (HBSS). |
| ROS Scavengers / Antioxidants | Quench reactive oxygen species generated during imaging, reducing indirect phototoxicity and protecting the biosensor. | Trolox (water-soluble vitamin E analog, 1-5 mM), Ascorbic Acid (Vitamin C, 0.5-1 mM). |
| Oxygen Scavenging System | Chemically reduces molecular oxygen (O₂) in the medium, mitigating Type II (singlet oxygen) photobleaching pathways. | ProtoCO2 (Oxyrase for cells), or Glucose Oxidase/Catalase (GOC) enzyme system. Use with caution as it creates hypoxia. |
| Live-Cell Imaging Chamber | Maintains precise physiological control (37°C, 5% CO₂, humidity) which is critical for redox homeostasis during long-term experiments. | Stage-top incubators (e.g., Tokai Hit) or micro-environment chambers (e.g., Ludin Chamber). |
| High-Sensitivity Detector | Enables the use of lower excitation intensities by maximizing photon collection efficiency. Essential for TCSPC-FLIM. | GaAsP or Hybrid PMT detectors, with high quantum efficiency (>40%) at emission wavelengths. |
| Pulsed LED or Diode Laser | Lower peak power and cost compared to Ti:Sapphire lasers, reducing nonlinear photodamage for FD- or widefield gated-FLIM. | 485 nm or 500 nm pulsed LED systems (e.g., Cairn Research) with fast modulation. |
| Immersion Oil (Low-Autofluorescence) | Reduces background signal, allowing lower excitation power. Critical for high-NA objective use. | Types specifically formulated for fluorescence imaging (e.g., Nikon ND50, Immersol 518F). |
In Fluorescence Lifetime Imaging (FLIM) biosensing for thiol-disulfide redox research, the lifetime (τ) of a genetically encoded or chemically conjugated probe (e.g., roGFP, rxYFP) serves as a quantitative readout of cellular redox potential. A shift in lifetime is interpreted as a change in the probe's microenvironment due to oxidation or reduction. However, the apparent lifetime is influenced by numerous factors beyond redox state. Misinterpretation of these shifts can lead to false conclusions about redox biology, compromising drug discovery efforts targeting oxidative stress. This application note details critical pitfalls and protocols for robust data acquisition and analysis.
Core Pitfalls & Validating the Lifetime Shift
Key non-redox factors causing lifetime shifts are summarized in Table 1.
Table 1: Common Non-Redox Factors Affecting FLIM Measurements in Redox Biosensing
| Factor | Mechanism of Impact on τ | Confounding Signal Mimicry |
|---|---|---|
| pH Variance | Protonation alters probe's electronic state. | Acidic pH can mimic an oxidized state for some probes. |
| Ionic Strength | Affects local dielectric constant and quenching. | High [Cl⁻] can shorten τ, obscuring a true reduction event. |
| Macromolecular Crowding | Alters rotational diffusion & collisional quenching. | Increased crowding may shorten τ, independent of redox. |
| Temperature Fluctuations | Changes non-radiative decay rates. | Temperature drop can increase τ, mimicking reduction. |
| Probe Concentration (Affinity) | Can enable homo-FRET or aggregation. | High local concentration may cause τ shortening. |
| Photobleaching | Creates irreversible, non-fluorescent species. | Alters biexponential fit, producing artifactual τ shifts. |
| Cellular Autofluorescence | Introduces background with distinct τ. | If unaccounted for, biases fit towards autofluorescence τ. |
Experimental Protocols for Pitfall Mitigation
Protocol 1: System Calibration & Environmental Control
Objective: Isolate redox-dependent lifetime changes from environmental artifacts.
- In vitro Calibration: Perform a full redox titration (using DTT/H₂O₂ or glutathione redox buffers) of the probe under controlled conditions.
- Prepare 10+ buffers spanning defined redox potentials (e.g., -320 mV to -220 mV).
- Acquire FLIM data for each buffer at precisely controlled temperature (e.g., 37°C ± 0.2°C), pH (e.g., 7.4 ± 0.05), and ionic strength.
- Generate a reference table of τ vs. Eh. This is your ground-truth calibration curve.
- Intracellular Environmental Mapping: In parallel experiments, use control biosensors (e.g., pH-insensitive redox probe mutants, Cl⁻ sensors) to map intracellular pH and ion concentrations under your experimental conditions.
- Cross-Reference: Compare cellular τ shifts to your in vitro calibration curves, correcting for measured intracellular pH/ion variations.
Protocol 2: Phasor Plot Analysis for Multi-Exponential Validation
Objective: Detect and separate multiple lifetime populations (e.g., probe in different states, background) without a priori fitting models.
- Data Transformation: Transform time-domain or frequency-domain FLIM data into phasor coordinates (G, S) for every pixel.
- G = (∫ I(t) cos(ωt) dt) / (∫ I(t) dt)
- S = (∫ I(t) sin(ωt) dt) / (∫ I(t) dt)
- Where ω is the laser repetition angular frequency.
- Population Identification: Plot all phasors. A single exponential lifetime falls on the "universal semicircle." Complex decays fall inside it.
- Artifact Flagging: Identify clusters corresponding to:
- Pure probe states (oxidized/reduced): Should align along the "redox trajectory" defined by in vitro calibration points.
- Autofluorescence: Will appear as a distinct cluster. Mask and exclude these pixels.
- Photobleached species: Appear as a time-dependent drift of the cluster position.
- Selective Gating: Apply a graphical gate to isolate pixels corresponding only to the healthy probe population for subsequent τ analysis.
Protocol 3: Lifetime-Targeted Control for Expression & Crowding
Objective: Control for artifacts from variable probe expression and local crowding.
- Co-express an inert τ reference: Use a redox-insensitive fluorescent protein (e.g., GFP-S65T) with a known, stable lifetime as an internal transfection/maturation control.
- Dual-Channel FLIM Acquisition: Acquire FLIM simultaneously or sequentially for the redox biosensor (Channel 1) and the inert reference (Channel 2).
- Ratio-metric Lifetime Analysis: Calculate a pixel-wise ratio: τ(biosensor) / τ(reference). This normalized "pseudoratio" helps cancel out global effects affecting both probes (e.g., temperature, scattering, some crowding effects), highlighting changes specific to the redox probe's microenvironment.
Visualization of Analysis Workflow & Pathways
Title: FLIM Redox Data Validation Workflow
Title: Pathways to Real vs. Artifactual Lifetime Shifts
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Robust FLIM Redox Biosensing
| Reagent / Material | Function & Rationale |
|---|---|
| Genetically Encoded Redox Biosensors (e.g., roGFP2, rxYFP, Grx1-roGFP2) | Target-specific probe. Grx1-roGFP2 offers direct coupling to the glutathione pool. |
| Redox Buffer Kits (Glutathione or DTT/H₂O₂ based) | For in vitro calibration curves. Allow precise definition of solution Eh. |
| Inert Lifetime Reference Probe (e.g., GFP-S65T, mCherry) | Internal control for Protocol 3. Must have spectrally separable emission from biosensor. |
| Environmental Control Sensors (pHluorin, Cl⁻ sensors) | To quantify and correct for parallel changes in pH and chloride concentration. |
| Two-Photon FLIM-Compatible Live-Cell Imaging Medium | Phenol-red free, HEPES-buffered medium to maintain pH without CO₂, minimizing optical interference. |
| Specific Pharmacological Agents (e.g., Paraquat, Menadione, N-Acetylcysteine) | Well-characterized redox modulators for positive and negative experimental controls. |
| Quenching Agents (e.g., Potassium Iodide) | For collision quenching experiments to probe solvent accessibility changes in the biosensor. |
Best Practices for Sample Preparation and Control Experiments
Fluorescence Lifetime Imaging Microscopy (FLIM) is a powerful quantitative technique for investigating cellular thiol-disulfide redox states, a crucial aspect of cellular signaling, oxidative stress, and drug mechanisms. Accurate and reproducible FLIM data is highly dependent on rigorous sample preparation and the implementation of robust control experiments. This protocol details best practices tailored for redox biosensing using genetically encoded probes like roGFP or rxYFP, whose fluorescence lifetime is sensitive to local redox potential.
Critical Sample Preparation Protocols
Cell Culture and Transfection for FLIM Biosensing
Objective: To prepare living cells expressing the redox biosensor with high viability and minimal inherent oxidative stress. Detailed Protocol:
- Cell Line Selection: Use low-passage-number cells (< passage 20). HeLa, HEK293, or primary cells relevant to the disease model are common.
- Transfection: Transfect with the plasmid encoding the redox biosensor (e.g., roGFP2-Orp1) using a low-cytotoxicity method (e.g., lipid-based transfection, electroporation optimized for >70% viability).
- Expression Timing: Perform FLIM experiments 24-48 hours post-transfection. Avoid prolonged expression (>72h) to prevent artifacts from overexpression.
- Seeding for Imaging: Seed cells onto #1.5 high-precision glass-bottom dishes 24 hours before imaging at a confluency of 50-60%.
- Serum Starvation: If required by the experiment, reduce serum concentration to 0.5-1% for 4-16 hours prior to imaging to minimize basal growth factor-induced signaling.
Preparation of Redox-Modulating Solutions
Objective: To create precise buffers for calibration and experimental perturbation. Detailed Protocol for Calibration Buffers:
- Base Imaging Buffer: 135 mM NaCl, 5 mM KCl, 1 mM MgCl₂, 2 mM CaCl₂, 10 mM Glucose, 10 mM HEPES, pH 7.4.
- Fully Oxidizing Buffer: Base buffer + 10 mM Diamide (a thiol-oxidizing agent). Prepare fresh.
- Fully Reducing Buffer: Base buffer + 10 mM Dithiothreitol (DTT). Prepare fresh.
- Physiological Buffer Control: Base buffer + 1-10 µM Cysteine/Cystine to mimic extracellular redox couples. Note: Use buffers pre-warmed to 37°C and equilibrated in a cell culture incubator with 5% CO₂ for at least 30 minutes before use.
Live-Cell Imaging Preparation
Objective: To minimize environmental stress during FLIM acquisition. Detailed Protocol:
- Phenol Red-Free Media: Replace growth media with pre-warmed, phenol-red-free imaging buffer 30 minutes before measurement.
- Environmental Control: Maintain the stage at 37°C using a heated stage incubator with precise temperature feedback control (±0.5°C). Maintain 5% CO₂ if using bicarbonate buffer.
- Anti-Photobleaching: Add an oxygen-scavenging system (e.g., Oxyrase, 0.3 units/mL) to reduce photobleaching and associated oxidative stress during long acquisitions.
Essential Control Experiments and Protocols
In-Situ Sensor Calibration Protocol
Objective: To determine the dynamic range (τₘᵢₙ, τₘₐₓ) of the FLIM biosensor within the cellular environment. Detailed Protocol:
- Image cells in physiological buffer and acquire FLIM data for baseline lifetime (τᵦₐₛₑₗᵢₙₑ).
- Gently replace buffer with fully oxidizing buffer (10 mM Diamide). Incubate for 5 minutes. Acquire FLIM data (τₒₓ).
- Gently wash cells 3x with base imaging buffer.
- Gently replace buffer with fully reducing buffer (10 mM DTT). Incubate for 5 minutes. Acquire FLIM data (τᵣₑd).
- Data Analysis: Calculate the redox ratio (R) for each pixel or cell: R = (τₘₑₐₛᵤᵣₑd - τᵣₑd) / (τₒₓ - τᵣₑd). This normalizes lifetime changes to the sensor's dynamic range.
Specificity and Viability Controls
Objective: To confirm observed lifetime changes are due to specific thiol-disulfide redox changes and not artifacts. Detailed Protocols:
- Sensorless Control: Perform identical treatments on non-transfected cells. Acquire FLIM at identical laser power/detection settings to measure autofluorescence background, which must be negligible (<5% of sensor signal).
- Pharmacological Inhibition Control: Pre-treat cells with a specific antioxidant (e.g., 5 mM N-acetylcysteine (NAC) for 1 hour) or pro-oxidant (e.g., 100 µM H₂O₂ for 10 min) before the experimental stimulus. This confirms the sensor's response is coupled to the relevant redox pathways.
- Cell Viability Control: Co-stain cells with a viability marker (e.g., 1 µg/mL Propidium Iodide) after FLIM acquisition. Correlate FLIM data from PI-negative (live) cells only.
Technical Controls for FLIM Instrumentation
Objective: To ensure instrument stability and data validity. Detailed Protocols:
- Reference Standard Measurement: Daily, measure the fluorescence lifetime of a stable fluorophore with a known lifetime (e.g., Fluorescein in 0.1 M NaOH, τ ≈ 4.0 ns). Record values; drift >±50 ps requires system realignment.
- "No-Laser" Control: Acquire a measurement with the laser beam blocked to assess electronic/dark noise of the detection system.
- Instrument Response Function (IRF) Measurement: Acquire the IRF weekly using a scattering solution (e.g., Ludox) or a instantly decaying dye. This is critical for accurate lifetime fitting, especially for sub-nanosecond components.
Data Presentation: Quantitative Reference Tables
Table 1: Typical FLIM Parameters for Common Redox Biosensors
| Biosensor | Excitation (nm) | Emission (nm) | Lifetime Range (Reduced, ns) | Lifetime Range (Oxidized, ns) | Key Control Experiment |
|---|---|---|---|---|---|
| roGFP2 | 488 | 500-550 | 2.6 - 2.8 | 2.1 - 2.3 | In-situ diamide/DTT calibration |
| rxYFP | 514 | 525-575 | 2.7 - 2.9 | 2.0 - 2.2 | Glutaredoxin-1 coupling validation |
| HyPer | 488/405 | 500-550 | 2.9 / 2.8* | 2.5 / 3.1* | H₂O₂ specificity control (Catalase) |
| Grx1-roGFP2 | 488 | 500-550 | 2.6 - 2.8 | 2.1 - 2.3 | Glutathione redox potential calibration |
*Lifetime is excitation-ratio dependent for HyPer.
Table 2: Common Redox Perturbation Agents & Recommended Controls
| Agent | Target/Effect | Working Concentration | Critical Control Experiment |
|---|---|---|---|
| Diamide | Thiol oxidizer | 1-10 mM | Reversibility check with subsequent DTT |
| DTT | Thiol reductant | 1-10 mM | Check for cellular acidification artifact |
| H₂O₂ | Physiological oxidant | 10-500 µM | Co-treatment with PEG-Catalase |
| Menadione | ROS generator (Complex II) | 10-100 µM | Viability assay (MTT/PI) parallel run |
| BSO (Buthionine sulfoximine) | Depletes GSH | 0.1-1 mM for 24h | Measure total glutathione levels biochemically |
| NAC | Increases GSH | 1-5 mM for 1-24h | Confirm increased GSH via independent assay |
Visualizing Workflows and Pathways
Diagram 1: FLIM Redox Experiment Core Workflow
Diagram 2: Thiol Redox Signaling & Biosensor Coupling
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function & Rationale | Example Product/Catalog # |
|---|---|---|
| Genetically Encoded Redox Sensor | Directly reports cellular thiol-disulfide state via fluorescence lifetime. | pCX-roGFP2-Orp1 (Addgene #64993) |
| Thiol-Oxidizing Agent (Diamide) | Induces complete, rapid oxidation of sensor for in-situ calibration. | Diamide, Sigma-Aldrich (D3648) |
| Thiol-Reducing Agent (DTT) | Induces complete reduction of sensor for in-situ calibration. | Dithiothreitol, Thermo Fisher (R0861) |
| Phenol Red-Free Imaging Medium | Eliminates background fluorescence from culture media. | FluoroBrite DMEM, Gibco (A1896701) |
| Live-Cell Imaging-Optimized Dish | #1.5 glass ensures optimal optical resolution for high-NA objectives. | MatTek Dish (P35G-1.5-14-C) |
| Stage Top Incubator | Precisely maintains 37°C and 5% CO₂ for physiological measurements. | Tokai Hit (STX) or Okolab (H301) |
| Oxygen-Scavenging System | Reduces photobleaching and phototoxicity during prolonged FLIM. | Oxyrase (OB-0000) |
| Fluorescence Lifetime Reference | Validates daily instrument performance and calibration. | Fluorescein (0.1M NaOH), Sigma (F6377) |
| Cell Viability Stain | Post-FLIM viability assessment to exclude dead cells from analysis. | Propidium Iodide, Thermo Fisher (P3566) |
Benchmarking FLIM: Validation Against Gold Standards and Comparative Advantages
Correlating FLIM Data with Biochemical Assays (e.g., HPLC, Ellman's Assay)
Within the broader thesis on FLIM biosensing for thiol-disulfide redox research, this application note details the critical integration of Fluorescence Lifetime Imaging (FLIM) with established biochemical endpoint assays. FLIM provides spatially resolved, quantitative maps of cellular redox states using biosensors like roGFP or Grx1-roGFP, which are sensitive to glutathione redox potential (Eh). However, validation and absolute quantification require correlation with bulk biochemical methods such as High-Performance Liquid Chromatography (HPLC) for glutathione quantification and Ellman's Assay for total free thiol determination. This protocol outlines a workflow to perform these techniques in parallel on matched samples, enabling the calibration of FLIM lifetime readings to concrete biochemical concentrations.
Key Quantitative Correlations
The following table summarizes typical data from correlated studies, demonstrating the relationship between FLIM-derived parameters and biochemical measurements.
Table 1: Correlation of FLIM Parameters with Biochemical Assay Data in Redox Studies
| Cell Line / Condition | FLIM Mean Lifetime (τ, ns) (roGFP biosensor) | HPLC [GSH] (nmol/mg protein) | HPLC [GSSG] (nmol/mg protein) | Ellman's Assay (Total Thiols, nmol/mg protein) | Calculated Eh (mV) |
|---|---|---|---|---|---|
| Control (HEK293) | 2.65 ± 0.10 | 25.4 ± 2.1 | 1.8 ± 0.3 | 28.9 ± 2.5 | -315 ± 10 |
| Oxidized (1 mM H2O2, 30 min) | 2.15 ± 0.08 | 12.1 ± 1.5 | 8.7 ± 1.1 | 15.3 ± 1.8 | -250 ± 8 |
| Reduced (10 mM DTT, 30 min) | 2.95 ± 0.12 | 38.7 ± 3.0 | 0.5 ± 0.2 | 40.5 ± 3.2 | -345 ± 12 |
Experimental Protocols
Protocol 1: FLIM of Live Cells Expressing Redox Biosensors
Objective: To acquire fluorescence lifetime data of cells expressing a genetically encoded redox biosensor (e.g., Grx1-roGFP2).
Materials:
- Cells expressing Grx1-roGFP2
- Imaging medium (e.g., FluoroBrite DMEM, no phenol red)
- Oxidizing/Reducing controls (e.g., 1 mM H2O2, 10 mM DTT)
- Confocal microscope with time-correlated single-photon counting (TCSPC) FLIM capability
Method:
- Seed cells on 35mm glass-bottom dishes and culture for 24-48 hrs.
- Replace medium with pre-warmed imaging medium.
- For calibration, treat control dishes with H2O2 (oxidized state) or DTT (reduced state) for 30 min.
- Mount dish on microscope stage maintained at 37°C/5% CO2.
- FLIM Acquisition:
- Excite roGFP at 488 nm using a pulsed laser (e.g., 40 MHz repetition rate).
- Collect emission using a bandpass filter (500-550 nm).
- Acquire images until peak photon count in the brightest pixel reaches ~10,000 counts to ensure good lifetime fitting.
- Use a 405 nm excitation scan to perform the ratiometric calibration of the biosensor if required.
- FLIM Analysis:
- Fit the fluorescence decay at each pixel to a double-exponential model using dedicated software (e.g., SPCImage, FLIMfit).
- Calculate the mean fluorescence lifetime (τm) as τm = (a1τ1 + a2τ2) / (a1 + a2), where a are amplitudes.
- Generate pseudocolor lifetime maps and extract average τm values for regions of interest (whole cells).
Protocol 2: HPLC Analysis of Glutathione (GSH/GSSG)
Objective: To quantitatively determine reduced (GSH) and oxidized (GSSG) glutathione levels from the same cell population used for FLIM trends.
Materials:
- Cell pellet from matched culture conditions (≥ 1x106 cells)
- Lysis buffer: 0.1% Triton X-100, 0.1 M Phosphate-EDTA buffer, pH 7.5
- 10% Metaphosphoric acid (MPA)
- Derivatization agent: 10 mM O-phthalaldehyde (OPA) in methanol
- HPLC system with fluorescence detector
Method:
- Sample Preparation: Lyse cell pellet in ice-cold lysis buffer. For total glutathione (GSH+GSSG), use half the lysate. For GSSG-specific measurement, pre-incubate the other half of lysate with 2-vinylpyridine (2% v/v) for 1 hr to derivative GSH.
- Protein Precipitation: Mix lysate with an equal volume of cold 10% MPA. Centrifuge at 13,000 x g for 10 min at 4°C. Collect supernatant.
- Derivatization: Mix 50 µL of supernatant with 50 µL OPA. Incubate at room temperature for 2 min.
- HPLC Analysis:
- Column: C18 reverse-phase column (e.g., 4.6 x 150 mm, 5 µm).
- Mobile Phase A: 0.1% (v/v) trifluoroacetic acid in water. Phase B: 100% methanol.
- Gradient: 10% B to 90% B over 15 min.
- Flow rate: 1 mL/min. Fluorescence detection: Ex 340 nm / Em 420 nm.
- Quantification: Calculate GSH and GSSG concentrations from standard curves. Normalize to total protein content of the initial lysate.
Protocol 3: Ellman's Assay for Total Free Thiols
Objective: To measure the total concentration of free sulfhydryl groups as a complementary redox metric.
Materials:
- Cell lysate (in PBS, pH 7.4)
- Ellman's Reagent: 5,5'-Dithio-bis-(2-nitrobenzoic acid) (DTNB, 4 mg/mL in PBS)
- L-Cysteine standard solution (0-100 µM)
Method:
- Prepare a standard curve of L-cysteine (0, 10, 25, 50, 75, 100 µM) in PBS.
- Mix 50 µL of cell lysate (or standard) with 150 µL of PBS in a 96-well plate.
- Add 10 µL of DTNB reagent to each well. Incubate at room temperature for 15 min protected from light.
- Measure the absorbance at 412 nm using a plate reader.
- Calculate the total thiol concentration from the cysteine standard curve. Normalize to total protein content.
Visualization of Workflow and Signaling
Title: Integrated FLIM and Biochemical Assay Workflow for Redox Research
Title: roGFP Redox Sensing and FLIM-Biochemical Correlation
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Correlated FLIM-Biochemical Redox Studies
| Reagent / Material | Function / Role in Experiment | Key Notes |
|---|---|---|
| Grx1-roGFP2 Plasmid | Genetically encoded biosensor; fusion of human glutaredoxin-1 and redox-sensitive GFP for rapid, specific equilibration with the glutathione pool. | Enables specific measurement of glutathione Eh in live cells via FLIM. |
| Metaphosphoric Acid (MPA) | Protein precipitant and acidifying agent for HPLC sample prep. Preserves thiols from auto-oxidation during processing. | Critical for accurate GSH/GSSG ratio measurement; must be freshly prepared or stored at -20°C. |
| O-Phthalaldehyde (OPA) | Fluorescent derivatization agent for primary amines and thiols. Forms adducts with GSH for sensitive HPLC detection. | Prepare fresh in methanol; stable for ~1 week at 4°C in the dark. |
| 5,5'-Dithio-bis-(2-nitrobenzoic acid) (DTNB) | Ellman's Reagent. Reacts with free thiol groups to produce 2-nitro-5-thiobenzoate (TNB²⁻), a yellow chromophore. | Standard method for total free thiol quantification; absorbance read at 412 nm. |
| 2-Vinylpyridine | Thiol-scavenging agent used to derivative GSH in samples for specific measurement of GSSG by HPLC. | Must be used in a fume hood. Allows measurement of the low-abundance GSSG pool without interference. |
| TCSPC FLIM Module | Instrumentation (e.g., Becker & Hickl, PicoQuant) attached to confocal microscope. Measures nanosecond fluorescence decay kinetics. | Essential for acquiring lifetime data (τ); requires pulsed laser and fast detectors. |
| C18 Reverse-Phase HPLC Column | Stationary phase for separation of OPA-derivatized GSH and GSSG based on hydrophobicity. | Standard column for glutathione analysis; requires equilibration and regular cleaning. |
Application Notes
Within the context of advancing FLIM biosensing for thiol-disulfide redox research, choosing the optimal imaging modality is critical for accurate quantification of dynamic biochemical states. This analysis compares Fluorescence Lifetime Imaging (FLIM) and Intensity-Based Ratiometric Imaging (IBRI) for sensing redox potentials, primarily using genetically encoded biosensors like roGFP or rxYFP.
Core Principle & Sensitivity: FLIM measures the exponential decay time of fluorescence after excitation, a parameter intrinsically independent of probe concentration, excitation intensity, and light path artifacts. It is exquisitely sensitive to the molecular microenvironment, including Förster Resonance Energy Transfer (FRET) efficiency, which changes with biosensor conformation (e.g., reduced vs. oxidized). In contrast, IBRI relies on the ratio of fluorescence intensities at two excitation or emission wavelengths. While it corrects for some technical variables, it remains vulnerable to photobleaching, scattering, and autofluorescence, which can distort ratios.
Quantification Advantages for Redox Research: For thiol-disulfide redox sensing, the reduced and oxidized states of a biosensor often exhibit distinct lifetimes. FLIM provides an absolute, quantitative measure (nanoseconds) that can directly report on the fractional population of each state within a pixel, enabling precise determination of redox potential. IBRI provides a unitless ratio that must be calibrated against fully reduced and oxidized states for each experiment. FLIM's independence from sensor concentration is particularly advantageous in heterogeneous cellular environments or over long-term experiments where expression levels may vary.
Key Application Insight: Recent studies highlight FLIM's superior sensitivity in detecting small changes in FRET efficiency, making it the method of choice for quantifying subtle redox shifts in subcellular compartments, such as mitochondrial matrix versus cytosol, or for high-content screening where artifact minimization is paramount. IBRI remains widely used due to its speed and instrumental simplicity but requires stringent controls for quantitative rigor.
Quantitative Data Comparison
Table 1: Comparative Metrics of FLIM and IBRI for Redox Biosensing
| Metric | Intensity-Based Ratiometric Imaging (IBRI) | Fluorescence Lifetime Imaging (FLIM) |
|---|---|---|
| Primary Readout | Intensity Ratio (e.g., I₄₈₈/I₄₀₅ for roGFP2) | Fluorescence Lifetime (τ, in nanoseconds) |
| Concentration Dependence | Independent (in theory) | Fully Independent |
| Excitation Intensity Dependence | Low (corrected by ratio) | None |
| Photobleaching Sensitivity | High (can distort ratio) | Low (lifetime often bleaches slower than intensity) |
| Temporal Resolution | High (ms to s) | Moderate to Low (seconds to minutes) |
| Typical Dynamic Range (Δ) | ~5-10 fold change in ratio | ~1.5-2.5 fold change in lifetime (e.g., 2.8 ns to 1.6 ns) |
| Quantitative Output | Relative ratio; requires in situ calibration | Absolute τ; can calculate fraction of species directly |
| Key Vulnerability | Autofluorescence, inner filter effects, scattering | Complex/expensive instrumentation, data fitting complexity |
Table 2: Example Data from a Redox Biosensor (Hypothetical rxYFP)
| Redox State | IBRI Ratio (Ex405/Ex488) | FLIM Average Lifetime (τ, ns) | Estimated FRET Efficiency |
|---|---|---|---|
| Fully Reduced (DTT) | 0.2 ± 0.05 | 3.10 ± 0.15 | ~0% |
| Fully Oxidized (H₂O₂) | 1.8 ± 0.1 | 1.65 ± 0.10 | ~45% |
| Cellular Cytosol (Resting) | 0.6 ± 0.15 | 2.50 ± 0.20 | ~20% |
Experimental Protocols
Protocol 1: Intensity-Based Ratiometric Imaging of roGFP2 for Cellular Redox State Objective: To quantify the glutathione redox potential (E_GSH) in live cells.
- Cell Preparation: Seed cells expressing roGFP2 targeted to your compartment of interest (e.g., roGFP2-Mito) in an imaging chamber.
- Microscope Setup: Use a confocal or widefield microscope capable of sequential excitation at 405 nm and 488 nm. Collect emission between 500-540 nm.
- Calibration Imaging: a. Acquire a baseline image set (Ex405, Em; Ex488, Em). b. Perfuse with calibration buffer containing 10 mM DTT (reducing agent) for 15 min. Acquire image set (Fully Reduced signal). c. Wash and perfuse with calibration buffer containing 10 mM H₂O₂ (oxidizing agent) for 15 min. Acquire image set (Fully Oxidized signal).
- Experimental Imaging: Acquire ratio images (I₄₀₅ / I₄₈₈) over time during treatment.
- Data Analysis: Calculate the normalized oxidation degree: (R - Rred) / (Rox - Rred), where R is the measured ratio. Convert to EGSH using the Nernst equation.
Protocol 2: FLIM-FRET Imaging of a Redox Biosensor (e.g., Grx1-roGFP2) Objective: To measure fluorescence lifetime changes reporting on thiol-disulfide status via FRET.
- Sample Preparation: Seed cells expressing the FRET-based redox biosensor.
- FLIM System Setup: Use a time-correlated single-photon counting (TCSPC) FLIM system coupled to a multiphoton or confocal microscope. Set excitation to 920 nm (for two-photon) or 488 nm. Set emission filter to collect donor emission (e.g., 500-550 nm for roGFP2).
- Lifetime Calibration: Measure a reference standard with a known, single-exponential lifetime (e.g., fluorescein at known pH).
- Data Acquisition: Acquire photon count images until sufficient counts per pixel are reached (typically 1000-2000) for a reliable fit. Maintain low laser power to avoid photobleaching.
- Lifetime Analysis: Fit the fluorescence decay curve per pixel using a bi-exponential model: I(t) = α₁ exp(-t/τ₁) + α₂ exp(-t/τ₂), where τ₁ and τ₂ represent lifetimes of the reduced (donor-only) and oxidized (FRETing) populations. The amplitude-weighted average lifetime τ_avg = (α₁τ₁ + α₂τ₂) / (α₁ + α₂).
- Quantification: Plot τ_avg or the fractional amplitude of the short-lifetime component (α₂) as a map of redox state. Correlate lifetime shifts with biological perturbations.
Visualization
Title: Comparison of IBRI and FLIM-FRET Redox Sensing Workflows
Title: FLIM Data Processing and Parameter Extraction
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for FLIM & Ratiometric Redox Imaging
| Item / Reagent | Function / Explanation | Example/Catalog Consideration |
|---|---|---|
| Genetically Encoded Redox Biosensor | Reports thiol-disulfide status via fluorescence changes. | roGFP2 (general redox), rxYFP (specific to H₂O₂), Grx1-roGFP2 (for glutathione redox). |
| Live-Cell Imaging Chamber | Maintains cells at 37°C, 5% CO₂ during time-lapse imaging. | Stage-top incubators or perfusion chambers with environmental control. |
| Calibration Reagents | Define minimum (reduced) and maximum (oxidized) sensor response. | DTT (10mM): Strong reducing agent. H₂O₂ (1-10mM): Direct oxidant. Alamethicin (μg/mL): Permeabilizes membranes for calibration. |
| FLIM Reference Standard | Validates instrument performance and fitting accuracy. | Fluorescein (pH 9): τ ~4.0 ns. Rose Bengal: τ ~0.8 ns. |
| TCSPC FLIM System | Instrumentation for lifetime measurement. | Includes: Pulsed laser (e.g., Ti:Sapphire, picosecond diode), PMT detectors, timing electronics (SPC module). |
| Ratiometric Imaging Software | For sequential acquisition, ratio calculation, and calibration. | Microscope vendor software (e.g., ZEN, NIS-Elements) or open-source (ImageJ/Fiji with Ratio Plus plugin). |
| Lifetime Analysis Software | Fits exponential decays to photon arrival data. | SPCImage, SymPhoTime, FLIMfit, or open-source alternatives like FLIMJ. |
Comparative Analysis with Other Optical Techniques (BRET, Phosphorescence)
This application note is framed within a thesis exploring Fluorescence Lifetime Imaging Microscopy (FLIM) as a biosensing platform for thiol-disulfide redox dynamics in live cells. Understanding redox balance is crucial in drug development for diseases like cancer and neurodegeneration. While FLIM offers unique advantages, its practical utility is best understood through comparison with established techniques such as Bioluminescence Resonance Energy Transfer (BRET) and phosphorescence-based oxygen sensing. This document provides a comparative analysis, detailed protocols, and a toolkit to guide researchers in selecting and implementing the optimal method for their redox studies.
Comparative Quantitative Analysis
Table 1: Comparative Analysis of Optical Techniques for Redox Biosensing
| Feature | FLIM (e.g., roGFP-based) | BRET (e.g., Redox-Sensitive Luciferase) | Phosphorescence Quenching (O2 Sensing) |
|---|---|---|---|
| Primary Readout | Fluorescence lifetime (τ), ns | Ratio of acceptor/donor emission intensity | Phosphorescence lifetime/intensity |
| Excitation Source | Pulsed laser (e.g., 405 nm, 488 nm) | Substrate (e.g., Coelenterazine h) | Pulsed LED/laser (e.g., 520 nm) |
| Key Measurand | Molecular environment, redox state via sensor oxidation | Protein-protein interaction/conformational change | Local oxygen concentration ([O₂]) |
| Quantitative Output | τ (2-4 ns range), redox ratio (τox/τred) | BRET ratio (e.g., 535 nm/470 nm) | τ (10-100 µs), [O₂] in µM or mmHg |
| Spatial Resolution | High (confocal/multiphoton) | Low to moderate (typically plate reader) | Moderate (microscopy possible) |
| Temporal Resolution | Moderate (seconds-minutes for imaging) | High (milliseconds-seconds, plate-based) | High (milliseconds) |
| Key Advantage | Ratiometric, insensitive to concentration/expression, provides spatial maps | No external light, low autofluorescence, high temporal resolution | Direct, quantitative measure of metabolic activity (O₂ consumption) |
| Key Limitation | Instrumentally complex, slower acquisition | Requires substrate addition, lower spatial resolution, potential substrate toxicity | Probe can be phototoxic, requires calibration, indirect redox readout |
| Best for Redox Research | Spatially resolved, ratiometric measurement of specific redox couples (e.g., GSH/GSSG) in subcellular compartments. | High-throughput screening of redox-modulating compounds or monitoring kinetics of global redox changes in populations. | Monitoring mitochondrial respiration, hypoxia, and cellular metabolic status as an indirect redox indicator. |
Detailed Experimental Protocols
Protocol 1: FLIM for roGFP2 Redox Sensing in Live Cells
Objective: To quantify the glutathione redox potential (Eh) in the cytosol using the rationetric redox sensor roGFP2 via FLIM.
Materials: See "Research Reagent Solutions" below. Workflow:
- Cell Preparation: Seed HeLa cells expressing roGFP2 targeted to the desired compartment (e.g., cytosol) in a glass-bottom dish.
- Calibration: 24h post-seeding, perform a two-point calibration.
- Full Oxidation: Treat cells with 2 mM diamide in HBSS for 5 min.
- Full Reduction: Treat cells with 10 mM dithiothreitol (DTT) in HBSS for 5 min.
- FLIM Acquisition:
- Use a confocal or multiphoton microscope equipped with a TCSPC module.
- Excite roGFP2 at 900 nm (2P) or 488 nm (confocal) using a pulsed laser (e.g., 40 MHz repetition rate).
- Collect emission using a bandpass filter (500-550 nm).
- Acquire images until peak photon count reaches 10^3-10^4 for reliable lifetime fitting.
- Lifetime Analysis:
- Fit fluorescence decay curves per pixel to a double-exponential model: I(t) = α₁exp(-t/τ₁) + α₂exp(-t/τ₂).
- Calculate the amplitude-weighted mean lifetime: τm = (α₁τ₁ + α₂τ₂) / (α₁ + α₂).
- Compute the redox ratio: R = (τm - τred) / (τox - τred), where τox and τred are from calibration steps.
- Data Interpretation: Map R values to Eh using the Nernst equation and known E0 for roGFP2.
Protocol 2: BRET Assay for Real-Time Redox Sensing Using a Redox-Sensitive Luciferase
Objective: To monitor global cellular redox changes in real-time using a BRET-based redox sensor (e.g., Luciferase-Nanoluc with redox-sensitive YFP).
Workflow:
- Plate Preparation: Seed cells expressing the redox-sensitive BRET construct in a white-walled, clear-bottom 96-well plate.
- Substrate Addition: Dilute the cell-permeable luciferase substrate (e.g., Furimazine for Nanoluc) in pre-warmed assay medium. Add to cells to a final concentration of 5-10 µM.
- Kinetic Reading:
- Immediately place plate in a plate reader equipped with dual-emission detectors.
- Measure donor emission (e.g., 460 nm ± 20) and acceptor emission (e.g., 535 nm ± 20) sequentially every 2-10 seconds for the desired period.
- Treatment: After establishing a baseline (5-10 reads), inject a test compound (redox modulator, drug candidate) using the plate reader's injector.
- Data Processing:
- Calculate the BRET ratio for each time point: BRET = (Acceptor Emission / Donor Emission).
- Normalize data as ΔBRET/Baseline or fold change over control.
- Plot BRET ratio versus time to visualize redox dynamics.
Visualizations
Diagram 1: FLIM vs. BRET Redox Sensing Pathways
Diagram 2: Integrated Redox Experiment Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for FLIM-based Redox Biosensing
| Item | Function & Rationale |
|---|---|
| Genetically Encoded Sensor (e.g., roGFP2, Grx1-roGFP2) | Ratiometric, reversible redox probe. Targeted to specific organelles (cytosol, mitochondria, ER). Provides specific readout of the glutathione redox couple. |
| Live-Cell Imaging Medium (e.g., FluoroBrite DMEM, HBSS with HEPES) | Phenol-red free to minimize background fluorescence. Contains buffering agents for stable pH during time-lapse imaging. |
| Redox Calibrators (Diamide, DTT, H₂O₂) | Used for two-point in situ calibration to define 100% oxidized and reduced states of the sensor, enabling quantitative Eh calculation. |
| TCSPC FLIM System | Instrumentation comprising a pulsed laser, fast detector, and timing electronics. Essential for precise measurement of nanosecond fluorescence lifetimes. |
| FLIM Analysis Software (e.g., SPCImage, FLIMfit, τ-SPARK) | Specialized software to fit photon decay histograms to exponential models, calculate lifetime parameters, and generate lifetime maps. |
| Mitochondrial Uncoupler (FCCP) & Inhibitor (Antimycin A) | Pharmacological tools to perturb mitochondrial respiration, thereby altering ROS production and redox state for functional validation. |
Fluorescence Lifetime Imaging Microscopy (FLIM) biosensing of genetically encoded probes, such as roGFP and rxYFP, provides a quantitative, compartment-specific readout of cellular thiol-disulfide redox states. This document presents application notes and protocols for validating FLIM redox findings within published preclinical disease models, contextualized within the broader thesis that FLIM is a critical tool for dissecting redox pathophysiology and therapeutic mechanisms.
Application Note 1: Neurodegenerative Disease Model (Alzheimer’s Disease)
Study Context: Validation of neuronal oxidative stress and glutathione (GSH) redox potential (Eh) shifts in APP/PS1 transgenic mouse models. FLIM Probe: roGFP2 targeted to the neuronal cytosol and mitochondrial matrix. Key Finding: A significant oxidative shift (more positive Eh) in the mitochondrial matrix precedes amyloid plaque deposition, implicating mitochondrial redox dysregulation as an early pathogenic event.
Table 1: FLIM-roGFP Redox Metrics in APP/PS1 Model Hippocampal Neurons
| Cellular Compartment | Probe | Wild-Type Eh (mV) | APP/PS1 Eh (mV) | ΔEh (Oxidative Shift) | p-value |
|---|---|---|---|---|---|
| Cytosol | roGFP2 | -299 ± 5 | -287 ± 6 | +12 mV | <0.01 |
| Mitochondrial Matrix | roGFP2 | -331 ± 4 | -305 ± 7 | +26 mV | <0.001 |
Protocol 1.1: Acute Brain Slice Preparation and FLIM Imaging for Redox Biosensing Objective: To prepare viable acute brain slices from transgenic mice for FLIM redox imaging. Materials:
- Vibratome: For generating 300μm thick acute hippocampal slices.
- Carbogenated (95% O2/5% CO2) Artificial Cerebrospinal Fluid (aCSF): 126 mM NaCl, 2.5 mM KCl, 2 mM CaCl2, 2 mM MgCl2, 1.25 mM NaH2PO4, 26 mM NaHCO3, 10 mM glucose (pH 7.4).
- Two-photon FLIM System: Ti:Sapphire laser tuned to 850-900 nm for roGFP excitation.
- Environmental Chamber: Maintains slice at 32°C during imaging.
Procedure:
- Euthanize age-matched wild-type and APP/PS1 mice (e.g., 3-4 months old) via isoflurane overdose and rapid decapitation.
- Dissect the brain and place in ice-cold, carbogenated aCSF.
- Mount the brain on a vibratome stage and cut 300μm thick coronal sections containing the hippocampus.
- Transfer slices to an incubation chamber with aCSF at 34°C for 30 min, then room temperature for at least 1 hour for recovery.
- Place a single slice in the FLIM imaging chamber under a nylon harp, continuously perfused with oxygenated aCSF at 32°C.
- Identify neurons expressing roGFP2 via epifluorescence.
- Acquire FLIM data using time-correlated single-photon counting (TCSPC) with a 40x water-immersion objective. Collect 100-200 image frames until photon counts per pixel reach >1000 for robust lifetime fitting.
- Perform biexponential decay fitting for each pixel to calculate the fluorescence lifetime.
- For roGFP, calculate the fractional oxidation from the amplitude-weighted mean lifetime (τm) using a standard curve generated from fully reduced (10mM DTT) and fully oxidized (1mM H2O2) conditions on control slices.
- Convert fractional oxidation to Eh using the Nernst equation for a 2-thiol system (Eh = E0 - (59.1/n)*log((1-ox)/(ox)) at 30°C, where E0 for roGFP2 is -280 mV).
Application Note 2: Cancer Model (Pancreatic Ductal Adenocarcinoma)
Study Context: Mapping the heterogeneous and oxidized redox landscape of pancreatic tumors in KPC (KrasG12D/+; Trp53R172H/+; Pdx1-Cre) mouse models. FLIM Probe: rxYFP expressed in cancer-associated fibroblasts (CAFs) and tumor cells via viral delivery. Key Finding: The tumor stroma, particularly CAFs, exhibits a more reduced state compared to malignant epithelial cells, suggesting compartment-specific redox roles in tumor progression and chemoresistance.
Table 2: FLIM-rxYFP Redox Heterogeneity in KPC Pancreatic Tumors
| Cell Type / Region | Probe | Fractional Oxidation (%) | Inferred Redox State |
|---|---|---|---|
| Malignant Epithelial Cells | rxYFP | 68 ± 9 | Oxidized |
| Cancer-Associated Fibroblasts | rxYFP | 32 ± 7 | Reduced |
| Adjacent Normal Acini | rxYFP | 45 ± 6 | Intermediate |
Protocol 2.1: Intravital Window Chamber Preparation and Tumor Redox Imaging Objective: To image redox dynamics in live, orthotopic pancreatic tumors in real-time. Materials:
- Orthotopic Tumor Model: KPC-derived tumor cells expressing rxYFP, injected into the mouse pancreas.
- Abdominal Imaging Window: Titanium dorsal skinfold window chamber modified for pancreatic access.
- Inverted Multi-photon FLIM Microscope: For deep-tissue imaging.
- Anesthesia System: Isoflurane vaporizer with nose cone for prolonged imaging.
Procedure:
- Implant an abdominal imaging window following survival surgery protocols. Allow tumor growth for 10-14 days post-implantation of rxYFP-expressing cells.
- Anesthetize the tumor-bearing mouse with 1-2% isoflurane and secure it on a heated (37°C) microscope stage.
- Use the multi-photon laser (920 nm) to locate the tumor through the window.
- Acquire FLIM stacks (Z-stacks, 5-10 planes, 5μm steps) in both tumor epithelial regions and stromal regions.
- Process FLIM data to generate lifetime maps. rxYFP lifetime is inversely proportional to oxidation. Use control scans with 2mM DTT (reduced) and 2mM diamide (oxidized) in vivo to calibrate the fractional oxidation scale.
- Co-register FLIM redox maps with second harmonic generation (SHG) signals for collagen (stroma) and fluorescence from nuclear labels to identify cell types.
- Perform quantitative analysis by segmenting images based on cell morphology and SHG signal to compare redox states between epithelial and stromal compartments.
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in FLIM Redox Validation |
|---|---|
| Genetically Encoded Redox Probes (roGFP, rxYFP, Grx1-roGFP2) | Target-specific biosensors whose fluorescence lifetime changes with thiol-disulfide oxidation state. |
| Adeno-Associated Viral (AAV) Vectors (Serotypes 1-9, PHP.eB) | For cell-type-specific in vivo delivery and expression of redox probes in rodent disease models. |
| Two-Photon FLIM System with TCSPC Module | Enables deep-tissue, high-resolution lifetime imaging with minimal phototoxicity in live specimens. |
| Lifetime Reference Standard (e.g., Fluorescein) | Provides a known single-exponential decay for daily calibration of the FLIM system. |
| Redox Calibration Reagents (DTT, H2O2, Diamide) | Used ex vivo and in vivo to define the fully reduced and oxidized limits of the probe for quantitative Eh calculation. |
| Tissue-Specific Promoters (e.g., Synapsin, GFAP, Collagen1a1) | Drive probe expression in specific cell types (neurons, astrocytes, fibroblasts) for compartmentalized analysis. |
| Metabolic Modulators (Antimycin A, Rotenone, Paraquat) | Pharmacological tools to perturb specific redox pathways (mitochondrial ETC, ROS production) for functional validation. |
Visualizations
Diagram Title: FLIM Redox Validation Workflow
Diagram Title: Redox Dysregulation in AD Neuron
Within the broader thesis on FLIM (Fluorescence Lifetime Imaging Microscopy) biosensing for thiol-disulfide redox research, the unique value proposition centers on the ability to capture dynamic biochemical events with high spatial and temporal precision while providing absolute quantitative metrics. Unlike intensity-based methods, FLIM measures the exponential decay rate of fluorescence, which is independent of probe concentration, photobleaching, or excitation light intensity. This enables precise, quantitative mapping of cellular redox states, such as glutathione (GSH)/glutathione disulfide (GSSG) ratios or protein thiol oxidation, within living cells and tissues.
Key Quantitative Data in Thiol-Disulfide Redox FLIM Biosensing
Table 1: Representative FLIM Biosensors for Redox Research
| Biosensor Name | Target Redox Parameter | Lifetime Range (τ, ns) | Dynamic Range | Key Application |
|---|---|---|---|---|
| roGFP2 | Glutathione Redox Potential (Eh) | ~2.9 (oxidized) ~3.7 (reduced) | ~-280 mV to -350 mV (Eh) | Cytosolic/mitochondrial GSH/GSSG |
| Grx1-roGFP2 | Glutathione Redox Potential (Eh) | ~2.9 - 3.7 | -280 mV to -350 mV | Specific, equilibrium-based Eh measurement |
| HyPer | H2O2 | ~3.1 - 3.9 | nM to µM H2O2 | Hydrogen peroxide dynamics |
| mrLP1 | Peroxiredoxin-2 Oxidation | ~2.2 - 2.8 | N/A | Mitochondrial matrix H2O2 |
| cpYFP | pH / Redox (crosstalk) | ~0.7 - 3.0 | pH 6-9 | pH control sensor for redox studies |
Table 2: FLIM vs. Intensity-Based Imaging for Redox Sensing
| Parameter | Intensity-Based Ratiometric Imaging (e.g., roGFP) | FLIM-Based Redox Sensing |
|---|---|---|
| Quantification | Relative ratio (e.g., 405/488 nm excitation) | Absolute lifetime (τ) in nanoseconds |
| Probe Concentration Dependency | Highly dependent; requires ratiometric calibration | Largely independent |
| Spatial Resolution | Diffraction-limited (~250 nm lateral) | Diffraction-limited, but functional maps |
| Temporal Resolution | Seconds to minutes (for full ratio) | Seconds (TCSPC/Frequency domain) |
| Absolute Measure | No, requires calibration for Eh | Yes, τ is an absolute physical property |
| Artifact Susceptibility | High (pH, concentration, photobleaching) | Low (insensitive to concentration, excitation intensity) |
Detailed Experimental Protocols
Protocol 1: FLIM Measurement of Glutathione Redox Potential in Live Cells Using Grx1-roGFP2
Objective: To quantitatively map the absolute glutathione redox potential (Eh) in the cytosol of live mammalian cells using FLIM.
Materials:
- HeLa or HEK293 cells expressing Grx1-roGFP2 (targeted to cytosol).
- Imaging medium: Phenol red-free DMEM, buffered with 20 mM HEPES.
- Control reagents: 2 mM DTT (reducing agent), 100 µM Diamide (thiol oxidant).
- FLIM system: Confocal or multiphoton microscope with time-correlated single photon counting (TCSPC) module. Pulsed laser at 488 nm.
Procedure:
- Cell Preparation: Seed cells expressing Grx1-roGFP2 on glass-bottom dishes 24-48h before imaging. Use ~70% confluency.
- System Calibration: Before measurement, calibrate the FLIM system using a known fluorescence standard (e.g., fluorescein in pH 11 buffer, τ ~4.0 ns) to ensure temporal accuracy.
- Lifetime Acquisition: Place dish on pre-warmed (37°C) microscope stage. Use a 60x oil immersion objective. Set 488 nm pulsed laser for excitation at low power to minimize phototoxicity. Collect emission using a bandpass filter (500-550 nm). Acquire photons until peak count reaches 10,000-20,000 in the brightest pixel or for a fixed time (e.g., 90 seconds).
- Data Fitting: Fit the fluorescence decay curve at each pixel to a double-exponential model: I(t) = α₁exp(-t/τ₁) + α₂exp(-t/τ₂) + C. The amplitude-weighted mean lifetime (τm = (α₁τ₁ + α₂τ₂)/(α₁+α₂)) is used for further analysis.
- In-situ Calibration: After baseline imaging, treat cells sequentially with: a. 100 µM Diamide in imaging medium for 5 min, image. b. Wash twice with fresh medium. c. 2 mM DTT in imaging medium for 5 min, image.
- Calculation of Eh: For each pixel, the fraction of oxidized sensor (OxD) is calculated from the lifetime: OxD = (τm - τred)/(τox - τred), where τred and τox are the mean lifetimes from the fully reduced (DTT) and fully oxidized (Diamide) conditions, respectively. Calculate Eh using the Nernst equation: Eh = E0 - (RT/nF)ln([Red]/[Ox]), where E0 for roGFP2 is -280 mV at pH 7.0.
- Spatiotemporal Analysis: Generate lifetime (τm) and calculated Eh maps. Use time-series acquisition to track changes upon addition of redox-modulating drugs.
Protocol 2: Quantifying Protein Thiol Oxidation via FLIM with a Maleimide-based Conjugatable Probe
Objective: To absolutely quantify the oxidation state of protein cysteine thiols in fixed cells.
Materials:
- Cells of interest grown on coverslips.
- Fixative: 4% formaldehyde in PBS.
- Labeling solution: 50 µM BODIPY FL Maleimide (or similar lifetime-sensitive thiol-reactive probe) in PBS with 0.5% DMSO. Note: Prepare fresh.
- Quenching/blocking solution: 20 mM N-ethylmaleimide (NEM) in PBS.
- Reducing control solution: 10 mM Tris(2-carboxyethyl)phosphine (TCEP) in PBS, followed by BODIPY FL Maleimide labeling.
Procedure:
- Fixation: Wash cells with PBS and fix with 4% formaldehyde for 15 min at room temperature. Wash 3x with PBS.
- Free Thiol Blocking: Incubate cells with 20 mM NEM in PBS for 30 min at RT to block all reduced (free) thiols. Wash 3x with PBS.
- Reduction of Oxidized Thiols (for control): For a fully reduced control sample, incubate one set of coverslips with 10 mM TCEP for 30 min after NEM blocking. Wash thoroughly.
- Probe Labeling: Incubate all coverslips (experimental and reduced control) with 50 µM BODIPY FL Maleimide in PBS for 1 hour at RT in the dark.
- Washing: Wash coverslips 5x with PBS over 30 minutes to remove unreacted dye.
- Mounting: Mount coverslips using a slow-fade mounting medium.
- FLIM Acquisition: Image using a 488 nm excitation laser and collect emission >500 nm. Acquire lifetime data with TCSPC.
- Quantitative Analysis: The fluorescence lifetime of the maleimide-bound probe is sensitive to the local microenvironment. A shift to a shorter lifetime may indicate labeling of a thiol in a more hydrophobic or constrained environment (potentially disulfide-bonded prior to reduction). Compare the lifetime distribution (histogram) of the experimental sample vs. the fully reduced control. The absolute lifetime value provides a quantitative measure of the thiol's chemical status, independent of the amount of labeling.
Visualization Diagrams
Title: Thiol-Disulfide Redox Signaling Pathway
Title: FLIM Biosensing Workflow for Redox
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for FLIM-based Redox Biosensing
| Item | Function & Relevance to FLIM Redox Research | Example Product/Source |
|---|---|---|
| Genetically Encoded Biosensor Plasmids | Enable specific, subcellular targeting of redox measurements. Crucial for spatiotemporal resolution. | roGFP2, Grx1-roGFP2, HyPer (Addgene). |
| Lifetime-sensitive Chemical Probes | Conjugatable dyes (e.g., maleimides) for quantifying protein thiol oxidation states. | BODIPY FL Maleimide, Cy3/Cy5 Maleimide (Thermo Fisher). |
| TCSPC FLIM Module | The core hardware for measuring fluorescence decay with picosecond precision. Enables absolute quantification. | Becker & Hickl SPC-150; PicoQuant PicoHarp 300. |
| Pulsed Laser Source | Provides the time-defined excitation pulses required for lifetime measurement. | Ti:Sapphire laser (for multiphoton); picosecond diode lasers (485 nm, 540 nm). |
| Redox Control Reagents | For system calibration and experimental manipulation in live cells. | Dithiothreitol (DTT, reductant), Diamide (oxidant), Tert-Butyl Hydroperoxide (TBHP, ROS inducer). |
| Environment Control Chamber | Maintains live cells at 37°C, 5% CO2 during time-lapse FLIM for physiological relevance. | Okolab stage-top incubator; Tokai Hit chamber. |
| FLIM Analysis Software | For fitting decay curves, calculating lifetime maps, and extracting quantitative data. | Becker & Hickl SPClmage; Fluofit (PicoQuant); open-source (FLIMfit). |
| Immersion Oil (Matched) | Essential for maintaining optimal spatial resolution and photon collection efficiency. | Nikon Type NF; Leica Immersol, with specified dispersion formula. |
Conclusion
FLIM biosensing represents a transformative technology for the quantitative, dynamic, and compartment-specific analysis of thiol-disulfide redox states in living systems. By moving beyond static, population-level snapshots, it provides unparalleled insight into the real-time metabolic and signaling fluxes that govern cell fate. The methodological robustness, when paired with rigorous troubleshooting and validation, establishes FLIM as a gold standard for redox imaging. Future directions include the development of next-generation, target-specific biosensors, integration with high-content screening platforms for drug discovery, and translation towards clinical applications, such as intraoperative redox monitoring. Embracing FLIM empowers researchers to decode the redox language of cells, opening new avenues for understanding disease mechanisms and developing precision therapeutics.