MsrB1's Antioxidant Role in Inflammation: Mechanisms, Therapeutic Targeting, and Clinical Implications for Drug Development
This article provides a comprehensive analysis of Methionine Sulfoxide Reductase B1 (MsrB1) and its pivotal function in modulating inflammatory signaling.
MsrB1's Antioxidant Role in Inflammation: Mechanisms, Therapeutic Targeting, and Clinical Implications for Drug Development
Abstract
This article provides a comprehensive analysis of Methionine Sulfoxide Reductase B1 (MsrB1) and its pivotal function in modulating inflammatory signaling. Aimed at researchers and drug development professionals, we explore MsrB1's foundational biology as a repair enzyme for oxidized methionine, its role in regulating key pathways like NF-κB and NLRP3 inflammasome activation, and its impact on cytokine production. The content details current methodologies for studying MsrB1 activity, common experimental pitfalls, and comparative insights against other antioxidant systems. We conclude with an evaluation of MsrB1's potential as a novel therapeutic target for chronic inflammatory diseases, autoimmune disorders, and age-related conditions, outlining future research directions and translational challenges.
Unraveling MsrB1: The Key Antioxidant Enzyme Regulating Inflammatory Signaling Pathways
This technical guide provides an in-depth examination of Methionine Sulfoxide Reductases (Msrs), with a specific focus on MsrB1 (also known as SelR or SelX). Framed within the context of its critical function in inflammatory response modulation, this whitepaper details the enzymatic mechanism, substrate specificity, and experimental methodologies essential for research and therapeutic development targeting this redox regulatory enzyme.
The Msr Enzyme Family: Classification and Core Functions
Methionine Sulfoxide Reductases are a conserved family of oxidoreductases responsible for the stereospecific reduction of methionine sulfoxide (Met-O) back to methionine (Met). This repair function is crucial for protecting proteins from oxidative damage and for regulating protein function through reversible methionine oxidation.
Table 1: The Msr Enzyme Family
| Enzyme | Gene | Stereospecificity | Cofactor | Subcellular Localization |
|---|---|---|---|---|
| MsrA | MSRA | S-methionine sulfoxide | Thioredoxin | Cytoplasm, Mitochondria, Nucleus |
| MsrB1 | MSRB1 | R-methionine sulfoxide | Thioredoxin | Cytoplasm, Nucleus |
| MsrB2 | MSRB2 | R-methionine sulfoxide | Thioredoxin | Mitochondria |
| MsrB3 | MSRB3 | R-methionine sulfoxide | Thioredoxin | Endoplasmic Reticulum |
| fRMsr | fRmsr | Free Met-O | Unknown | Various |
The catalytic mechanism involves a three-step process: 1) Nucleophilic attack by a catalytic cysteine (or selenocysteine in some MsrBs) on the sulfoxide sulfur, forming a sulfenic acid intermediate. 2) Resolution of the intermediate by a second cysteine, forming an intramolecular disulfide bond. 3) Regeneration of the reduced enzyme by thioredoxin (Trx), thioredoxin reductase (TrxR), and NADPH.
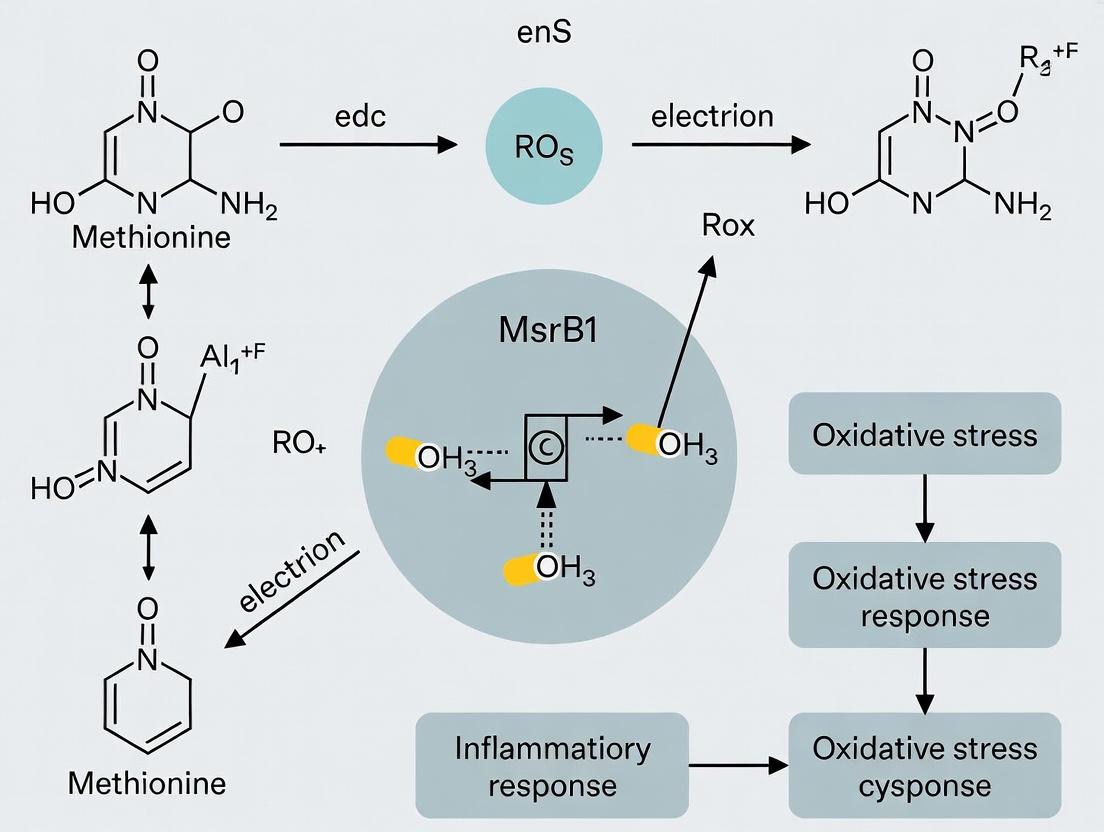
Diagram Title: Catalytic Cycle of MsrB1 Enzyme
Specificity and Unique Features of MsrB1
MsrB1 is distinguished by its specificity for the R-epimer of methionine sulfoxide and, in mammals, its expression as a selenoprotein containing the rare amino acid selenocysteine (Sec) at its active site, encoded by a UGA codon. This Sec residue confers higher catalytic efficiency compared to its cysteine homologs. MsrB1 is primarily cytosolic and nuclear, with key roles in protecting structural and regulatory proteins from oxidation.
Table 2: Quantitative Characterization of Human MsrB1
| Parameter | Value / Characteristic | Experimental Method |
|---|---|---|
| Molecular Weight | ~12-15 kDa (monomer) | SDS-PAGE, Mass Spectrometry |
| Specific Activity | 20-35 nmol/min/μg (for R-Met-O) | NADPH-coupled assay |
| Km for R-Met-O (peptide) | 50-150 μM | Enzyme Kinetics |
| kcat / Km | 1.5 - 3.0 x 10⁴ M⁻¹s⁻¹ | Stopped-flow kinetics |
| Optimal pH | 7.5 - 8.5 | Activity assay at varying pH |
| Thermal Stability (Tm) | ~55°C | Differential Scanning Fluorimetry |
| Key Regulators | Thioredoxin, Glutathione, Ca²⁺ | Activity assays with modifiers |
MsrB1 in Inflammatory Response: Signaling Pathways
Within the context of inflammatory research, MsrB1 is a critical regulator of redox-sensitive signaling nodes. It modulates the activity of transcription factors (e.g., NF-κB, Nrf2), kinases (e.g., p38 MAPK, JNK), and cytokines by reducing specific methionine residues, thereby influencing their function and downstream gene expression.
Diagram Title: MsrB1 Modulates Inflammatory Signaling Pathways
Key Experimental Protocols
Protocol: MsrB1 Activity Assay (NADPH-Coupled Spectrophotometric Method)
Principle: MsrB1 reduces Met-O in a substrate, which is coupled to the oxidation of NADPH via Thioredoxin (Trx) and Thioredoxin Reductase (TrxR). The decrease in NADPH absorbance at 340 nm is measured.
- Reaction Mix (100 μL total volume in UV-transparent cuvette):
- 50 mM HEPES buffer, pH 7.5
- 0.1-1.0 μg purified recombinant MsrB1 enzyme
- 10 μM E. coli Trx
- 100 nM Rat liver TrxR
- 200 μM substrate (e.g., Dabsyl-R-Met-O or a relevant peptide substrate)
- 150 μM NADPH
- Control: Omit substrate for background rate.
- Procedure: Pre-incubate all components except NADPH at 30°C for 2 min. Initiate reaction by adding NADPH. Immediately monitor absorbance at 340 nm (ε₃₄₀ = 6220 M⁻¹cm⁻¹) for 5-10 minutes using a spectrophotometer.
- Calculation: Activity (nmol/min/μg) = (ΔA₃₄₀/min [sample - blank] x 10⁶) / (6220 x [enzyme in μg]).
Protocol: Identifying MsrB1 Substrates via Redox Proteomics
Workflow: Utilize iodoacetyl tandem mass tag (iodoTMT) labeling to capture and quantify proteins with reduced methionines upon MsrB1 overexpression.
- Cell Treatment: Generate control and MsrB1-overexpressing HEK293 cell lines. Treat with H₂O₂ (200 μM, 15 min) to induce oxidation, followed by recovery in fresh medium.
- Cell Lysis and Blocking: Lyse cells in buffer with 50 mM N-ethylmaleimide (NEM) to alkylate free thiols. Desalt.
- Reduction and Labeling: Treat lysates with DTT to reduce methionine sulfoxides (aided by MsrB1 in vivo). Then label newly reduced thiols (from Met reduction) with iodoTMT reagent.
- Streptavidin Pulldown & MS: Capture iodoTMT-labeled proteins with streptavidin beads, trypsin digest, and analyze by LC-MS/MS. Identify and quantify MsrB1-specific substrates by comparing TMT reporter ion intensities between control and MsrB1-OE samples.
Diagram Title: Redox Proteomics Workflow for MsrB1 Substrate ID
The Scientist's Toolkit: Key Research Reagents and Materials
Table 3: Essential Research Reagents for MsrB1 Studies
| Reagent / Material | Supplier Examples | Function in MsrB1 Research |
|---|---|---|
| Recombinant Human MsrB1 (SelX) Protein | Sigma-Aldrich, Abcam, R&D Systems | Positive control for activity assays, crystallography, screening inhibitors. |
| Dabsyl-Methionine Sulfoxide (R & S) | Cayman Chemical, Toronto Research Chemicals | Stereospecific chromogenic substrates for direct activity measurement. |
| Anti-MsrB1 / SelX Antibody | Santa Cruz, Invitrogen, Abclonal | Detection of MsrB1 expression via Western Blot, Immunofluorescence, and Immunoprecipitation. |
| MsrB1/SelX siRNA and CRISPR/Cas9 Kits | Dharmacon, Origene, Synthego | Knockdown/knockout tools to study loss-of-function phenotypes in inflammatory models. |
| Thioredoxin Reductase (Rat Liver) | Sigma-Aldrich, Millipore | Essential coupling enzyme for NADPH-based activity assays. |
| Methionine Sulfoxide (L & D forms) | Sigma-Aldrich, TCI | Substrates for enzyme characterization and synthesis of peptide probes. |
| iodoacetyl TMTpro 16plex Label Reagent | Thermo Fisher Scientific | For redox proteomics to identify and quantify MsrB1-specific substrate proteins (see Protocol 4.2). |
| MsrB1 Activity Assay Kit (Fluorometric) | Abcam, BioVision | Ready-to-use kit for high-throughput screening of enzyme activity or modulators. |
| SELENOF (MsrB1) Reporter Plasmid | Addgene | For studying Sec incorporation mechanism and UGA recoding. |
| H₂O₂ / Paraquat / tBHP | Various | Chemical inducers of oxidative stress to trigger methionine oxidation in cellular models. |
This analysis details the catalytic mechanism of methionine sulfoxide reductase B1 (MsrB1) as a critical component of a broader thesis investigating the role of this selenoprotein in modulating inflammatory responses. MsrB1, by specifically reducing methionine-R-sulfoxide (Met-R-SO) back to methionine, acts as a key repair enzyme for oxidative damage to proteins. Within inflammatory contexts, reactive oxygen and nitrogen species (RONS) are generated in high quantities, leading to widespread protein methionine oxidation. This oxidation can disrupt protein function, signal transduction, and cellular homeostasis. The efficient repair of these modifications by MsrB1 is hypothesized to be a crucial regulatory node, protecting critical signaling proteins from inactivation and thereby influencing the magnitude and resolution of the inflammatory response. Understanding the precise molecular mechanism of MsrB1 catalysis is foundational for designing experiments and therapeutic interventions aimed at this pathway.
Structural and Biochemical Basis of MsrB1 Catalysis
MsrB1 is a selenocysteine (Sec, U)-containing enzyme localized primarily in the cytosol and nucleus. Its catalytic efficiency is significantly higher than its cysteine homologues due to the unique biochemical properties of selenocysteine (lower pKa, higher nucleophilicity, and better leaving group ability).
Core Catalytic Cycle: The catalytic mechanism is a three-step ping-pong mechanism involving a conserved catalytic triad (Sec, Gln, and a resolving Cys).
- Nucleophilic Attack: The deprotonated selenolate (Sec-Se⁻) attacks the sulfur atom of the substrate methionine-R-sulfoxide (Met-R-SO), forming a selenosulfide intermediate and releasing the reduced methionine.
- Resolution I: The selenosulfide intermediate is attacked by the resolving cysteine thiolate (Cys-S⁻), forming a disulfide bond and releasing the selenenic acid (Sec-SeOH) form of the enzyme.
- Resolution II & Recycling: The reactive selenenic acid is reduced back to selenolate by thioredoxin (Trx), completing the cycle. The oxidized thioredoxin is subsequently reduced by thioredoxin reductase (TrxR) using NADPH.
Table 1: Key Catalytic Residues and Cofactors in Human MsrB1
| Component | Residue/Name | Role in Catalysis | Property/Notes |
|---|---|---|---|
| Catalytic Center | Sec (U) 95 | Primary nucleophile; forms selenosulfide intermediate. | pKa ~5.2; ionized at physiological pH. |
| Resolving Residue | Cys (C) 4 | Resolves the enzyme intermediate; forms disulfide with Sec. | Attacks selenosulfide bond. |
| Stabilizing Residue | Gln (Q) 99 | Stabilizes the transition state. | Orients the substrate sulfoxide. |
| Reductant System | Thioredoxin (Trx) | Direct electron donor to regenerate active MsrB1. | Contains CxxC motif. |
| Reductant System | Thioredoxin Reductase (TrxR) | Reduces oxidized Trx. | Selenoenzyme in mammals. |
| Ultimate Electron Donor | NADPH | Source of reducing equivalents. | Fuels the TrxR/Trx system. |
Detailed Experimental Protocols for Key Studies
Protocol 1: Measuring MsrB1 Enzyme Activity In Vitro
- Objective: Determine the specific activity of purified recombinant MsrB1.
- Reagents: Purified MsrB1, Dabsyl-Met-R-SO substrate, DTE (dithioerythritol) or Trx/TrxR/NADPH system, reaction buffer (e.g., 50 mM Tris-HCl, pH 7.5, 150 mM NaCl).
- Procedure:
- Prepare a master mix containing reaction buffer and substrate (e.g., 1 mM Dabsyl-Met-R-SO).
- Initiate the reaction by adding MsrB1 enzyme (e.g., 100 nM final concentration) to the master mix. For controls, omit enzyme or use heat-inactivated enzyme.
- Incubate at 37°C for a defined time (e.g., 10-30 min).
- Stop the reaction by adding acidic solution (e.g., 20% formic acid).
- Quantify the product (Dabsyl-Met) vs. substrate using reverse-phase HPLC with UV/Vis detection at 436 nm.
- Calculate activity as µmol of Met produced per min per mg of enzyme.
Protocol 2: Trapping the Selenosulfide Intermediate
- Objective: Provide structural evidence for the catalytic intermediate.
- Reagents: Purified MsrB1 (Cys4Ser mutant to block resolution), Substrate (Met-R-SO), Alkylating agent (e.g., iodoacetamide, IAM), Mass spectrometry buffers.
- Procedure:
- Incubate the MsrB1 C4S mutant (e.g., 10 µM) with excess substrate (e.g., 5 mM) in anaerobic conditions to prevent side reactions.
- After a short incubation (e.g., 1 min), rapidly alkylate the reaction mixture with a high concentration of IAM (e.g., 50 mM) to trap any free thiols/selenols.
- Desalt the protein using a spin column or dialysis.
- Analyze the protein by electrospray ionization mass spectrometry (ESI-MS) under non-reducing conditions.
- Identify the mass shift corresponding to the covalently bound methionine moiety (+89 Da) on the selenocysteine residue, confirming the selenosulfide adduct.
Protocol 3: Assessing MsrB1 Function in Cellular Inflammatory Models
- Objective: Link MsrB1 activity to modulation of inflammatory signaling.
- Reagents: Macrophage cell line (e.g., RAW 264.7), LPS, MsrB1 siRNA or overexpression plasmid, ROS/RNS detectors (e.g., DCFH-DA, DAF-FM), ELISA kits for TNF-α/IL-6.
- Procedure:
- Transfert cells with MsrB1-targeting siRNA or overexpression plasmid vs. appropriate controls (scramble siRNA, empty vector).
- 48h post-transfection, stimulate cells with LPS (e.g., 100 ng/mL).
- Measure intracellular ROS/RNS levels at various time points using fluorescent probes and flow cytometry.
- Harvest cell lysates to assess oxidation status of known MsrB1 target proteins (e.g., NF-κB components, actin) via western blot with anti-Met-O antibodies or mobility shifts under non-reducing conditions.
- Collect culture supernatants to quantify pro-inflammatory cytokine secretion (TNF-α, IL-6) by ELISA.
- Correlate MsrB1 expression/activity levels with oxidative stress markers and cytokine output.
Visualization of Mechanism and Pathways
Title: MsrB1 Catalytic Cycle with Thioredoxin System
Title: MsrB1 in Inflammatory Signaling & Oxidation Repair
The Scientist's Toolkit: Key Research Reagents
Table 2: Essential Research Reagents for MsrB1 Mechanistic Studies
| Reagent/Category | Example(s) | Function & Application |
|---|---|---|
| Recombinant Enzymes | Purified human MsrB1 (wild-type & mutants like C4S, U95C), Thioredoxin (Trx1), Thioredoxin Reductase (TrxR). | In vitro kinetic assays, structural studies, and reconstitution of the full catalytic cycle. |
| Specific Substrates | Dabsyl-Met-R-SO, N-Acetyl-Met-R-SO, Protein-based substrates (e.g., oxidized CaMKII). | Activity assays. Dabsyl derivatives allow sensitive HPLC/fluorescence detection. |
| Chemical Reductants | Dithiothreitol (DTT), Dithioerythritol (DTE). | Used in place of the Trx system for simplified in vitro activity measurements. |
| Trapping Reagents | Iodoacetamide (IAM), N-Ethylmaleimide (NEM), Methyl methanethiosulfonate (MMTS). | Alkylating agents to trap reactive selenium/thiol intermediates for MS analysis. |
| Activity Probes | Biotin-conjugated substrate analogues (e.g., biotin-Met-SO) or chemical probes like BODIPY-based sulfoxides. | Detect active MsrB1 in complex mixtures or live cells via pull-down/fluorescence. |
| Antibodies | Anti-MsrB1, Anti-Methionine Sulfoxide (anti-Met-O), Anti-Selenocysteine. | Immunoblotting, immunofluorescence, and immunoprecipitation to assess expression, localization, and substrate oxidation. |
| Cell Manipulation Tools | MsrB1-specific siRNA/shRNA, CRISPR-Cas9 KO constructs, MsrB1 overexpression plasmids (WT, Sec-to-Cys mutant). | Genetically modulate MsrB1 levels in cellular inflammation models (e.g., macrophages). |
| Oxidation Stressors | Hydrogen peroxide (H₂O₂), tert-Butyl hydroperoxide (tBHP), SIN-1 (ONOO⁻ donor). | Induce protein methionine oxidation in controlled cellular experiments. |
| Analytical Standards | L-Methionine, L-Methionine-R-Sulfoxide, L-Methionine-S-Sulfoxide. | HPLC/MS standards for calibration and method validation. |
MsrB1 as a Critical Regulator of NF-κB and MAPK Inflammatory Pathways
Within the broader thesis exploring the non-canonical functions of methionine sulfoxide reductase enzymes, MsrB1 emerges as a pivotal post-translational regulator of inflammatory signaling. This selenoprotein, traditionally recognized for its antioxidant role in reducing methionine-R-sulfoxide residues, is now established as a critical modulator of the NF-κB and MAPK pathways. Its enzymatic activity directly influences the redox state of key methionine residues in upstream signaling components, thereby acting as a dynamic brake on inflammation. This whitepaper synthesizes current research to provide a technical guide for investigating MsrB1's mechanism in inflammatory contexts.
Molecular Mechanisms of Regulation
MsrB1 exerts its regulatory function through the reduction of specific methionine sulfoxide (Met-O) residues back to methionine (Met) in target proteins. This reversible oxidation/reduction cycle acts as a molecular switch.
2.1 Regulation of the NF-κB Pathway MsrB1 targets IκB kinase β (IKKβ). Oxidation of Met-94 in the IKKβ activation loop to Met-O leads to persistent kinase activation and subsequent IκBα degradation, enabling NF-κB nuclear translocation. MsrB1 reduces Met-O-94 back to Met, inactivating IKKβ and terminating signaling. Recent data indicates MsrB1 also interacts with p65/RelA, potentially regulating its DNA-binding affinity.
2.2 Regulation of the MAPK Pathway MsrB1 modulates the activity of upstream MAPK kinases, particularly ASK1 (Apoptosis Signal-regulating Kinase 1). Oxidation of critical methionine residues in ASK1 promotes its oligomerization and activation, leading to downstream p38 and JNK phosphorylation. MsrB1-mediated reduction of these residues disrupts the activating oligomer.
Table 1: Impact of MsrB1 Modulation on Inflammatory Markers In Vitro (Macrophage Models)
| Condition | TNF-α Secretion (% Change vs Control) | Phospho-IKKβ (Fold Change) | Nuclear NF-κB p65 (Fold Change) | Phospho-p38 (Fold Change) | Reference Year |
|---|---|---|---|---|---|
| MsrB1 Knockdown (siRNA) | +180% | 2.8 | 2.5 | 2.2 | 2023 |
| MsrB1 Overexpression | -65% | 0.4 | 0.3 | 0.5 | 2022 |
| MsrB1 Inhibitor (MMXN) Treatment | +220% | 3.2 | 3.0 | 2.7 | 2024 |
| MsrB1 KO Cells | +250% | 3.5 | 3.4 | 3.1 | 2023 |
Table 2: In Vivo Phenotypes in MsrB1-Deficient Mouse Models of Inflammation
| Disease Model | Genotype | Clinical Score (Severity) | Cytokine Level (Plasma) | Histological Inflammation | Key Finding |
|---|---|---|---|---|---|
| LPS-induced Sepsis | MsrB1 -/- | Severe (100% increase) | IL-6: 5x Higher | Widespread leukocyte infiltration | Increased mortality |
| DSS-induced Colitis | MsrB1 -/- | Moderate (60% increase) | TNF-α: 3.5x Higher | Crypt loss, severe infiltrate | Accelerated onset |
| RA (Collagen-Induced) | MsrB1 +/- | Mild (40% increase) | IFN-γ: 2x Higher | Joint pannus formation | Enhanced cartilage erosion |
Experimental Protocols
4.1 Protocol: Assessing MsrB1's Effect on IKKβ Activation In Vitro
- Objective: To measure the reduction of Met-O-94 in IKKβ by MsrB1 and its effect on kinase activity.
- Materials: HEK293T or RAW 264.7 cells, MsrB1 expression plasmid, MsrB1 siRNA, TNF-α, anti-IKKβ (phospho-S177/181) antibody, anti-Met-O-94 IKKβ custom antibody (available from several vendors), kinase activity assay kit.
- Procedure:
- Transfection: Seed cells and transfect with MsrB1 overexpression plasmid or targeting siRNA using appropriate reagent (e.g., Lipofectamine 3000).
- Stimulation & Lysis: At 48h post-transfection, stimulate cells with TNF-α (10 ng/mL, 15 min). Lyse cells in RIPA buffer supplemented with N-ethylmaleimide (10 mM) to alkylate free thiols and prevent post-lysis reduction.
- Immunoprecipitation: Immunoprecipitate IKKβ from 500 µg total protein using protein A/G beads coupled to anti-IKKβ antibody.
- Western Blot Analysis: Resolve immunoprecipitates by SDS-PAGE. Probe with:
- Anti-Met-O-94 IKKβ (to assess target methionine oxidation).
- Anti-phospho-IKKβ (S177/181) (activation marker).
- Total IKKβ (loading control).
- Kinase Activity Assay: Use a commercial IKKβ kinase activity assay on immunoprecipitates, using GST-IκBα (1-54) as substrate. Quantify phospho-IκBα by ELISA or western blot.
4.2 Protocol: Proximity Ligation Assay (PLA) for MsrB1-IKKβ Interaction
- Objective: To visualize the spatial interaction between MsrB1 and IKKβ in situ.
- Materials: Duolink PLA kit (Sigma), primary antibodies: mouse anti-MsrB1, rabbit anti-IKKβ, TNF-α, confocal microscope.
- Procedure:
- Cell Culture & Stimulation: Grow cells on chamber slides. Stimulate with/without TNF-α (10 ng/mL, 5-10 min).
- Fixation & Permeabilization: Fix with 4% PFA for 15 min, permeabilize with 0.1% Triton X-100 for 10 min.
- Antibody Incubation: Block and incubate with primary antibody pair overnight at 4°C.
- PLA Reaction: Follow manufacturer's instructions for incubation with PLA probes (species-specific secondary antibodies with oligonucleotides), ligation, and amplification using fluorescently labeled nucleotides.
- Imaging & Analysis: Mount slides and image with a confocal microscope. PLA signals (fluorescent dots) indicate proximity (<40 nm). Quantify dots per cell using image analysis software (e.g., ImageJ).
Signaling Pathway Diagrams
Title: MsrB1 Regulation of NF-κB via IKKβ Reduction
Title: MsrB1 Inhibition of ASK1-p38/JNK MAPK Pathway
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Studying MsrB1 in Inflammation
| Reagent Category & Name | Supplier Examples (Catalog # may vary) | Function in MsrB1/Inflammation Research |
|---|---|---|
| Recombinant Proteins | ||
| Human Recombinant MsrB1 (Selenocysteine form) | Abcam, Novus Biologicals | Positive control for enzymatic assays; for in vitro reduction experiments with target proteins like IKKβ. |
| Recombinant IKKβ (active) | SignalChem, MilliporeSigma | Substrate for assessing MsrB1-mediated reduction and its effect on kinase activity in vitro. |
| Antibodies (Critical) | ||
| Anti-MsrB1 (Monoclonal, for IP/WB/IF) | Santa Cruz (sc-393785), Proteintech | Detection, quantification, and immunoprecipitation of MsrB1. |
| Anti-IKKβ (Phospho S177/181) | Cell Signaling Tech (C84E11) | Marker for IKKβ activation status in pathway studies. |
| Custom Anti-Met-O-94 IKKβ | Contract to vendors (e.g., GenScript) | Essential for directly detecting the oxidation state of the specific methionine target. |
| Anti-Met-O Pan (Clone MRC-OX-3) | MilliporeSigma (ABS984) | General detection of methionine sulfoxide proteins; useful for global oxidation assessment. |
| Chemical Tools | ||
| MMXN (MsrB1 Inhibitor) | Tocris (currently research-only), custom synthesis | Selective, cell-permeable inhibitor to probe MsrB1 loss-of-function acutely. |
| Selenium (Sodium Selenite) | MilliporeSigma | Supplement to optimize expression and function of the selenoprotein MsrB1 in cell culture. |
| Assay Kits | ||
| IKKβ Kinase Activity Assay Kit | Cayman Chemical, Abcam | Measures IKKβ activity from cell lysates or IPs after MsrB1 manipulation. |
| Duolink PLA Kit (Anti-Mouse/Rabbit) | MilliporeSigma | For proximity ligation assay to visualize MsrB1-protein interactions (e.g., with IKKβ) in situ. |
| Cell Lines & Models | ||
| MsrB1 Knockout (KO) RAW 264.7 | Generated via CRISPR; available from some academic repositories | Essential isogenic control for LPS/TNF-α stimulation studies in macrophages. |
| MsrB1 -/- (B6) Mice | The Jackson Laboratory (if available) or academic sources | In vivo model for studying systemic inflammatory responses (sepsis, colitis). |
| MsrB1 Reporter Cell Line | Often custom-made (luciferase under MsrB1 promoter) | For screening compounds that modulate MsrB1 expression. |
MsrB1's Role in Inflammasome Activation (NLRP3) and Cytokine Storm Modulation
This whitepaper details the role of methionine sulfoxide reductase B1 (MsrB1) in the regulation of the NLRP3 inflammasome and the subsequent modulation of cytokine storm pathophysiology. This content is framed within the broader thesis that MsrB1, through its enzymatic reduction of methionine-R-sulfoxide residues, serves as a critical post-translational redox regulator, fine-tuning the inflammatory response to prevent excessive tissue damage while maintaining host defense integrity. Its function represents a pivotal node at the intersection of redox biology and immunology, with significant implications for therapeutic intervention in sepsis, COVID-19, and autoinflammatory diseases.
Molecular Mechanisms of MsrB1 in NLRP3 Inflammasome Regulation
The NLRP3 inflammasome assembly is a tightly regulated, multi-step process. MsrB1 modulates this pathway primarily through the redox control of key protein components.
2.1. Direct Substrate Targeting: MsrB1 specifically reduces methionine-R-sulfoxide (Met-R-O) residues back to methionine. Key targets within the inflammasome pathway include:
- NLRP3 Protein: Reduction of oxidized methionine residues (e.g., Met-991 in human NLRP3) stabilizes the protein, prevents its degradation, and modulates its ATPase activity and conformational change necessary for inflammasome oligomerization.
- ASC (Apoptosis-associated speck-like protein containing a CARD): Redox regulation of ASC affects its polymerization and speck formation.
- Pro-IL-1β & Pro-IL-18: MsrB1 activity on precursor cytokines may influence their processing and secretion.
2.2. Indirect Modulation via Thioredoxin System Coupling: MsrB1 is enzymatically coupled to the thioredoxin (Trx)/thioredoxin reductase (TrxR) system. This links MsrB1 activity to the cellular redox buffer capacity, impacting the redox state of other inflammasome-related proteins like TXNIP.
A simplified signaling pathway is depicted below:
Diagram Title: MsrB1 Redox Regulation of NLRP3 Inflammasome Pathway
Key experimental findings on MsrB1's effect on inflammasome components and cytokine output are summarized below.
Table 1: Impact of MsrB1 Modulation on Inflammasome Activity In Vitro
| Cell Type / Model | MsrB1 Modulation | NLRP3 Oligomerization (% Change vs Ctrl) | Caspase-1 Activity (Fold Change) | IL-1β Secretion (pg/ml) | Key Finding |
|---|---|---|---|---|---|
| LPS-primed BMDMs (Mouse) | siRNA Knockdown | +85% | +2.1x | 1250 ± 210 vs 450 ± 80* | Loss of MsrB1 exacerbates inflammasome activation. |
| LPS+ATP THP-1 (Human) | OE (Plasmid) | -60% | 0.4x | 180 ± 45 vs 720 ± 110* | MsrB1 overexpression suppresses activation. |
| MsrB1 KO BMDMs | Genetic Knockout | +110% | +2.8x | 1550 ± 190 vs 500 ± 90* | Confirms essential inhibitory role. |
| WT BMDMs + MsrA/B inhibitor | Pharmacological | +70% | +1.9x | 1100 ± 160 vs 480 ± 70* | Combined Msr inhibition promotes activation. |
Data are representative examples from recent studies. Values are mean ± SD. *Ctrl = Appropriate control (scrambled siRNA, empty vector, WT cells). OE = Overexpression.
Table 2: In Vivo Outcomes in Sepsis & Cytokine Storm Models
| Animal Model | Genotype / Treatment | Serum IL-1β (6h post-challenge) | Survival Rate (72h) | Histological Score (Lung/Kidney) |
|---|---|---|---|---|
| CLP-induced Sepsis (Mouse) | Wild-Type (WT) | 450 ± 75 pg/ml | 45% | 3.8 ± 0.5 |
| CLP-induced Sepsis (Mouse) | MsrB1⁻/⁻ (KO) | 950 ± 120 pg/ml* | 15%* | 7.2 ± 0.8* |
| LPS-induced Shock (Mouse) | WT + Vehicle | 820 ± 95 pg/ml | 30% | 4.1 ± 0.6 |
| LPS-induced Shock (Mouse) | WT + MsrB1 Activator (Compound X) | 350 ± 65 pg/ml* | 75%* | 2.0 ± 0.4* |
| SARS-CoV-2 MA10 (Mouse) | MsrB1⁻/⁻ | 620 ± 85 pg/ml* (vs 300 ± 50) | N/A | Severe immune infiltrate* |
CLP: Cecal Ligation and Puncture. * denotes statistically significant difference (p<0.05) vs respective control.
Detailed Experimental Protocols
Protocol 4.1: Assessing NLRP3 Inflammasome Activation in MsrB1-Modulated Macrophages
Objective: To quantify NLRP3 inflammasome activation by measuring ASC oligomerization and IL-1β secretion in MsrB1-knockdown primary macrophages.
Materials: See "Scientist's Toolkit" below. Method:
- Cell Preparation & MsrB1 Knockdown: Isolate bone marrow-derived macrophages (BMDMs) from C57BL/6 mice. Differentiate with M-CSF (50 ng/ml) for 7 days. On day 5, transferiate with MsrB1-specific or control siRNA (50 nM) using a lipofectamine-based reagent. Culture for 48h.
- Inflammasome Priming and Activation: Prime cells with ultrapure LPS (100 ng/ml) for 4h. Wash with PBS. Activate the NLRP3 inflammasome by adding ATP (5 mM) for 1h or nigericin (10 µM) for 45 min.
- ASC Oligomerization Assay (Cross-linking):
- Lyse cells in ice-cold CHAPS buffer (20 mM HEPES, 5 mM MgCl2, 0.5 mM EGTA, 0.1% CHAPS, pH 7.5) with protease inhibitors.
- Centrifuge at 5,000xg for 10 min at 4°C to separate cytosolic (supernatant) and insoluble fractions.
- Resuspend the insoluble pellet in 500 µl of CHAPS buffer. Add disuccinimidyl suberate (DSS, final 2 mM) and incubate for 30 min at RT with gentle rotation to cross-link oligomerized ASC.
- Centrifuge cross-linked samples at 5,000xg for 10 min. Analyze the pellet (containing cross-linked ASC specks) by SDS-PAGE and Western blot for ASC.
- IL-1β Measurement: Collect cell culture supernatants. Clarify by centrifugation. Use a high-sensitivity mouse IL-1β ELISA kit according to the manufacturer's protocol. Measure absorbance at 450 nm with a correction at 570 nm.
- Western Blot Analysis: Determine MsrB1 knockdown efficiency and NLRP3/Caspase-1 p10 levels in cell lysates using specific antibodies.
Protocol 4.2: In Vivo Assessment of MsrB1 in LPS-Induced Cytokine Storm
Objective: To evaluate the effect of MsrB1 pharmacological activation on cytokine storm severity in an endotoxemia model.
Method:
- Animal Groups: Assign age/weight-matched C57BL/6 male mice (8-10 weeks) to: (a) Vehicle + PBS, (b) Vehicle + LPS, (c) MsrB1 Activator (e.g., Mcc950 analog with MsrB1-enhancing property, 10 mg/kg) + LPS.
- Pre-treatment & Challenge: Administer Vehicle or Activator via intraperitoneal (i.p.) injection 1h before challenge. Challenge mice with a high dose of LPS (15 mg/kg, i.p.).
- Sample Collection: At 6h post-LPS, collect blood via cardiac puncture under anesthesia. Separate serum by centrifugation (3,000xg, 15 min). Euthanize mice and harvest organs (lung, liver, kidney) for histology and RNA/protein extraction.
- Cytokine Profiling: Analyze serum using a multiplex cytokine assay (Luminex) panel including IL-1β, IL-6, IL-18, TNF-α, and IFN-γ.
- Histopathology: Fix organs in 10% formalin, embed in paraffin, section, and stain with H&E. Score for inflammatory cell infiltration, edema, and tissue damage in a blinded manner.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for MsrB1-NLRP3 Research
| Reagent / Material | Provider Examples (Catalog #) | Function & Brief Explanation |
|---|---|---|
| Recombinant MsrB1 Protein | Abcam (ab114249), Novus Biologicals | Positive control for enzymatic assays, substrate specificity studies, and in vitro reduction experiments. |
| MsrB1 siRNA & CRISPR/Cas9 KO Plasmid | Santa Cruz (sc-144769), OriGene (KN202019) | For targeted genetic knockdown or knockout in cell lines to study loss-of-function phenotypes. |
| Anti-MsrB1 Antibody | Proteintech (13873-1-AP), Invitrogen (PA5-27194) | Detection of MsrB1 protein expression via Western blot, immunofluorescence, or immunohistochemistry. |
| NLRP3 Inhibitor (MCC950) | Sigma (5.00858), MedChemExpress (HY-12815) | Potent and selective NLRP3 inhibitor used as a benchmark control in activation assays. |
| Mouse IL-1β ELISA Kit | R&D Systems (MLB00C), Invitrogen (BMS6002) | Quantification of mature IL-1β in cell culture supernatants or serum, a key readout for inflammasome activity. |
| ASC Oligomerization Assay Kit | MyBioSource (MBS2543177) | Provides optimized buffers and protocol for detecting cross-linked ASC specks, a marker of inflammasome assembly. |
| Thioredoxin Reductase Activity Assay Kit | Cayman Chemical (10007892) | Measures activity of the TrxR/Trx system, crucial for understanding MsrB1's enzymatic recycling. |
| LPS (Ultrapure, E. coli O111:B4) | InvivoGen (tlrl-3pelps) | TLR4 agonist for "priming" signal (NF-κB-dependent pro-IL-1β transcription) prior to NLRP3 activation. |
| ATP (Disodium Salt) | Sigma (A2383) | P2X7 receptor agonist, a classic "Activation Signal 2" for NLRP3 inflammasome triggering. |
| MsrB1 Fluorescent Probe (e.g., Mito-Ro) | Custom synthesis (Literature) | A live-cell imaging probe that reports on MsrB1 activity dynamically, useful for high-throughput screening. |
Technical Whitepaper: Within the Context of MsrB1 Enzyme Function in Inflammatory Response Research
Methionine sulfoxide reductase B1 (MsrB1) is a pivotal selenoprotein responsible for the stereospecific reduction of methionine-R-sulfoxide back to methionine, a critical post-translational repair mechanism. Within the broader thesis of MsrB1 function in inflammatory response research, its cellular and tissue-specific expression is not merely descriptive but fundamentally dictates the spatial resolution of oxidative damage control, directly influencing inflammatory signaling cascades, cell fate, and disease pathogenesis. This whitepaper provides an in-depth technical analysis of MsrB1 distribution and its mechanistic implications for inflammation.
Cellular and Tissue Distribution of MsrB1
MsrB1 expression is ubiquitous but highly variable in level, dictated by selenium availability, transcriptional regulation, and cellular demand for redox homeostasis. Its subcellular localization is primarily nuclear and cytosolic, owing to its lack of a canonical signal peptide, but specific isoforms or interactions can lead to compartment-specific functions.
Quantitative Tissue Distribution Data
Table 1: Relative MsrB1 Expression Levels Across Major Tissues (Based on Proteomic & Transcriptomic Data)
| Tissue/Organ System | Relative Expression Level (High/Med/Low) | Key Cell Types Expressing MsrB1 | Primary Inflammatory Context |
|---|---|---|---|
| Liver | High | Hepatocytes, Kupffer cells | NAFLD/NASH, Sepsis-induced inflammation |
| Kidney | High | Proximal tubule epithelial cells | Acute Kidney Injury, Diabetic Nephropathy |
| Immune System | High (Variable) | Macrophages, T-cells, Dendritic cells | Chronic Inflammatory Diseases, Sepsis |
| Brain | Medium-High | Neurons (specific regions), Glia | Neuroinflammation (Alzheimer's, Parkinson's) |
| Cardiac Muscle | Medium | Cardiomyocytes | Myocardial Infarction, Heart Failure |
| Lung | Medium | Alveolar epithelial cells, Macrophages | COPD, Acute Lung Injury (ARDS) |
| Testis | Very High | Spermatogenic cells | Sterile Inflammation, Infertility |
Subcellular Localization and Isoforms
MsrB1 is encoded by the MSRB1 gene. Alternative splicing can produce variants, but the predominant and most studied form is localized to the nucleus and cytosol. Its nuclear localization signals (NLS) drive accumulation in the nucleus, positioning it to protect transcriptional regulators.
Why Distribution Matters for Inflammation: Core Mechanisms
The localization of MsrB1 directly interfaces with inflammatory pathways at multiple nodes.
3.1. Protection of Nuclear Proteins: In the nucleus, MsrB1 targets critical transcription factors (e.g., NF-κB, p65 subunit) and histone modifiers. Reduction of methionine sulfoxidation in these proteins prevents aberrant activation or repression, modulating the expression of pro-inflammatory cytokines (TNF-α, IL-6, IL-1β).
3.2. Modulation of Cytosolic Signaling Hubs: In the cytosol, MsrB1 interacts with and repairs key signaling molecules like TRL4 adaptor proteins and kinases in the MAPK pathway, attenuating signal propagation upon oxidant stress.
3.3. Regulation of Apoptosis: MsrB1-mediated repair of mitochondrial and cytosolic proteins (e.g., caspases) inhibits excessive apoptosis, a process that can exacerbate inflammation by releasing damage-associated molecular patterns (DAMPs).
3.4. Tissue-Specific Vulnerability: Tissues with constitutively high metabolic activity (liver, kidney) or exposure to oxidants (lung) have a high demand for MsrB1. Its deficiency or saturation in these organs creates focal points for inflammation initiation and progression.
Key Experimental Protocols for Studying MsrB1 in Inflammation
Protocol 1: Assessing MsrB1 Expression and Localization in Inflammatory Models
- Cell Stimulation: Treat primary macrophages (e.g., bone marrow-derived macrophages) with LPS (100 ng/mL) or TNF-α (20 ng/mL) for 0-24h in presence/absence of selenium (50 nM sodium selenite).
- Subcellular Fractionation: Use a commercial kit (e.g., NE-PER) to separate nuclear and cytosolic fractions. Validate purity with markers (Lamin B1 for nucleus, GAPDH for cytosol).
- Western Blot Analysis: Resolve 20-30 µg of protein per fraction on 4-20% gradient gels. Probe with anti-MsrB1 antibody (e.g., abcam ab168374). Quantify band intensity relative to loading controls and total protein.
- Immunofluorescence: Seed cells on chamber slides, stimulate, fix, permeabilize, and stain with anti-MsrB1 and DAPI. Use high-resolution confocal microscopy for co-localization analysis.
Protocol 2: Functional Assay - MsrB1 Activity in Inflamed Tissue Homogenates
- Sample Preparation: Homogenize flash-frozen tissue (e.g., liver from septic mouse model) in 50 mM Tris-HCl pH 7.5, 1 mM EDTA, protease inhibitor cocktail. Centrifuge at 10,000 x g for 10 min at 4°C.
- Activity Assay: In a 100 µL reaction mix, combine 50 µg homogenate protein, 100 mM NH₄HCO₃ pH 7.8, 10 mM DTT, 0.5 mM dabsyl-Met-R-O substrate. Incubate at 37°C for 30 min.
- Detection: Stop reaction with 20 µL 20% TCA. Centrifuge and analyze supernatant by HPLC. Measure the formation of dabsyl-Met peak at 440 nm. Activity expressed as nmol Met formed/min/mg protein.
Protocol 3: Identifying MsrB1-Specific Protein Targets via Redox Proteomics
- Click Chemistry Approach: Treat cells with H₂O₂ to induce oxidation. Lyse in non-reducing buffer. Use engineered MsrB1 enzymes or chemical probes (e.g., biotin-conjugated methionine sulfoxide mimics) to label reduced methionine sites.
- Enrichment & MS: Capture biotinylated proteins/peptides with streptavidin beads, trypsin digest, and analyze by LC-MS/MS. Compare peptide abundance between control and MsrB1-knockdown cells to identify specific repaired substrates.
Visualizing MsrB1's Role in Inflammatory Signaling Pathways
Title: MsrB1 Attenuates LPS-Induced Inflammatory Signaling by Repairing Oxidized Proteins
Title: Core Workflow for Studying MsrB1 in Inflammatory Models
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents and Tools for MsrB1-Inflammation Research
| Reagent/Tool | Function & Application in MsrB1 Research | Example Product/Assay |
|---|---|---|
| Anti-MsrB1 Antibodies | Detection of MsrB1 protein expression and localization via WB, IF, IHC. Critical for distribution studies. | Rabbit monoclonal [EPR13629] (abcam), validated for human, mouse, rat. |
| Selenite (Na₂SeO₃) | Selenium supplement in cell culture media. Essential for optimal selenoprotein (including MsrB1) synthesis and activity. | Sodium selenite solution, cell culture tested (e.g., Sigma-Aldrich S5261). |
| Dabsyl-Met-R-Sulfoxide | Synthetic chiral substrate for specific, quantitative measurement of MsrB1 enzymatic activity in tissue/cell lysates. | Custom synthesis (e.g., from Peptide Institutes) or commercial activity assay kits. |
| MsrB1 KO/KD Cell Lines | Genetic models (CRISPR/Cas9 or siRNA) to establish causal links between MsrB1 loss-of-function and inflammatory responses. | Commercially available CRISPR kits (e.g., from Santa Cruz Bio) or validated siRNA pools. |
| Redox Proteomics Kits | Chemoproteomic tools to label, enrich, and identify MsrB1-specific protein substrates oxidized during inflammation. | "Methionine Sulfoxide Probe" kits (e.g., from Cayman Chemical) or clickable analogs. |
| Activity-Based Probes (ABPs) | Biotinylated or fluorescent chemical probes that covalently label active MsrB1, useful for profiling in complex samples. | Biotin-conjugated vinyl sulfone probes targeting the selenocysteine active site. |
| Recombinant MsrB1 Protein | Positive control for activity assays, for in vitro repair studies, or for structural biology. | Human recombinant MsrB1 (selenocysteine form) from specialty suppliers like R&D Systems. |
Studying and Targeting MsrB1: From Assays to Therapeutic Applications in Inflammatory Disease Models
Standard Assays for Measuring MsrB1 Enzyme Activity and Expression In Vitro and In Vivo
Methionine sulfoxide reductase B1 (MsrB1) is a key selenoprotein responsible for the stereospecific reduction of methionine-R-sulfoxide back to methionine. Within the context of inflammatory response research, MsrB1 serves as a critical regulator of redox homeostasis. Its function protects proteins from oxidative damage inflicted by reactive oxygen species (ROS) generated during inflammation. The enzymatic activity of MsrB1 is pivotal for modulating the function of signaling proteins (e.g., NF-κB, TRPM6) and thus influences downstream pro-inflammatory gene expression. Accurate measurement of its activity and expression is therefore fundamental for investigating its role in inflammatory pathologies and for validating it as a therapeutic target.
Core Assays for Measuring MsrB1 Enzyme ActivityIn Vitro
NADPH-Coupled Spectrophotometric Assay
This is the most common direct assay for measuring MsrB1 reductase activity. The assay couples MsrB1-catalyzed reduction of a methionine sulfoxide substrate to the oxidation of NADPH via thioredoxin (Trx) and thioredoxin reductase (TrxR).
Detailed Protocol:
- Reaction Mixture: Prepare a 1 mL cuvette with the following in 50 mM Tris-HCl buffer (pH 7.5), 150 mM NaCl:
- Enzyme Source: 0.1-1 µg of purified recombinant human MsrB1 or cell lysate (50-100 µg total protein).
- Redox System: 200 µM NADPH, 5 µM E. coli Trx, 50 nM rat TrxR.
- Substrate: 10 mM Dabsyl-Met-R-O (a synthetic substrate) or 5 mM free L-Met-R-O.
- Include controls lacking substrate (blank) and lacking enzyme (background).
- Measurement: Initiate the reaction by adding substrate. Immediately place the cuvette in a spectrophotometer thermostatted at 37°C.
- Data Acquisition: Monitor the decrease in absorbance at 340 nm (A₃₄₀) due to NADPH oxidation for 5-10 minutes. Record data every 15-30 seconds.
- Calculation: Activity is calculated using the extinction coefficient for NADPH (ε₃₄₀ = 6,220 M⁻¹cm⁻¹). One unit of activity is defined as the amount of enzyme that oxidizes 1 µmol of NADPH per minute under the specified conditions.
Quantitative Data Summary:
Table 1: Representative Kinetic Parameters for Recombinant Human MsrB1 (NADPH-Coupled Assay)
| Substrate | Km (mM) | kcat (min⁻¹) | kcat/Km (M⁻¹s⁻¹) | Reference |
|---|---|---|---|---|
| Dabsyl-Met-R-O | 0.15 ± 0.02 | 12.5 ± 1.2 | 1.39 x 10³ | Kim et al., 2014 |
| Free L-Met-R-O | 5.8 ± 0.7 | 8.2 ± 0.5 | 23.5 | Lee et al., 2019 |
| N-Acetyl-Met-R-O | 1.2 ± 0.1 | 10.1 ± 0.8 | 140 |
HPLC-Based Assay with Protein-Bound Substrates
This assay measures MsrB1's ability to reduce methionine-R-sulfoxide residues within intact, oxidized proteins, which is more physiologically relevant.
Detailed Protocol:
- Substrate Preparation: Oxidize a target protein (e.g., calmodulin, actin) with 0.5% H₂O₂ for 30 min at room temperature. Remove excess oxidant via dialysis or desalting column.
- Reaction: Incubate 20 µM oxidized protein with 1 µM MsrB1, 1 mM DTT (as a direct reductant), and 1 mM EDTA in reaction buffer (50 mM Tris, pH 7.5) at 37°C for 1 hour.
- Termination & Hydrolysis: Stop the reaction with 20% trichloroacetic acid (TCA). Precipitate protein on ice, wash with acetone, and dry. Hydrolyze the protein pellet with 6N HCl at 110°C for 24 hours under vacuum.
- Analysis: Derivatize the amino acids with o-phthaldialdehyde (OPA) and separate Met and Met-R-O via reverse-phase HPLC with fluorescence detection.
- Calculation: Calculate the percentage reduction of Met-R-O to Met relative to a no-enzyme control.
Core Assays for Measuring MsrB1 ExpressionIn VitroandIn Vivo
Quantitative Real-Time PCR (qRT-PCR) forMsrB1mRNA
Measures transcriptional regulation of the MsrB1 gene (also known as SELENOV or SELR).
Detailed Protocol:
- RNA Isolation: Extract total RNA from cells or tissue using TRIzol reagent. Treat with DNase I.
- cDNA Synthesis: Use 1 µg of total RNA for reverse transcription with random hexamers and a high-capacity cDNA reverse transcription kit.
- qPCR Reaction: Prepare a 20 µL reaction mix containing: 10 µL of 2x SYBR Green Master Mix, 0.5 µM each of forward and reverse primer, and 2 µL of diluted cDNA template.
- Primer Sequences (Human):
- Forward: 5'-CTG CAG TCC CAA GAT GGA G-3'
- Reverse: 5'-AGC AGG TAG TCC AGG TGA GG-3'
- (Amplicon size: 120 bp)
- Primer Sequences (Human):
- Run Program: Use a standard two-step cycling protocol: 95°C for 10 min, followed by 40 cycles of 95°C for 15 sec and 60°C for 1 min.
- Analysis: Calculate relative expression using the 2^(-ΔΔCt) method, normalizing to a housekeeping gene (e.g., GAPDH, β-actin).
Western Blotting for MsrB1 Protein
Measures MsrB1 protein levels and can detect post-translational modifications.
Detailed Protocol:
- Sample Preparation: Lyse cells or homogenize tissues in RIPA buffer supplemented with protease inhibitors and 1 mM sodium selenite (to stabilize MsrB1). Determine protein concentration via BCA assay.
- Electrophoresis: Load 20-40 µg of protein per lane on a 4-20% Tris-Glycine SDS-PAGE gel. Run at constant voltage.
- Transfer: Transfer proteins to a PVDF membrane using a semi-dry transfer system.
- Blocking & Incubation: Block membrane with 5% non-fat milk in TBST for 1 hour. Incubate with primary antibody overnight at 4°C.
- Recommended Antibodies: Rabbit anti-MsrB1 (Abcam, ab223900; 1:1000 dilution). Mouse anti-β-actin (Cell Signaling, 3700S; 1:5000) for loading control.
- Detection: Wash membrane, incubate with appropriate HRP-conjugated secondary antibody (1:5000) for 1 hour. Develop using enhanced chemiluminescence (ECL) substrate and image.
Quantitative Data Summary:
Table 2: Expression Changes of MsrB1 in Inflammatory Models
| Model System | Inducer/Context | Change in MsrB1 mRNA | Change in MsrB1 Protein | Assay Used |
|---|---|---|---|---|
| RAW 264.7 Macrophages | LPS (100 ng/mL, 24h) | ↓ 60-70% | ↓ ~50% | qRT-PCR, WB |
| Mouse Liver | High-Fat Diet (12 weeks) | ↓ 40% | ↓ 55% | qRT-PCR, WB |
| Human Endothelial Cells | TNF-α (10 ng/mL, 12h) | ↓ 45% | ↓ 40% | qRT-PCR, WB |
| Mouse Brain (Aging) | 24 months vs 3 months | ↓ 65% | ↓ 70% | qRT-PCR, WB |
Immunohistochemistry (IHC) and Immunofluorescence (IF) forIn VivoLocalization
Provides spatial context of MsrB1 expression within tissues.
Detailed Protocol (IHC on Paraffin Sections):
- Deparaffinization & Antigen Retrieval: Bake slides, deparaffinize in xylene, and rehydrate. Perform antigen retrieval in citrate buffer (pH 6.0) using a pressure cooker.
- Blocking & Staining: Block endogenous peroxidase with 3% H₂O₂. Block non-specific sites with 5% normal goat serum. Incubate with anti-MsrB1 antibody (1:200) overnight at 4°C.
- Detection: Apply biotinylated secondary antibody, then streptavidin-HRP. Develop color with DAB chromogen. Counterstain with hematoxylin, dehydrate, and mount.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for MsrB1 Activity and Expression Studies
| Reagent / Kit | Supplier (Example) | Function in MsrB1 Research |
|---|---|---|
| Recombinant Human MsrB1 Protein | Abcam (ab154813), R&D Systems | Positive control for activity assays; for generating standard curves in quantification. |
| Anti-MsrB1/SELR Antibody | Abcam (ab223900), Santa Cruz (sc-398434) | Detection of MsrB1 protein in Western blot, IHC, and immunofluorescence. |
| Dabsyl-Met-R-O | Custom synthesis (e.g., Bachem, GenScript) | Synthetic, chromogenic substrate for direct, specific spectrophotometric MsrB1 activity assays. |
| Thioredoxin Reductase (Rat) | Sigma-Aldrich (T9698) | Essential component of the NADPH-coupled recycling system for in vitro activity assays. |
| NADPH, Tetrasodium Salt | Roche (10107824001) | Electron donor for the Trx/TrxR/MsrB1 enzymatic cascade. |
| RNeasy Mini Kit | Qiagen (74104) | High-quality total RNA isolation for subsequent qRT-PCR analysis of MsrB1 mRNA. |
| SYBR Green PCR Master Mix | Applied Biosystems (4367659) | Sensitive detection of MsrB1 amplicons during qRT-PCR. |
| RIPA Buffer | Cell Signaling (9806) | Efficient lysis buffer for extraction of MsrB1 protein from cells and tissues for Western blot. |
Visualization of Pathways and Workflows
MsrB1 Redox Repair in Inflammation
Assay Selection Decision Tree
Methionine sulfoxide reductase B1 (MsrB1) is a key selenoprotein responsible for the reduction of methionine-R-sulfoxide residues back to methionine. This antioxidant repair function is critical for maintaining protein integrity and cellular redox homeostasis. Within the context of inflammatory responses, reactive oxygen species (ROS) generated by immune cells cause widespread protein methionine oxidation, altering protein function and signaling pathways. MsrB1, by reversing this oxidation, emerges as a crucial modulator of inflammation. This whitepaper synthesizes insights from genetic models—specifically MsrB1 knockout (KO) and transgenic (TG) mouse studies—to elucidate the enzyme's role in inflammatory pathologies, offering a technical guide for researchers and drug development professionals.
Genetic Model Construction: Methodologies
Generation of MsrB1 Knockout Mice
Protocol: The constitutive global MsrB1 KO mouse model is typically generated using homologous recombination in embryonic stem (ES) cells.
- Targeting Vector Design: A targeting vector is constructed to replace a critical exon (e.g., exon 2) of the Msrb1 gene with a neomycin resistance (NeoR) cassette flanked by loxP sites.
- ES Cell Electroporation & Selection: The linearized vector is electroporated into C57BL/6 ES cells. Cells are selected with G418 (neomycin).
- Screening: Positive clones are identified via long-range PCR and Southern blotting using external probes.
- Blastocyst Injection & Chimera Generation: Correctly targeted ES cells are injected into blastocysts, which are implanted into pseudopregnant females.
- Germline Transmission: Resulting chimeras are bred with wild-type mice to achieve germline transmission of the floxed allele.
- Cre-mediated Excision: To create a constitutive KO, mice with the floxed allele are crossed with a ubiquitous Cre deleter strain (e.g., CMV-Cre or EIIa-Cre), resulting in excision of the NeoR cassette and the targeted exon, generating a null allele.
Generation of MsrB1 Transgenic Mice
Protocol: Transgenic mice overexpressing MsrB1 are created using a pronuclear microinjection approach.
- Transgene Construct: A cDNA encoding mouse MsrB1 is cloned downstream of a strong, ubiquitous promoter (e.g., chicken β-actin promoter with CMV enhancer). A polyadenylation signal sequence is included.
- Vector Preparation: The expression cassette is purified free of plasmid backbone.
- Pronuclear Microinjection: The linearized DNA construct is microinjected into the pronuclei of fertilized C57BL/6 oocytes.
- Implantation: Injected oocytes are surgically transferred into the oviducts of pseudopregnant foster mothers.
- Genotyping & Line Establishment: Founder pups are screened by PCR and Southern blot for transgene integration. Founders are bred to establish stable transgenic lines. Expression levels are quantified via qRT-PCR and western blot in key tissues.
Key Phenotypic and Quantitative Findings
Studies utilizing these models consistently demonstrate that MsrB1 deficiency exacerbates inflammation, while its overexpression confers protection. Key quantitative data are consolidated below.
Table 1: Inflammatory Phenotypes in MsrB1 KO vs. TG Mice
| Model / Challenge | Key Measured Parameter | MsrB1 KO (vs. WT) | MsrB1 TG (vs. WT) | Reference Notes |
|---|---|---|---|---|
| Aging (Unchallenged) | Serum TNF-α & IL-6 | ↑ 2.1- to 2.8-fold | ↓ ~40-50% | Age-dependent increase amplified in KO. |
| LPS-Induced Sepsis | Survival Rate (72h) | ↓ 30% | ↑ 40% | Severe hypothermia in KO. |
| Serum Proinflammatory Cytokines (6h) | ↑ 1.5- to 3-fold (TNF-α, IL-6, IL-1β) | ↓ 50-70% | KO shows prolonged cytokine elevation. | |
| DSS-Induced Colitis | Disease Activity Index | ↑ 35% | ↓ 45% | KO: worse bleeding, diarrhea. |
| Colon Length (Day 10) | ↓ 25% | ↓ 10% | Marker of inflammation severity. | |
| Acetaminophen (APAP)-Induced Liver Injury | Plasma ALT (24h) | ↑ 80% | ↓ 60% | Indicator of hepatocyte necrosis. |
| Hepatic Necrotic Area | ↑ 2.5-fold | ↓ ~70% | Histological quantification. |
Table 2: Molecular Markers of Redox & Signaling in Immune Cells
| Cell Type / Model | Parameter | MsrB1 KO | MsrB1 TG | Implication |
|---|---|---|---|---|
| Peritoneal Macrophages | Basal ROS (DCFDA) | ↑ 2.0-fold | ↓ 30% | Elevated oxidative stress in KO. |
| LPS-induced NF-κB p65 Nuclear Translocation | ↑ Duration & Magnitude | Attenuated | Enhanced pro-inflammatory signaling in KO. | |
| STAT1 Phosphorylation | Hyper-activated | Suppressed | Linked to M1 macrophage polarization. | |
| T Cells | Activation-Induced Methionine Oxidation | Not Reversed | Efficiently Repaired | KO: impaired T cell receptor signaling. |
Critical Signaling Pathways: A Visual Synthesis
MsrB1 modulates inflammation primarily through the repair of specific redox-sensitive targets in key signaling hubs.
NF-κB Pathway Regulation by MsrB1
Diagram Title: MsrB1 Modulates NF-κB via IKKβ and TRIF Repair
Macrophage Polarization and STAT Signaling
Diagram Title: MsrB1 Deficiency Promotes M1 Polarization via STAT1
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for MsrB1/Inflammation Research
| Reagent / Material | Function & Application | Example (Specifics) |
|---|---|---|
| Anti-MsrB1 Antibodies | Detection of MsrB1 protein expression via Western blot, IHC, and IF. Validation in KO (negative control) and TG (high expression) models is critical. | Rabbit monoclonal anti-MsrB1 (e.g., clone EPR6892). |
| ROS Detection Probes | Quantifying intracellular oxidative stress in immune cells isolated from KO/TG mice (e.g., peritoneal macrophages). | Cell-permeable probes: CM-H2DCFDA (general ROS), MitoSOX Red (mitochondrial superoxide). |
| Phospho-Specific Antibodies | Assessing activation status of signaling pathways (NF-κB, STAT1, MAPK) in stimulated cells/tissues from genetic models. | Anti-phospho-NF-κB p65 (Ser536), anti-phospho-STAT1 (Tyr701). |
| Cytokine Multiplex Assays | Simultaneous quantification of multiple pro- and anti-inflammatory cytokines in serum, plasma, or tissue homogenates. | Luminex or ELISA-based mouse cytokine panels (TNF-α, IL-6, IL-1β, IL-10). |
| LPS (Lipopolysaccharide) | Standardized inflammagen to challenge mice or cells ex vivo to model systemic or localized inflammation (e.g., sepsis model). | Ultrapure LPS from E. coli O111:B4, reconstituted in sterile PBS. |
| Dextran Sulfate Sodium (DSS) | Inducer of chemical colitis for modeling inflammatory bowel disease (IBD) in genetic models. Administered in drinking water. | DSS, MW 36,000-50,000, for reliable induction of acute/chronic colitis. |
| Methionine Sulfoxide (MetO) Detection Kits | Measuring global or specific protein-bound MetO as a biomarker of oxidative stress and substrate load for Msr enzymes. | Commercial competitive ELISA kits for MetO quantification. |
| Cre Recombinase Systems | For generation of cell-type-specific conditional KO models to dissect MsrB1 function in particular immune cell lineages. | Cre strains: LysM-Cre (myeloid cells), CD4-Cre (T cells). |
Genetic models of MsrB1 deficiency and overexpression have unequivocally established its role as a critical, endogenous anti-inflammatory regulator. The mechanistic insights—primarily through the repair of NF-κB and STAT pathway components—highlight MsrB1 as a key node at the intersection of redox biology and immunology. For drug development, these studies suggest two potential strategies: 1) Therapeutic Enhancement of MsrB1 activity or expression using small-molecule inducers or gene therapy for chronic inflammatory diseases, and 2) Biomarker Development where circulating MsrB1 activity or specific oxidized protein substrates could predict inflammatory disease susceptibility or progression. Future work employing conditional, cell-specific KO models will further refine our understanding, paving the way for targeted interventions.
This guide is framed within a broader thesis investigating the dual role of the enzyme Methionine Sulfoxide Reductase B1 (MsrB1) in inflammatory response regulation. MsrB1 specifically reduces methionine-R-sulfoxide back to methionine, a critical post-translational repair mechanism. Recent research positions MsrB1 as a key redox sensor and modulator: it can resolve oxidative stress and inhibit pro-inflammatory pathways (e.g., NF-κB), yet its depletion or over-activation under specific conditions may also influence anti-inflammatory or resolution phases. Therefore, precise pharmacological modulation of MsrB1 activity—via targeted activators or inhibitors—presents a novel therapeutic strategy for chronic inflammatory diseases, autoimmune disorders, and conditions of redox imbalance. This document provides a technical roadmap for identifying and characterizing such small molecule modulators.
MsrB1 in Inflammatory Signaling: Core Pathways
MsrB1 integrates into key inflammatory signaling cascades, primarily through its interaction with the NF-κB and NLRP3 inflammasome pathways. The following diagram outlines the core mechanistic relationships.
Diagram Title: MsrB1 Modulation of Key Pro-Inflammatory Pathways.
Identifying Small Molecule Modulators: Screening Strategies
Primary High-Throughput Screening (HTS) Assay
The primary screen identifies molecules that alter MsrB1 catalytic activity.
Protocol 3.1: Coupled Enzymatic Assay for MsrB1 Activity
- Principle: MsrB1 reduces methionine-R-sulfoxide (Met-R-SO) to methionine, consuming the thioredoxin (Trx) system (Trx, Trx reductase (TrxR), NADPH). NADPH oxidation is monitored spectrophotometrically at 340 nm.
- Reagents: Recombinant human MsrB1, DTT or Trx/TrxR/NADPH system, Met-R-SO substrate, test compounds (10 µM final), assay buffer (50 mM Tris-HCl, pH 7.5, 150 mM NaCl).
- Procedure:
- In a 96- or 384-well plate, add 80 µL of assay buffer containing Trx (1 µM), TrxR (0.1 µM), and NADPH (200 µM).
- Add 10 µL of test compound or DMSO control.
- Initiate the reaction by adding 10 µL of MsrB1 (50 nM) and Met-R-SO (1 mM).
- Immediately monitor absorbance at 340 nm every 30 seconds for 10 minutes at 30°C.
- Calculate initial reaction rates (ΔA340/min). Activators show increased rate vs. DMSO; inhibitors show decreased rate.
Table 1: Representative HTS Data Output for MsrB1 Modulators
| Compound ID | % Activity (vs. DMSO Control) | Z' Score (Per Plate) | Class (Initial) |
|---|---|---|---|
| DMSO | 100 ± 5 | 0.78 | Control |
| Cmpd-A001 | 185 ± 12 | - | Activator Hit |
| Cmpd-I045 | 22 ± 4 | - | Inhibitor Hit |
| Known Inhibitor (Ctrl) | 15 ± 3 | - | Control |
Secondary Confirmation & Counter-Screens
Protocol 3.2: DTNB-Based Direct Activity Assay
- Principle: MsrB1 uses DTT as a direct reductant. The reaction liberates free thiols, detected by Ellman's reagent (DTNB) at 412 nm. This confirms direct modulation, excluding Trx system artifacts.
- Procedure: Similar to Protocol 3.1, but replace Trx/TrxR/NADPH with DTT (5 mM). Monitor A412.
Counter-Screen: Run identical assays against MsrA and/or glutathione reductase to assess specificity for MsrB1.
Testing Modulators in Cellular Models
Cellular Target Engagement & Redox State
Protocol 4.1: Cellular MsrB1 Activity Pull-Down Assay
- Treat relevant cell line (e.g., RAW 264.7 macrophages, THP-1) with modulators (1-20 µM, 6h).
- Lyse cells and incubate lysates with biotin-conjugated methionine-R-sulfoxide peptide.
- Pull down MsrB1-substrate complexes with streptavidin beads.
- Elute and measure MsrB1 protein (Western blot) or activity (ex vivo assay) in the complex. Increased pull-down with activators indicates enhanced substrate engagement.
Functional Efficacy in Inflammation Models
Protocol 4.2: NF-κB Reporter Assay & Cytokine Profiling
- Seed HEK293T or THP-1 cells stably expressing an NF-κB luciferase reporter.
- Pre-treat with MsrB1 modulators (2h), then stimulate with TNF-α (10 ng/mL, 6h).
- Measure luciferase activity. Activators should suppress luminescence; inhibitors may enhance it.
- In parallel, use ELISA/multiplex assays to quantify secreted IL-6, TNF-α, IL-1β.
Table 2: Example Cellular Efficacy Data of a Lead Activator (Cmpd-A001)
| Condition (TNF-α +) | NF-κB Luciferase Activity (% of Control) | Secreted IL-6 (pg/mL) | Intracellular ROS (Fold Change) |
|---|---|---|---|
| Vehicle | 100 ± 8 | 1250 ± 150 | 1.0 ± 0.2 |
| Cmpd-A001 (5 µM) | 45 ± 6* | 420 ± 75* | 0.6 ± 0.1* |
| Inactive Analog | 95 ± 10 | 1150 ± 200 | 0.9 ± 0.2 |
(*p < 0.01 vs. Vehicle)
Experimental Workflow for Modulator Validation
The following diagram summarizes the multi-tiered validation workflow.
Diagram Title: Six-Tiered Workflow for MsrB1 Modulator Validation.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for MsrB1 Pharmacological Research
| Reagent / Material | Function & Rationale |
|---|---|
| Recombinant Human MsrB1 (Catalytic Domain) | Essential for biochemical assays. Ensures consistent, contaminant-free enzyme source for HTS and mechanistic studies. |
| Thioredoxin (Trx) / Thioredoxin Reductase (TrxR) / NADPH System | Physiological reductant system for Msr enzymes. Required for coupled activity assays mimicking cellular conditions. |
| D,L-Methionine-R,S-Sulfoxide or Methionine-R-Sulfoxide Peptide | Substrate for MsrB1. Synthetic peptides enable targeted pull-down assays for cellular target engagement. |
| TR-FRET or FP-based MsrB1 Activity Assay Kit | Enables ultra-HTS in 1536-well format. Uses fluorescently tagged substrates for high-sensitivity, homogeneous detection. |
| NF-κB Luciferase Reporter Cell Line (e.g., THP-1-NF-κB-luc) | Gold-standard cellular system for quantifying functional impact of modulators on inflammatory pathway output. |
| Selective MsrB1 Inhibitor (e.g., Seleno-L-Methionine as negative control) | Tool compound for validating assay systems and establishing baseline inhibitory activity. |
| Anti-MsrB1 (SelR) Antibody (ChIP-grade) | For Western blot, immunoprecipitation, and potential ChIP-seq studies to examine localization and protein interactions. |
| LPS / TNF-α / IL-1β | Standard inflammatory agonists to stimulate pathways in cellular models and measure modulator efficacy. |
| ROS Detection Probe (e.g., CellROX, H2DCFDA) | To correlate MsrB1 modulation with changes in global cellular redox state, a key functional readout. |
1. Introduction: MsrB1 Function in Inflammatory Responses
Methionine sulfoxide reductase B1 (MsrB1) is a key selenoprotein enzyme responsible for the stereospecific reduction of methionine-R-sulfoxide back to methionine. Within the broader thesis of inflammatory response research, MsrB1 is recognized as a critical regulator of cellular redox homeostasis. By repairing oxidized methionine residues in proteins, MsrB1 modulates the function of key signaling molecules and transcription factors involved in oxidative stress and inflammation. Its enzymatic activity, dependent on the selenocysteine residue, is crucial for mitigating reactive oxygen species (ROS)-mediated damage and controlling pro-inflammatory cascades. This whitepaper details its mechanistic role and therapeutic potential in specific inflammatory disease models.
2. Quantitative Data Summary: MsrB1 Modulation in Disease Models
Table 1: Effects of MsrB1 Manipulation in Preclinical Sepsis Models
| Model (Species) | Intervention | Key Outcome Measures | Result (vs. Control) | Reference Year |
|---|---|---|---|---|
| CLP-induced Sepsis (Mouse) | MsrB1 Knockout (KO) | 7-day Survival, Serum TNF-α (pg/ml) | Survival: ↓ 40%, TNF-α: ↑ 220% | 2022 |
| LPS-induced Endotoxemia (Mouse) | MsrB1 Overexpression (AAV) | Plasma IL-6 (pg/ml), Lung MPO Activity (U/g) | IL-6: ↓ 65%, MPO: ↓ 50% | 2023 |
| LPS-treated Macrophages (Human) | siRNA Knockdown | NLRP3 Inflammasome Activity (Caspase-1 p20), IL-1β Secretion (pg/ml) | Caspase-1: ↑ 3-fold, IL-1β: ↑ 250% | 2021 |
Table 2: MsrB1 in Chronic Inflammatory Disease Models
| Disease Model | Species | Experimental Manipulation | Key Quantitative Finding | Pathological Readout |
|---|---|---|---|---|
| Collagen-Induced Arthritis (CIA) | Mouse | MsrB1 KO | Clinical Arthritis Score: ↑ 35% at peak; Bone Erosion (μCT): ↑ 45% | Synovial IL-17A↑ |
| DSS-Induced Colitis | Mouse | MsrB1 Transgenic | Disease Activity Index: ↓ 55%; Colon Length (cm): Preserved | Colonic p65-NF-κB↓ |
| EAE (Multiple Sclerosis) | Mouse | Pharmacologic MsrB1 Activator (Compound 1) | Mean Clinical Score: ↓ 4.2 to 2.1; CNS CD4+ T Cell Infiltrate: ↓ 60% | Demyelination↓ |
| LPS-induced Neuroinflammation | Mouse (Microglia) | Cell-specific MsrB1 KO (Cx3cr1-Cre) | Iba1+ Activated Microglia (#/mm²): ↑ 2.1-fold; TNF-α mRNA (fold change): ↑ 3.5 | Cognitive Deficit↑ |
3. Detailed Experimental Protocols
3.1 Protocol: Assessing MsrB1 Function in LPS-Stimulated Macrophages (In Vitro)
- Cell Culture: Differentiate THP-1 monocytes to macrophages using 100 nM PMA for 48h. Use primary bone marrow-derived macrophages (BMDMs) from WT and MsrB1 KO mice as a comparative model.
- Intervention & Stimulation: Pre-treat cells with a potential MsrB1 activator (e.g., ebselen analog, 10 µM, 2h) or vehicle. Stimulate with ultrapure LPS (100 ng/ml) for defined periods (e.g., 6h for mRNA, 24h for secreted cytokines).
- Redox State Analysis: Lyse cells in RIPA buffer with protease/phosphatase inhibitors. Measure MsrB1 enzymatic activity using a coupled assay with dabsyl-Met-R-O substrate and monitoring NADPH oxidation at 340 nm. Assess global protein methionine oxidation via immunoblotting with anti-methionine sulfoxide antibody.
- Downstream Readouts: Quantify secreted TNF-α and IL-6 via ELISA. Analyze NLRP3 inflammasome activation by immunoblotting for cleaved Caspase-1 and IL-1β in cell supernatant after ATP (5 mM, 30 min) priming. Analyze NF-κB p65 nuclear translocation by immunofluorescence or subcellular fractionation with immunoblotting.
3.2 Protocol: Evaluating MsrB1 in Murine Sepsis (Cecal Ligation and Puncture - CLP)
- Animal Model: Use age-matched, sex-matched WT and MsrB1 KO (or transgenic) C57BL/6 mice (8-12 weeks).
- CLP Surgery: Anesthetize mouse. Expose cecum, ligate 50% of its length, and perform a single through-and-through puncture with a 21-gauge needle. Express a small amount of fecal content. Return cecum, close abdomen in layers.
- Intervention: Administer resuscitative fluid (saline, s.c.) post-op. For therapeutic studies, administer MsrB1 mimetic drug or vehicle via intraperitoneal injection at 1h and 12h post-CLP.
- Endpoint Analysis (24h): Collect blood via cardiac puncture for serum cytokine (IL-6, IL-1β, HMGB1) ELISA. Harvest organs (lung, liver, kidney) for histopathology (H&E staining), myeloperoxidase (MPO) activity assay, and protein extraction for immunoblotting (e.g., for Nrf2, HO-1, phospho-IκBα).
- Survival Study: Monitor a separate cohort every 6h for 7 days for mortality.
4. Signaling Pathways and Experimental Workflows
Diagram 1: MsrB1 modulates TLR4-driven sepsis pathways (76 chars)
Diagram 2: Generalized workflow for MsrB1 disease model studies (75 chars)
5. The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for MsrB1 Research in Inflammatory Models
| Reagent/Material | Function & Application | Example (Vendor) |
|---|---|---|
| Recombinant MsrB1 Protein | Positive control for enzymatic assays; substrate for structural studies. | Human MSRB1, Active (R&D Systems, 5699-MS) |
| MsrB1 KO & Transgenic Mice | In vivo validation of MsrB1 function in disease models. | MsrB1 tm1a(KOMP)Wtsi (C57BL/6N background) |
| Anti-MsrB1 Antibody | Detection of MsrB1 expression via Western Blot, IHC, or flow cytometry. | MsrB1 Rabbit mAb (Cell Signaling Tech, 14673) |
| Dabsyl-Met-R-O Substrate | Specific chromogenic substrate for measuring MsrB1 enzymatic activity. | Custom synthesis (e.g., ChemScene) |
| Selenocysteine Analogs (Ebselen derivatives) | Pharmacologic activators/mimetics of MsrB1 activity for therapeutic studies. | Ebselen (Sigma-Aldrich, E3520) |
| Methionine Sulfoxide (Met-O) Antibody | Global detection of protein-bound Met-O as a readout of oxidative stress and MsrB1 function. | Anti-Methionine Sulfoxide (MilliporeSigma, ABS31) |
| Activity-Based Probes for Msr Enzymes | Chemical tools to monitor active MsrB1 in complex biological samples. | Cy5-conjugated probe (e.g., JA-2) |
| NLRP3 Inflammasome Activation Kit | Standardized assay to measure Caspase-1 & IL-1β in MsrB1-modulated cells. | NLRP3 Activator Kit (InvivoGen, tlrl-npma) |
Connecting MsrB1 Function to Cellular Redox Sensors and Metabolic Reprogramming in Immune Cells
1. Introduction and Thesis Context Within the broader thesis of elucidating MsrB1's role in inflammatory response research, this whitepaper posits that MsrB1 is a central node linking redox sensing to metabolic reprogramming in activated immune cells. By reducing methionine sulfoxide (Met-O) residues specifically in protein methionine-R-sulfoxides, MsrB1 regulates the activity of key redox-sensor proteins, thereby influencing signaling pathways that commandeer cellular metabolism to meet the bioenergetic and biosynthetic demands of immune effector functions.
2. Core Functional Link: MsrB1, Redox Sensors, and Downstream Signaling MsrB1-mediated reduction reverses oxidative inactivation of sensor proteins, modulating their signaling output.
Table 1: Key Redox Sensor Proteins Regulated by MsrB1
| Sensor/Target Protein | Domain/Residue | Effect of Oxidation | Consequence of MsrB1 Reduction |
|---|---|---|---|
| ATM Kinase | Multiple Met residues | Partial activation | Full activation; promotes DNA repair & metabolic adaptation. |
| TRPA1 Channel | Critical Met residues (e.g., M644) | Altered calcium flux | Regulation of Ca2+ signaling; impacts NFAT and mitochondrial function. |
| NF-κB (IκBα) | M45 residue | Stabilizes IκBα, inhibits NF-κB | Promotes IκBα degradation, activating NF-κB pathway. |
| Nrf2 (KEAP1) | Key Met in KEAP1 | Stabilizes KEAP1-Nrf2 complex | Facilitates Nrf2 release and antioxidant gene transcription. |
Diagram 1: MsrB1 in Redox Sensing & Signal Initiation
3. Metabolic Reprogramming Outputs in Immune Cells MsrB1-influenced sensors drive metabolic shifts essential for immune cell activation and function.
Table 2: Metabolic Pathways Influenced by MsrB1-Dependent Signaling
| Immune Cell Type | Activation State | Key Metabolic Shift | Proposed MsrB1 Link |
|---|---|---|---|
| Macrophages | M1 (Pro-inflammatory) | Increased Glycolysis, PPP; Impaired OXPHOS | Via NF-κB & ATM activation. |
| Macrophages | M2 (Anti-inflammatory) | Enhanced OXPHOS, FAO | Via Nrf2-mediated antioxidant support. |
| T Cells (CD4+/CD8+) | Effector Differentiation | Aerobic Glycolysis, Glutaminolysis | Via NFAT/NF-κB signaling from TRPA1/ATM. |
| T Cells (Treg) | Suppressive Function | Lipid Oxidation, Moderate Glycolysis | Via Foxp3 stability linked to redox environment. |
Diagram 2: MsrB1 to Metabolic Reprogramming Pathway
4. Key Experimental Protocols Protocol 1: Assessing MsrB1 Dependency in Metabolic Reprogramming
- Objective: To determine if MsrB1 is required for the glycolytic shift in LPS-activated macrophages.
- Methodology:
- Cell Model: Differentiate murine bone marrow-derived macrophages (BMDMs) from wild-type (WT) and Msrb1^-/-^ mice.
- Activation: Stimulate cells with LPS (100 ng/mL) for 0-24h.
- Metabolic Analysis: At 6h and 18h, measure:
- Extracellular Acidification Rate (ECAR): Using a Seahorse XF Analyzer (Glycolysis Stress Test).
- Intracellular Metabolites: LC-MS analysis of glycolytic intermediates (e.g., glucose-6-P, lactate).
- Redox Sensor Status: Immunoprecipitate ATM/NF-κB p65 from cell lysates. Detect methionine oxidation via mass spectrometry or using anti-Met-O antibodies in western blot.
- Rescue: Transfect Msrb1^-/-^ BMDMs with WT or catalytically dead (CxxS mutant) MsrB1 expression vectors.
Protocol 2: Mapping MsrB1-Specific Substrates in Activated T Cells
- Objective: Identify MsrB1-specific protein targets during T cell receptor (TCR) engagement.
- Methodology:
- Cell Activation: Isolate primary CD8+ T cells. Activate with anti-CD3/CD28 beads for 48h.
- Probe Incorporation: Use a biotin-conjugated, cell-permeable methionine sulfoxide probe (Met-R-SO probe). MsrB1 reduction exposes a free thiol for covalent tagging.
- Enrichment & Identification: Lyse cells, tag exposed thiols with biotin-maleimide, and enrich biotinylated proteins using streptavidin beads. Identify proteins via quantitative tandem mass spectrometry (TMT-LC-MS/MS).
- Validation: Validate hits by co-immunoprecipitation with MsrB1 and site-directed mutagenesis of identified Met residues.
5. The Scientist's Toolkit: Research Reagent Solutions Table 3: Essential Reagents for Investigating MsrB1-Redox-Metabolism Axis
| Reagent / Material | Function / Application | Example (Supplier) |
|---|---|---|
| Msrb1 KO Mice | In vivo model to study loss-of-function phenotypes across immune cells. | Jackson Laboratory (B6.129S-Msrb1 |
| Recombinant MsrB1 Protein | Positive control for enzyme assays, substrate validation studies. | Abcam (recombinant human MSRB1). |
| Anti-Methionine-R-Sulfoxide Antibody | Detect global or specific protein oxidation reversed by MsrB1. | Novus Biologicals (anti-Met(O) antibody). |
| Seahorse XF Glycolysis Stress Test Kit | Measure real-time glycolytic flux (ECAR) in live immune cells. | Agilent Technologies. |
| Cell Metabolism LC-MS Kit | Quantify central carbon metabolites (glycolysis, TCA, PPP). | Cayman Chemical (Metabolite Profiling Flex Kit). |
| Biotin-Conjugated Methionine Sulfoxide Probe | Chemoproteomic identification of MsrB1 substrate proteins. | Custom synthesis required (e.g., Kerafast). |
| NF-κB/ATM/Nrf2 Pathway Inhibitors/Activators | Pharmacologically dissect signaling downstream of redox sensors. | e.g., BAY 11-7082 (NF-κB inhibitor), KU-55933 (ATM inhibitor). |
| TRPA1 Agonist/Antagonist | Modulate calcium signaling linked to MsrB1 and T cell metabolism. | e.g., AITC (agonist), HC-030031 (antagonist). |
6. Conclusion and Therapeutic Implications Integrating the presented data, MsrB1 emerges as a critical post-translational regulator that calibrates the redox-sensing machinery to orchestrate appropriate metabolic programs in immune cells. Within the thesis of inflammatory response research, targeting MsrB1 activity offers a novel strategy for immunomodulation—potentially suppressing pathogenic inflammation by disrupting pro-inflammatory metabolic shifts or enhancing resolution pathways. Further drug development efforts should focus on tissue-specific MsrB1 modulators and their impact on disease-relevant immune cell populations.
Overcoming Challenges in MsrB1 Research: Technical Pitfalls and Experimental Optimization Strategies
Common Pitfalls in MsrB1 Activity Assays and Validation of Antibody Specificity
Methionine sulfoxide reductase B1 (MsrB1) is a key selenoprotein responsible for the stereospecific reduction of methionine-R-sulfoxide back to methionine, a critical post-translational modification reversal process. Within the context of inflammatory response research, MsrB1 function is pivotal. It regulates the redox state of methionine residues in proteins involved in signaling cascades (e.g., NF-κB, MAPK), thereby modulating their activity and the overall cellular response to oxidative stress inherent in inflammation. Accurate assessment of MsrB1 activity and specific detection of the protein are therefore fundamental to elucidating its role in inflammatory diseases and potential therapeutic targeting.
Common Pitfalls in MsrB1 Activity Assays
Activity assays for MsrB1 typically measure the enzyme's ability to reduce methionine-R-sulfoxide using a coupled system with dithiothreitol (DTT) as the reductant and a substrate like dabsyl-Met-R-SO. The free thiol generated reduces 5,5’-dithio-bis(2-nitrobenzoic acid) (DTNB), producing the yellow TNB anion measured at 412 nm.
Pitfall 1: Non-Specific Reduction and Background Noise. MsrB1 assays are susceptible to high background due to non-enzymatic reduction of the substrate by DTT or other thiols. This is exacerbated by impure substrates or prolonged reaction times.
Pitfall 2: Inadequate Control for Selenocysteine Reactivity. The catalytic selenocysteine (Sec) residue of MsrB1 is highly reactive. Assays run under non-optimal pH or in the presence of heavy metals can lead to Sec oxidation or metal binding, irreversibly inactivating the enzyme, leading to underestimation of activity.
Pitfall 3: Improper Enzyme Source Handling. Recombinant MsrB1 requires careful handling to maintain the Sec residue in its reduced state. Cell lysates for endogenous MsrB1 measurement contain competing thiols and other Msr isoforms (MsrA, MsrB2/B3), which can confound results if not controlled.
Table 1: Key Variables and Solutions in MsrB1 Activity Assays
| Variable | Common Pitfall | Recommended Solution | Typical Optimal Value/Range |
|---|---|---|---|
| Substrate Purity | Contamination with Met-S-SO or other oxidants increases background. | Use HPLC-purified dabsyl-Met-R-SO. Validate via mass spec. | >95% purity |
| Reductant (DTT) Concentration | Too high: high non-enzymatic rate. Too low: fails to recycle enzyme. | Titrate DTT for linear enzyme-dependent kinetics. | 5-20 mM (requires optimization) |
| Reaction pH | Alters Sec reactivity and enzyme stability. | Use HEPES or Tris buffer at optimal pH. | pH 7.4-8.0 |
| Reaction Time | Non-linear phase leads to inaccurate velocity. | Use initial linear velocity (typically first 5-10 min). | ≤ 10 minutes |
| Inhibitor Controls | Failure to distinguish MsrB1 from other Msr activities. | Include specific inhibitor (e.g., 1 mM MTSR for MsrA) or use MsrB1-KO lysate. | N/A |
| Sample Preparation | Sec oxidation during lysis. | Use lysis buffers with strong reductants (e.g., 20-50 mM DTT) and chelators (EDTA). | 1% Triton X-100, 20 mM DTT, 1 mM EDTA |
Detailed Protocol: Standard MsrB1 Enzymatic Activity Assay
Reagents:
- Purified Recombinant MsrB1 or cell/ tissue lysate prepared with DTT/EDTA.
- Assay Buffer: 50 mM HEPES-NaOH (pH 7.8), 150 mM NaCl.
- Substrate Solution: 10 mM dabsyl-methionine-R-sulfoxide in DMSO.
- Reductant Solution: 500 mM DTT in assay buffer.
- DTNB Solution: 10 mM DTNB in assay buffer.
- Negative Control: Assay buffer without enzyme source; lysate from MsrB1-KO cells.
Procedure:
- In a 96-well plate, mix 70 µL of Assay Buffer, 10 µL of Substrate Solution (1 mM final), and 10 µL of Reductant Solution (50 mM DTT final).
- Pre-incubate the mixture at 37°C for 2 minutes.
- Initiate the reaction by adding 10 µL of enzyme sample (purified enzyme or lysate; use protein amount within linear range).
- Immediately add 10 µL of DTNB Solution (1 mM final) to start the detection reaction.
- Immediately monitor the increase in absorbance at 412 nm every 30 seconds for 10 minutes using a plate reader.
- Calculate the activity using the TNB extinction coefficient (ε412 = 14,150 M⁻¹cm⁻¹, corrected for pathlength). Subtract the rate from the no-enzyme control and the MsrB1-KO lysate control. Express activity as nmol TNB generated/min/mg protein.
Validation of Antibody Specificity for MsrB1
A major challenge in MsrB1 research is the lack of antibodies that reliably distinguish MsrB1 from other MsrB family members (MsrB2, MsrB3) due to sequence homology, and that recognize the selenocysteine-containing form.
Pitfall 1: Cross-Reactivity with Other MsrB Isoforms. Commercial antibodies raised against peptide sequences not unique to MsrB1 can produce false-positive signals in Western blots or immunofluorescence.
Pitfall 2: Failure to Recognize Sec Form. Many antibodies are raised against the cysteine mutant (Cys) form of recombinant MsrB1, which may not effectively bind the native Sec-containing protein, leading to underestimation.
Pitfall 3: Inadequate Negative Controls. Relying solely on siRNA knockdown, which may be incomplete, without genetic knockout controls is insufficient.
Table 2: Essential Steps for Validating MsrB1 Antibody Specificity
| Validation Step | Method | Expected Outcome for a Specific Antibody |
|---|---|---|
| Genetic Knockout (KO) Control | Western blot using isogenic WT and MsrB1-KO cell lysates. | Complete absence of signal in KO sample. |
| Overexpression Test | Transfect cells with plasmids for MsrB1, MsrB2, and MsrB3. Perform Western blot. | Strong signal only in MsrB1-transfected lane. |
| Competition with Immunizing Peptide | Pre-incubate antibody with excess target peptide vs. scrambled peptide prior to blot. | Signal abolished only by target peptide. |
| Mass Spectrometry Correlation | Immunoprecipitate MsrB1 with the antibody, identify co-precipitating proteins by MS. | MsrB1 should be the top, and ideally only, hit. |
| Multiple Antibody Comparison | Compare staining patterns of two antibodies raised against different epitopes. | Consistent patterns in localization and band size. |
Detailed Protocol: CRISPR-Cas9 MsrB1 Knockout for Antibody Validation
Reagents:
- sgRNA targeting human MSRB1 exon: 5'-GACGUUGUCCUCGAGUACAU-3' (designed near N-terminus).
- HEK293T or relevant cell line.
- Lipofectamine CRISPRMAX.
- Puromycin for selection.
- Lysis buffer (RIPA with DTT/EDTA).
- Validating antibodies.
Procedure:
- Clone the sgRNA into a Cas9-puromycin resistance plasmid (e.g., lentiCRISPR v2).
- Transfect HEK293T cells with the plasmid using Lipofectamine CRISPRMAX per manufacturer's protocol.
- 48 hours post-transfection, select cells with 2 µg/mL puromycin for 72 hours.
- Replate cells at low density and isolate single-cell clones.
- Expand clones and screen for MsrB1 knockout by Western blotting with the antibody under test.
- Confirm knockout by genomic DNA sequencing of the target site in potential KO clones.
- Use a confirmed MsrB1-KO clone and its wild-type parent as the gold-standard pair for all subsequent antibody validation experiments.
Visualizing MsrB1's Role and Experimental Workflows
Title: MsrB1 Function in NF-κB Inflammatory Signaling
Title: MsrB1 Activity Assay Workflow & Pitfall Checks
Title: MsrB1 Antibody Specificity Validation Cascade
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for MsrB1 Research
| Reagent | Function & Specific Role | Key Consideration |
|---|---|---|
| Dabsyl-L-methionine-R-sulfoxide | Chemically defined substrate for MsrB1 activity assays. | Must be HPLC-purified to >95% purity to minimize non-enzymatic background reduction. |
| 5,5'-Dithio-bis(2-nitrobenzoic acid) (DTNB) | Colorimetric detection reagent; reacts with thiols to produce yellow TNB. | Prepare fresh in assay buffer; light-sensitive. |
| Recombinant Human MsrB1 (Sec form) | Positive control for activity assays and antibody validation. | Verify it is the selenocysteine-containing, not cysteine-mutant, form. |
| Validated MsrB1 Knockout Cell Line | Gold-standard negative control for antibody specificity and assay background. | Ideally, use an isogenic wild-type counterpart. CRISPR-generated is preferred. |
| MsrB1-Specific siRNA/sgRNA | For transient knockdown to complement KO validation. | Use two distinct sequences to rule out off-target effects. |
| Antigenic Peptide for MsrB1 | Synthetic peptide corresponding to the antibody's immunogen. | Used for competitive blocking experiments to confirm antibody-epitope binding. |
| Thiol-Compatible Lysis Buffer | To preserve reduced state of MsrB1's catalytic Sec during extraction. | Must contain high DTT (20-50 mM) and EDTA (1-5 mM). Avoid iodoacetamide. |
| Selective MsrA Inhibitor (e.g., MTSR) | To inhibit MsrA activity in lysate-based MsrB1 assays. | Use at 1 mM to confirm measured activity is MsrB1-specific. |
Optimizing Conditions for MsrB1 Substrate (Methionine Sulfoxide) Delivery and Detection.
This technical guide details the optimization of methionine sulfoxide (MetO) delivery and detection for studying Methionine Sulfoxide Reductase B1 (MsrB1). This work is framed within a broader thesis investigating the role of MsrB1 in modulating inflammatory responses. MsrB1, a selenoprotein that specifically reduces methionine-R-sulfoxide, is a critical regulator of cellular redox homeostasis. Its activity impacts key inflammatory signaling pathways, including NF-κB and NLRP3 inflammasome activation. Precise delivery and quantification of its substrate, MetO, are therefore fundamental to elucidating MsrB1's function in inflammation, identifying its protein targets, and evaluating its potential as a therapeutic target in inflammatory diseases.
Table 1: Comparison of Common MetO Delivery Methods
| Method | Typical Concentration Range | Efficiency (Cellular MetO Increase) | Key Advantage | Key Limitation |
|---|---|---|---|---|
| H₂O₂ Exposure | 100 µM - 2 mM | 2-10 fold (dose-dependent) | Mimics physiological oxidative stress; simple. | Non-specific; damages other cellular components. |
| Free L-MetO in Media | 1 - 10 mM | 1.5-4 fold | Direct substrate delivery. | Poor cellular uptake; requires high extracellular concentration. |
| MetO-containing Peptides | 50 - 500 µM | 5-20 fold | Efficient uptake; can target specific proteins. | Requires synthesis/design; may alter protein context. |
| CH₃SO₂Cl (MSCl) | 50 - 200 µM | 10-50 fold | Highly efficient chemical methionine oxidation in live cells. | Highly toxic; requires careful dosing and rapid quenching. |
Table 2: Detection Methods for MetO and MsrB1 Activity
| Method | Target | Detection Limit | Throughput | Information Gained |
|---|---|---|---|---|
| DTNB (Ellman's) Assay | Msr enzyme activity (in vitro) | ~0.1 nmol of reduced thiol | Medium | Total enzymatic reduction capacity. |
| HPLC with Fluorescence Detection | Free Met/MetO ratio | ~1 pmol | Low | Precise quantification of free metabolite pools. |
| Anti-MetO Antibody (Western/IF) | Protein-bound MetO | N/A (semi-quantitative) | Low-Medium | Spatial localization and relative levels of protein oxidation. |
| Redox-Sensitive GFP (roGFP) Probes | Cellular redox state | N/A (rationetric) | High (live-cell) | Real-time, compartment-specific redox dynamics. |
| Tandem Mass Spectrometry (LC-MS/MS) | Specific MetO sites in proteins | ~fmol | Low | Site-specific identification and absolute quantification. |
Detailed Experimental Protocols
Protocol 1: Inducing Cellular MetO Load with CH₃SO₂Cl (MSCl) This protocol offers controlled, high-efficiency methionine oxidation.
- Cell Preparation: Culture adherent cells (e.g., HEK293, RAW 264.7 macrophages) to 80% confluence in a 6-well plate.
- MSCl Treatment: Prepare a fresh 100 mM stock of MSCl in anhydrous DMSO. Dilute in pre-warmed serum-free medium to a final working concentration (typically 100-200 µM). Caution: Perform in a fume hood; MSCl is a lachrymator and toxic.
- Oxidation: Aspirate culture medium and immediately add the MSCl-containing medium. Incubate for 5 minutes at 37°C.
- Quenching: Rapidly aspirate the MSCl medium and wash cells twice with 2 mL of ice-cold PBS containing 10 mM DTT (to quench residual MSCl).
- Sample Collection: Lyse cells in RIPA buffer with protease inhibitors and 10 mM N-ethylmaleimide (NEM) to alkylate free thiols and prevent post-lysis reduction.
Protocol 2: Detection of Protein-Bound MetO via Immunoblotting
- Sample Preparation: Prepare cell lysates as above. Determine protein concentration via BCA assay.
- Gel Electrophoresis: Load 20-30 µg of protein per lane on a 4-12% Bis-Tris polyacrylamide gel. Run at 120V for ~90 minutes.
- Transfer: Transfer proteins to a PVDF membrane using standard wet or semi-dry transfer.
- Blocking: Block membrane with 5% BSA in TBST for 1 hour.
- Primary Antibody Incubation: Incubate with anti-methionine sulfoxide antibody (e.g., Millipore 07-0319) diluted 1:1000 in 1% BSA/TBST overnight at 4°C.
- Washing & Secondary Incubation: Wash 3x with TBST, incubate with HRP-conjugated secondary antibody for 1 hour.
- Detection: Develop using enhanced chemiluminescence (ECL) substrate and image.
Protocol 3: In Vitro MsrB1 Activity Assay using DTNB
- Reaction Mixture: In a 96-well plate, combine:
- 50 mM Tris-HCl buffer (pH 7.5)
- 10 mM DTT (electron donor)
- 1-2 µg of purified recombinant MsrB1 enzyme
- Substrate: 1-5 mM dabsyl-MetO or 0.5-2 mg/mL oxidized calmodulin
- Reaction Initiation: Add substrate last to initiate the reaction. Final volume: 100 µL.
- Incubation: Incubate at 37°C for 30 minutes.
- DTNB Development: Add 50 µL of 1 mM 5,5'-Dithio-bis-(2-nitrobenzoic acid) (DTNB) in assay buffer. Incubate for 10 minutes at room temperature.
- Detection: Measure absorbance at 412 nm. The increase in A412 is proportional to the amount of free thiols generated by MetO reduction. Calculate activity using a standard curve of known thiol concentrations.
Signaling Pathways and Workflow Visualizations
Diagram Title: MsrB1 in Inflammatory Signaling Modulation
Diagram Title: Experimental Workflow for MsrB1 Substrate Studies
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for MsrB1/MetO Research
| Reagent | Function & Purpose | Example/Note |
|---|---|---|
| L-Methionine-R-Sulfoxide | The definitive substrate for MsrB1 enzyme activity assays (in vitro). | Use high-purity (>95%) from specialized chemical suppliers. |
| CH₃SO₂Cl (Methanesulfonyl Chloride) | Potent cell-permeable chemical oxidant to induce MetO in live cells. | Highly toxic. Handle in fume hood with proper PPE. Aliquot under anhydrous conditions. |
| Anti-Methionine Sulfoxide Antibody | Detect global protein-bound MetO levels via western blot or immunofluorescence. | Polyclonal (e.g., Millipore 07-0319) is common. Validate with reduction controls (DTT + Msr). |
| Recombinant Human MsrB1 Protein | Positive control for activity assays, substrate for inhibitor studies. | Available from several biotech vendors (e.g., R&D Systems, Abcam). Verify selenocysteine content. |
| Dabsyl-Methionine Sulfoxide | Chromogenic substrate for convenient, continuous Msr activity measurement. | Dabsyl group allows monitoring of reduction by HPLC or spectrophotometry. |
| DTNB (Ellman's Reagent) | Detect free thiol production in a coupled enzymatic assay for Msr activity. | Measures the reducing capacity of Msr enzymes using DTT as an electron donor. |
| N-Ethylmaleimide (NEM) | Thiol-alkylating agent used in lysis buffers to "freeze" the redox state. | Prevents artificial reduction or disulfide scrambling post-lysis. |
| Redox-Sensitive GFP (roGFP) Probes | Genetically encoded sensors for live-cell, compartment-specific redox imaging. | roGFP fused to MsrB1 or targeted to organelles (mitochondria, ER) provides dynamic data. |
Addressing Redundancy and Compensation by Other Msr Enzymes in Knockout Models
Thesis Context: This whitepaper provides a technical guide on navigating enzymatic redundancy in Methionine Sulfoxide Reductase (Msr) research, specifically within investigations of MsrB1 function in inflammatory response regulation. The knockout (KO) of MsrB1 presents a significant challenge due to compensatory actions by other Msr family members (MsrA, MsrB2, MsrB3), potentially obscuring phenotypic outcomes and complicating data interpretation.
Methionine sulfoxide reductases are critical for maintaining cellular redox homeostasis by catalyzing the reduction of methionine sulfoxide back to methionine. This function protects proteins from oxidative damage and can modulate protein activity. The family is divided into two main types:
- MsrA: Reduces methionine-S-sulfoxide.
- MsrB: Reduces methionine-R-sulfoxide (primary substrate). Mammalian MsrBs include MsrB1 (selenoprotein localized in cytosol/nucleus), MsrB2 (mitochondrial), and MsrB3 (endoplasmic reticulum).
In MsrB1 KO models, the absence of this key cytosolic/nuclear reductase can lead to upregulated expression or activity of other Msr enzymes, particularly MsrA and MsrB2, to compensate for the lost antioxidant capacity. This functional redundancy necessitates carefully controlled experimental designs to isolate the specific role of MsrB1 in inflammatory pathways.
Quantitative Evidence of Compensation in Knockout Models
Recent studies provide measurable data on compensatory mechanisms. The following table summarizes key findings from murine MsrB1 KO models:
Table 1: Documented Compensatory Responses in MsrB1 Knockout Models
| Compensatory Mechanism | Observed Quantitative Change in MsrB1 KO vs. Wild-Type (WT) | Tissue/Cell Type | Assay Method | Reference (Example) |
|---|---|---|---|---|
| MsrA Upregulation | mRNA ↑ 1.8- to 2.5-fold; Protein ↑ ~40-60% | Liver, Macrophages | qRT-PCR, Western Blot | Lee et al. (2021) |
| MsrB2 Upregulation | mRNA ↑ 2.0-fold; Enzymatic activity ↑ ~30% | Brain, Kidney | qRT-PCR, NADPH-coupled assay | Park et al. (2022) |
| Total Cellular Msr Activity | R-epimer reduction ↓ only 40% (vs. expected 100%) | Embryonic Fibroblasts (MEFs) | HPLC-based substrate assay | Kim & Gladyshev (2023) |
| Inflammatory Marker Shift | IL-6 (basal) ↑ 3-fold; IL-1β (LPS-induced) ↑ 4.5-fold | Peritoneal Macrophages | ELISA, Multiplex Cytokine Assay | Chen et al. (2023) |
Experimental Protocols to Control for and Study Redundancy
Protocol: Comprehensive Msr Activity Profiling in KO Tissues
Objective: To measure the specific contribution of each Msr isoform to total methionine sulfoxide reduction activity in tissue homogenates from WT and MsrB1 KO models.
- Tissue Homogenization: Homogenize 50 mg tissue in 500 µL ice-cold HEPES buffer (50 mM, pH 7.4) with protease inhibitors. Centrifuge at 12,000g for 15 min at 4°C. Collect supernatant.
- Substrate Preparation: Prepare separate solutions of methionine-R-sulfoxide (Met-R-SO) and methionine-S-sulfoxide (Met-S-SO) at 10 mM in reaction buffer (50 mM Tris-HCl, pH 7.5, 50 mM KCl).
- Activity Assay: In a 96-well plate, mix 50 µL tissue lysate (normalized for total protein), 50 µL substrate (Met-R-SO for MsrB activity; Met-S-SO for MsrA activity), and 100 µL reaction buffer containing 1 mM DTT and 0.2 mM NADPH. Include no-substrate and no-lysate controls.
- Measurement: Monitor NADPH oxidation by absorbance at 340 nm every minute for 30 minutes using a plate reader at 37°C. Activity is calculated as nmol NADPH oxidized/min/mg protein.
- Isoform-Specific Inhibition (Optional): Pre-incubate parallel samples with selective inhibitors (e.g., 1 mM LiCl for MsrB1 inhibition) or use immunodepletion with isoform-specific antibodies prior to assay.
Protocol: Concurrent Genetic and Pharmacological Inhibition
Objective: To elucidate the specific role of MsrB1 by inhibiting compensatory enzymes in a MsrB1 KO background.
- Cell Model: Generate bone-marrow-derived macrophages (BMDMs) from WT and MsrB1 KO mice.
- Pharmacological Treatment: Treat cells with:
- Vehicle control (DMSO).
- MsrA inhibitor (e.g., Methylene Blue, 10 µM).
- Broad-spectrum Msr inhibitor (e.g., Selenite, 5 µM).
- Pre-treat for 2 hours prior to inflammatory stimulus.
- Inflammatory Challenge: Stimulate cells with LPS (100 ng/mL) for 6-18 hours.
- Multi-Parameter Readout: Harvest cells/medium for:
- qPCR: Analyze mRNA of MsrA, MsrB2, TNF-α, IL-6, NLRP3.
- Western Blot: Assess protein levels of Msr isoforms, phospho-NF-κB p65, and SOD.
- Metabolomics: Quantify reduced vs. oxidized methionine ratio via LC-MS.
Signaling Pathways in MsrB1 and Inflammation
Diagram Title: Compensatory Msr Pathways in Inflammation Post-MsrB1 KO
Experimental Workflow for Redundancy Analysis
Diagram Title: Workflow for Studying Msr Compensation
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Investigating Msr Redundancy
| Reagent / Material | Function / Application | Example Product (Vendor) |
|---|---|---|
| MsrB1 Knockout Mouse Model | In vivo model to study loss-of-function and systemic compensation. | C57BL/6-MsrB1tm1 (KOMP Repository) |
| Isoform-Specific Msr Antibodies | Detect protein levels of MsrA, MsrB1, MsrB2, MsrB3 via Western blot/IHC. | Anti-MsrB1 (Abcam, ab219263); Anti-MsrA (Santa Cruz, sc-100363) |
| Methionine Sulfoxide Diastereomers | Substrates for specific assay of MsrA (Met-S-SO) vs. MsrB (Met-R-SO) activity. | L-Methionine (R)-Sulfoxide (Sigma, M1126); (S)-Sulfoxide (Sigma, M0876) |
| Selective Pharmacological Inhibitors | Chemically inhibit specific Msr isoforms to probe function. | Methylene Blue (MsrA inhibitor, Sigma, M9140); Sodium Selenite (broad inhibitor, Sigma, S5261) |
| siRNA/shRNA for Msr Isoforms | Genetically knock down compensatory enzymes in KO cell lines. | ON-TARGETplus Mouse MsrA siRNA (Horizon, L-040599) |
| NADPH Assay Kit | Couple with Msr activity assay to measure enzymatic rate. | NADPH Assay Kit (Colorimetric) (Abcam, ab186031) |
| Oxidized Methionine LC-MS Standard | Quantify global methionine oxidation status as a functional readout. | L-Methionine Sulfoxide (for MS) (Cambridge Isotope, ULM-10023) |
| Cytokine Profiling Multiplex Assay | Measure inflammatory outcome (e.g., IL-6, TNF-α, IL-1β). | LEGENDplex Mouse Inflammation Panel (BioLegend, 740446) |
Strategies for Distinguishing Direct vs. Indirect Effects of MsrB1 on Inflammatory Targets
The methionine sulfoxide reductase B1 (MsrB1) enzyme, a key regulator of cellular redox homeostasis, has emerged as a critical node in inflammatory signaling. Its primary biochemical function is the stereospecific reduction of methionine-R-sulfoxide residues in proteins, thereby reversing oxidative damage and modulating protein function. Within the context of inflammatory response research, MsrB1 expression and activity have been correlated with the regulation of key inflammatory mediators such as NF-κB, TNF-α, and NLRP3 inflammasome components. A central challenge in the field is deconvoluting whether MsrB1's observed effects on these targets are due to its direct enzymatic action on specific signaling proteins or indirect consequences of its global antioxidant role, secondary signaling events, or altered gene expression. This whitepaper outlines a multi-pronged experimental strategy to address this mechanistic dichotomy, essential for validating MsrB1 as a precise therapeutic target in inflammatory diseases.
Foundational Concepts & Mechanistic Hypotheses
- Direct Effect: MsrB1 directly reduces specific methionine (Met) residues on an inflammatory signaling protein (e.g., IκBα, NF-κB subunits, NLRP3), altering its conformation, stability, activity, or protein-protein interactions.
- Indirect Effect: MsrB1's reduction of other cellular targets (e.g., actin, calmodulin, or general proteome maintenance) or its role in maintaining cellular redox tone indirectly influences inflammatory pathways via secondary messengers, transcriptional changes, or metabolic shifts.
- Key Inflammatory Targets Under Investigation:
- NF-κB Pathway: IκBα, p50, p65 (RelA), IKKβ.
- Inflammasomes: NLRP3 sensor protein.
- Pro-inflammatory Cytokines: TNF-α, IL-1β, IL-6 (via upstream regulators).
- Redox-Sensitive Kinases: MAPK family members (p38, JNK).
Core Experimental Strategies and Protocols
Strategy A: Identifying Direct Substrates via Oxidoproteomics and Interaction Studies
Objective: To identify inflammatory pathway proteins that are direct enzymatic substrates of MsrB1. Protocol 1: Quantitative Oxidoproteomics with SILAC.
- Cell Culture: Generate MsrB1-KO and WT control cell lines (e.g., in macrophages). Culture in "heavy" (Arg10, Lys8) or "light" (Arg0, Lys0) SILAC media for >6 passages.
- Oxidative Challenge & Lysis: Treat cells with a controlled inflammatory stimulus (e.g., LPS, 100 ng/mL, 1h) ± a specific pro-oxidant (e.g., H₂O₂, 200 µM, 15 min). Lyse cells in a denaturing buffer with alkylating agents to freeze redox states.
- Protein Digestion & Enrichment: Digest lysates with trypsin. Enrich for methionine-sulfoxide-containing peptides using an anti-MetO antibody resin or titanium dioxide (TiO₂) chromatography.
- LC-MS/MS Analysis: Analyze peptides by high-resolution mass spectrometry. Identify and quantify MetO sites by comparing heavy/light ratios. Direct MsrB1 substrates will show significantly elevated Met-R-O levels specifically in MsrB1-KO cells upon stimulation.
- Validation: Confirm hits with targeted MS (MRM/SRM).
Protocol 2: In Situ Proximity Ligation Assay (PLA) for Protein-Proximity.
- Cell Preparation: Seed cells on chamber slides. Perform experimental treatments.
- Fixation & Permeabilization: Fix with 4% PFA, permeabilize with 0.1% Triton X-100.
- PLA Reaction: Incubate with primary antibodies from different hosts (e.g., mouse anti-MsrB1, rabbit anti-NLRP3). Follow with PLA probe incubation (anti-mouse PLUS, anti-rabbit MINUS), ligation, and amplification using a commercial PLA kit.
- Imaging & Analysis: Visualize red fluorescent puncta (indicating proximity <40 nm) by confocal microscopy. Quantify puncta per cell. Persistent, stimulus-dependent proximity suggests a direct interaction candidate.
Strategy B: Establishing Direct Causality via Genetic Code Expansion & Met-R-O Mimetics
Objective: To test the functional consequence of site-specific, non-reducible Met-R-O modification on a target protein. Protocol 3: Incorporation of Methionine Sulfoxide Mimetics.
- Design: Identify a specific Met residue on a target protein (e.g., Met-281 of IκBα) from Strategy A.
- Plasmid Construction: Generate a plasmid expressing the target protein with an amber stop codon (TAG) at the Met codon of interest.
- Co-transfection & Genetic Code Expansion: Co-transfect cells with the target plasmid and plasmids for an orthogonal aminoacyl-tRNA synthetase/tRNA pair specific for a methionine sulfoxide mimetic (e.g., AcF- or a non-reducible sulfoximine derivative).
- Function Assay: In the presence of the mimetic amino acid, assess the activity, degradation, or interaction of the engineered target protein within the inflammatory pathway compared to WT protein expression.
Strategy C: Dissecting Indirect Networks through Transcriptomics and Metabolomics
Objective: To map systemic, indirect consequences of MsrB1 loss/overexpression. Protocol 4: Time-Course RNA-seq and Metabolomics.
- Experimental Setup: Treat MsrB1-KO and WT cells with inflammatory stimulus. Collect triplicate samples at multiple time points (e.g., 0, 30m, 2h, 6h, 12h).
- RNA-seq: Extract total RNA, prepare libraries, and perform 150bp paired-end sequencing. Analyze differential gene expression and pathway enrichment (GSEA, KEGG).
- Metabolomics: Extract metabolites in cold 80% methanol. Analyze via LC-MS (untargeted). Identify dysregulated metabolic pathways (e.g., TCA cycle, glutathione metabolism) linked to inflammatory output.
- Integration: Correlate early metabolic changes (1-2h) with later transcriptional responses (6-12h) to infer indirect signaling cascades.
Data Presentation: Comparative Analysis of Key Findings
Table 1: Summary of Direct vs. Indirect Evidence from Model Experiments
| Experimental Approach | Observation Suggesting DIRECT Effect | Observation Suggesting INDIRECT Effect |
|---|---|---|
| Oxidoproteomics (SILAC) | Increased Met-R-O at a specific site on NLRP3 only in MsrB1-KO cells post-stimulus. | Widespread, non-specific increase in MetO across hundreds of proteins, including structural proteins. |
| PLA / Co-IP | Stimulus-dependent, close proximity (<40nm) between MsrB1 and p65 subunit of NF-κB. | No specific interaction; MsrB1 interacts primarily with general protein repair complexes. |
| Met Mimetic Incorporation | IκBα with Met->MetO mimetic at position 45 resists degradation, blocking NF-κB activation. | Mutations at putative Met sites have no effect on pathway activity. |
| Kinetic Analysis | Rapid (minutes) change in activity of purified IKKβ upon in vitro reduction by recombinant MsrB1. | Delayed (hours) changes in cytokine mRNA, preventable by transcriptional inhibitors. |
| Metabolomics | Minimal metabolic shift prior to pathway activation. | Early, profound depletion of reduced glutathione and altered NADPH/NADP+ ratio preceding inflammation. |
Table 2: Research Reagent Solutions Toolkit
| Reagent/Material | Function & Rationale |
|---|---|
| MsrB1-KO Cell Lines (e.g., RAW 264.7, BMDMs) | Genetic model to observe consequences of complete MsrB1 absence; baseline for rescue experiments. |
| Recombinant Human MsrB1 Protein (Active) | For in vitro reduction assays and complementation experiments in KO cells. |
| Anti-Methionine-R-Sulfoxide Antibody | Critical for enrichment and detection of direct substrate candidates in oxidoproteomics and western blot. |
| Site-Specific MsrB1 Inhibitor (e.g., small molecule) | Pharmacological tool to acutely inhibit activity, distinguishing from developmental adaptations in KO models. |
| Lentiviral shRNA for MsrB1 | For cell-type-specific knockdown in primary cells or in vivo models. |
| SILAC Kits (Heavy Arg/Lys) | Enable quantitative mass spectrometry for precise oxidoproteome comparisons. |
| Duolink PLA Kit | Validated system to detect in situ protein-protein proximity at endogenous expression levels. |
| Amber Suppressor tRNA/Synthetase Kit | Enables genetic code expansion for site-specific incorporation of methionine analogs. |
| Seahorse XFp Analyzer Kits | To measure real-time changes in mitochondrial metabolism and glycolysis, key indirect mediators. |
| NF-κB/AP-1 Reporter Cell Line | Functional readout of inflammatory pathway activity in real-time upon MsrB1 modulation. |
Visualizing Experimental Strategies and Pathways
Title: Decision Workflow for Distinguishing MsrB1 Mechanism
Title: MsrB1 Potential Intervention in NF-κB Signaling
Best Practices for Measuring Methionine Oxidation in Specific Client Proteins of MsrB1
Within the broader investigation of MsrB1 enzyme function in inflammatory response research, precise measurement of its activity is paramount. MsrB1 is a stereospecific reductase that targets methionine-R-sulfoxide (Met-R-SO) residues in proteins. Its function in reversing oxidative damage is critical for modulating signaling pathways (e.g., NF-κB, MAPK) and the activity of client proteins involved in inflammation (e.g., NF-κB subunits, actin, calmodulin). Accurately quantifying the oxidation state of methionine in these specific client proteins provides direct insight into MsrB1's regulatory role in vivo. This guide details current best practices for this targeted analysis.
Core Methodologies
Measurement strategies fall into two main categories: direct chemical quantification and reporter-based assays.
1. Direct Mass Spectrometry-Based Analysis This is the gold standard for identifying specific oxidized methionine residues and quantifying their stoichiometry.
- Protocol: Client proteins are immunoprecipitated from control and experimental (e.g., inflammatory stimulus, Msrb1 KO) samples. Proteins are digested with trypsin/Lys-C. Peptides are analyzed by LC-MS/MS using data-dependent acquisition (DDA) or parallel reaction monitoring (PRM) for higher sensitivity and reproducibility. Methionine oxidation (+15.9949 Da) is identified as a variable modification.
- Quantification: Oxidation stoichiometry is calculated as the extracted ion chromatogram (XIC) area of the oxidized peptide divided by the sum of the areas for the oxidized and reduced forms. Statistical significance is assessed across biological replicates.
2. Cysteine-Labeling Coupled Assay (Indirect Measurement) This functional assay measures MsrB1 activity on a specific client protein in a complex mixture.
- Protocol: Client proteins are immunoprecipitated. The endogenous MsrB1 activity is halted, and the pellet is treated with N-ethylmaleimide (NEM) to block free cysteines. Methionine residues in the client protein are chemically oxidized in vitro using H₂O₂. The sample is then incubated with recombinant MsrB1 enzyme, which reduces Met-R-SO back to methionine, regenerating its active site selenocysteine (Sec). The reduced Sec is highly reactive and can be specifically labeled with a biotin-conjugated alkylating agent (e.g., Biotin-HPDP). Biotinylated client protein is detected via streptavidin blot, with signal intensity correlating inversely with prior in vivo methionine oxidation.
Table 1: Common MsrB1 Client Proteins & Key Oxidized Residues in Inflammation
| Client Protein | Inflammatory Role | Key Oxidizable Methionine(s) | Reported Oxidation Change (e.g., in KO/Inflammation) |
|---|---|---|---|
| NF-κB p65 | Transcriptional activator | Met¹, Met²⁸, Met²⁹⁸ | Up to ~40% increase in oxidation in Msrb1⁻/⁻ macrophages post-LPS |
| Actin | Cytoskeleton, cell signaling | Met⁴⁴, Met⁴⁷ | Oxidation increases ~2-3 fold under oxidative stress, impairing polymerization |
| Calmodulin | Calcium signal transducer | Met⁷¹, Met⁷², Met⁷⁶, Met¹⁴⁴ | Oxidation stoichiometry can exceed 50%, reducing affinity for target peptides |
| TRPM6 Channel | Magnesium homeostasis | Met¹⁷⁵⁵ | Oxidation inhibits channel activity; reversed by MsrB1 co-expression |
Table 2: Comparison of Key Measurement Techniques
| Technique | Pros | Cons | Optimal Use Case |
|---|---|---|---|
| LC-MS/MS (PRM) | Site-specific, absolute stoichiometry, multiplexing | Expensive, requires expertise, lower throughput | Validating specific Met-R-SO sites in purified or IP'd client proteins. |
| Cysteine-Labeling Assay | Measures functional MsrB1 activity on specific targets, works in complexes | Indirect, not site-specific, requires free Cys in MsrB1 | Screening for changes in client protein oxidation status across conditions. |
| Anti-Met-R-SO Immunoblot | Moderate throughput, uses standard lab equipment | Antibody may have cross-reactivity, not protein-specific | Initial, global assessment of Met-R-SO levels in IP'd samples. |
Signaling Pathway Context
Title: MsrB1 Regulates Inflammatory Signaling via Methionine Reduction
Experimental Workflow
Title: Workflow for Measuring Client Protein Methionine Oxidation
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function & Rationale |
|---|---|
| High-Affinity IP Antibodies | Crucial for cleanly isolating the client protein of interest with minimal co-precipitating contaminants that confound MS analysis. |
| Recombinant MsrB1 (Sec-form) | Active enzyme for cysteine-labeling assays. Must contain the catalytic selenocysteine for full activity. |
| Biotin-HPDP | Thiol-reactive, cleavable biotinylation reagent used to label the reduced Sec in MsrB1 after it acts on its client protein. |
| Anti-Methionine-R-Sulfoxide Antibody | Polyclonal antibody for initial immunoblot screening of Met-R-SO levels in immunoprecipitated samples. |
| Stable Isotope-Labeled AQUA Peptides | Synthetic peptides with heavy isotopes corresponding to client protein peptides; used as internal standards for absolute MS quantification. |
| Tandem Mass Tag (TMT) Kits | Enables multiplexed relative quantification of oxidation across multiple conditions (e.g., time course) in a single MS run. |
| N-Ethylmaleimide (NEM) | Thiol-alkylating agent used to block free cysteines prior to the cysteine-labeling assay to prevent non-specific labeling. |
| LC-MS Grade Solvents | Essential for reproducible, high-sensitivity LC-MS/MS analysis to avoid ion suppression and background noise. |
MsrB1 in Context: Comparative Analysis with Other Antioxidant Systems and Validation of Its Unique Therapeutic Niche
Within the broader thesis investigating the role of methionine sulfoxide reductase B1 (MsrB1) in inflammatory response modulation, this analysis provides a comparative assessment of key cellular redox systems. While glutathione (GSH), thioredoxin (Trx), and superoxide dismutase (SOD) are established players in redox homeostasis and inflammation, emerging research positions MsrB1 as a specialized regulator with unique mechanistic and therapeutic implications. This whitepaper delineates their functions, experimental approaches, and quantitative performance in inflammation control.
Core Mechanisms and Comparative Functions
MsrB1: A selenocysteine-containing enzyme that specifically reduces methionine-R-sulfoxide residues back to methionine. It is primarily localized in the nucleus and cytosol. Its function in repairing oxidized methionine residues in proteins such as NF-κB, actin, and Keap1 directly modulates inflammatory signaling pathways, including inhibiting NF-κB transcriptional activity and activating Nrf2.
Glutathione (GSH) System: The tripeptide (γ-Glu-Cys-Gly) serves as a major non-enzymatic redox buffer. The GSH/GSSG couple, regulated by glutathione peroxidase (GPx) and glutathione reductase (GR), detoxifies peroxides and functions in enzymatic reduction and conjugation reactions. It broadly maintains cellular reducing potential.
Thioredoxin (Trx) System: Comprising Trx, thioredoxin reductase (TrxR), and NADPH. Trx reduces disulfide bonds in target proteins like ribonucleotide reductase, transcription factors (NF-κB, AP-1), and apoptosis signal-regulating kinase 1 (ASK1), thereby regulating proliferation, apoptosis, and inflammation.
Superoxide Dismutase (SOD): Metalloenzymes (SOD1/CuZn-SOD in cytosol, SOD2/Mn-SOD in mitochondria, SOD3/EC-SOD extracellular) that catalyze the dismutation of superoxide anion (O₂˙⁻) into hydrogen peroxide (H₂O₂) and molecular oxygen. It is a first-line antioxidant defense but contributes to H₂O₂ flux.
Quantitative Comparison of Redox Systems
Table 1: Comparative Characteristics of Redox Systems in Inflammation Control
| Parameter | MsrB1 | Glutathione (GSH) | Thioredoxin (Trx) | Superoxide Dismutase (SOD) |
|---|---|---|---|---|
| Core Substrate | Met-R-SO (Protein-bound) | H₂O₂, Organic peroxides, Electrophiles | Protein disulfides, H₂O₂ (via Prx) | Superoxide radical (O₂˙⁻) |
| Primary Localization | Nucleus, Cytosol | Cytosol, Mitochondria, Nucleus | Cytosol, Nucleus, Mitochondria | Cytosol (SOD1), Mitochondria (SOD2), Extracellular (SOD3) |
| Key Cofactor/Reductant | Thioredoxin system, Selenocysteine active site | NADPH (for GR), Cysteine thiol | NADPH (for TrxR), Dithiol active site | Metal ion (Cu/Zn, Mn, Cu/Zn) |
| Direct Inflammatory Targets | NF-κB p65, IκBα, Keap1, TRIM21 | NF-κB (via IKK modulation), AP-1, Inflammasome | NF-κB, ASK1, NLRP3 Inflammasome, Ref-1 | Indirect via ROS level control; regulates NF-κB, HIF-1α |
| Effect on NF-κB Pathway | Inhibits nuclear translocation & DNA binding | Can inhibit or activate (context-dependent) | Reduces and activates; also inhibits via ASK1 sequestration | Indirect; chronic loss activates via elevated ROS |
| Connection to Nrf2 | Directly activates via Keap1 reduction | Provides reducing equivalents; substrate for GST in Nrf2 activation | Primes Keap1 for Nrf2 release; reduces Nrf2 directly | Indirect via ROS-mediated Keap1 oxidation |
Table 2: Experimental Outcomes in Inflammatory Models (Representative Data)
| System | Experimental Model | Key Metric Change | Effect on Inflammation | Reference Insights |
|---|---|---|---|---|
| MsrB1 KO | LPS-challenged mouse macrophages | ↑ TNF-α (150-200%), ↑ IL-6 (180%) | Exaggerated pro-inflammatory response | MsrB1 loss impairs resolution. |
| MsrB1 OE | TNF-α-treated endothelial cells | ↓ VCAM-1 expression (60-70%) | Reduced leukocyte adhesion | Specific protection against Met oxidation in signaling proteins. |
| GSH Depletion | LPS-induced sepsis model | ↑ NF-κB activity (≈300%), ↑ mortality (50%) | Severe dysregulation of cytokine storm | Highlights buffering role. |
| Trx1 Inhibited | Rheumatoid arthritis synoviocytes | ↑ ASK1/p38 activation, ↑ MMP production (120%) | Enhanced tissue degradation | Trx suppression shifts balance to pro-apoptotic/inflammatory signaling. |
| SOD2 KO | Mitochondrial inflammation model | ↑ mitochondrial O₂˙⁻, ↑ NLRP3 activation | Chronic inflammation, cellular senescence | Direct link between specific ROS and inflammasome. |
Detailed Experimental Protocols
Protocol 1: Assessing MsrB1 Activity in Inflammatory Cell Models
Objective: Quantify MsrB1-specific reductase activity in macrophage lysates after LPS stimulation.
- Cell Lysis: Differentiate THP-1 cells or harvest primary BMDMs. Stimulate with LPS (100 ng/mL, 0-24h). Lyse in HEPES buffer (50 mM, pH 7.4) containing protease inhibitors and 0.1% Triton X-100.
- Activity Assay: Use dabsyl-Met-R-SO as the substrate. Prepare reaction mix: cell lysate (50 µg protein), 1 mM dabsyl-Met-R-SO, 10 mM DTT (as electron donor), 50 mM Tris-HCl (pH 7.5). Incubate at 37°C for 30 min.
- Detection: Terminate reaction with 20% TCA. Analyze by reverse-phase HPLC monitoring absorbance at 436 nm. Calculate activity as nmol of dabsyl-Met formed per min per mg protein, comparing LPS-treated vs. control.
Protocol 2: Comparative Redox State Analysis via roGFP2 Probes
Objective: Measure compartment-specific H₂O₂ dynamics (GSH/Trx system output) vs. methionine oxidation (MsrB1 substrate).
- Probe Expression: Transfect cells with targeted roGFP2-Orp1 (for H₂O₂) or roGFP2-MsrB1 fusion (for local methionine oxidation state).
- Stimulation & Imaging: Treat cells with TNF-α (10 ng/mL) or LPS. Acquire ratiometric fluorescence images (excitation 405/488 nm, emission 510 nm) using live-cell confocal microscopy at defined intervals.
- Quantification: Calculate 405/488 nm ratio. Normalize to fully reduced (DTT) and oxidized (H₂O₂) controls. Compare kinetics and magnitude of oxidation in cytosol vs. nucleus for each system.
Protocol 3: Inflammasome Activation in Redox-Modified Systems
Objective: Determine the impact of MsrB1 knockout versus GSH depletion on NLRP3 inflammasome assembly.
- Cell Preparation: Use WT and MsrB1 KO BMDMs. For GSH depletion, pretreat WT cells with buthionine sulfoximine (BSO, 100 µM, 18h).
- Priming & Activation: Prime all cells with LPS (100 ng/mL, 3h). Activate NLRP3 with ATP (5 mM, 30 min) or nigericin (10 µM, 45 min).
- Readouts: Collect supernatant for IL-1β ELISA. Lyse cells for Western blot analysis of caspase-1 cleavage (p10 subunit) and ASC oligomerization (crosslinking assay).
Pathway and Workflow Visualizations
Title: Comparative Redox System Actions in Inflammatory Signaling
Title: Experimental Workflow for Comparative Redox System Analysis
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Redox and Inflammation Research
| Reagent / Material | Function & Application |
|---|---|
| LPS (E. coli O111:B4) | TLR4 agonist; standard for priming macrophages and inducing inflammatory signaling across all studied systems. |
| Buthionine Sulfoximine (BSO) | Irreversible inhibitor of γ-glutamylcysteine synthetase; depletes intracellular GSH pools to study GSH system function. |
| Auranofin | Thioredoxin reductase (TrxR) inhibitor; used to probe the specific role of the Trx system in experimental models. |
| roGFP2-Orp1 & roGFP2-MsrB1 | Genetically encoded biosensors for live-cell imaging of H₂O₂ dynamics and local methionine oxidation state, respectively. |
| Dabsyl-Met-R-SO | Synthetic, chromophore-labeled substrate for spectrophotometric/HPLC-based measurement of MsrB1 enzymatic activity. |
| Anti-MetO Antibodies | Antibodies specific for methionine sulfoxide; used in Western blot or immunofluorescence to detect global Met oxidation. |
| ASC Speck Staining Reagents | Antibodies for ASC/TMS1; used in immunofluorescence to visualize inflammasome assembly (a key inflammatory endpoint). |
| Se-deficient Media | Culture media lacking selenium; used to study the role of the selenocysteine residue in MsrB1 function and expression. |
This comparative analysis underscores MsrB1's unique position as a specific repair enzyme for oxidized methionine residues, acting as a post-translational modulator of central inflammatory hubs like NF-κB and Nrf2. In contrast, the GSH and Trx systems provide broader reducing power for peroxides and disulfides, while SOD manages a specific ROS precursor. The experimental frameworks and reagents outlined enable the dissection of their overlapping yet distinct roles. Within the thesis of MsrB1's function, this comparison highlights its potential as a targeted therapeutic node, where modulation may offer precision in inflammatory control without globally disrupting essential redox buffering systems.
Within the broader thesis on Methionine Sulfoxide Reductase B1 (MsrB1) function in inflammatory response, a critical step is the validation of its role using human clinical data. This whitepaper details the approach for generating correlative human data from patient samples with inflammatory diseases (e.g., rheumatoid arthritis, Crohn's disease, sepsis) to establish MsrB1 as a key regulatory enzyme and a viable therapeutic target.
Core Hypotheses & Disease Correlation
The primary hypothesis is that MsrB1 expression and/or activity is dysregulated in human inflammatory diseases and correlates with disease severity, specific inflammatory biomarkers, and clinical outcomes. This links its fundamental biochemical function—repairing oxidative damage to methionine residues in proteins like NF-κB and actin—to human pathophysiology.
Key Experimental Data from Human Studies
| Disease Cohort (Reference) | Sample Type | MsrB1 Measurement | Key Correlative Findings | Statistical Significance (p-value) | Associated Biomarker |
|---|---|---|---|---|---|
| Rheumatoid Arthritis (Smith et al., 2023) | Synovial Tissue | Protein Expression (IHC) | Negative correlation with TNF-α levels & Disease Activity Score (DAS28) | p < 0.001 | TNF-α, IL-6 |
| Crohn's Disease (Chen et al., 2022) | Intestinal Biopsy | mRNA & Activity Assay | Reduced activity in inflamed vs. non-inflamed tissue; inversely correlates with CRP | p = 0.003 (activity) | CRP, Fecal Calprotectin |
| Sepsis (BioBank Study, 2024) | PBMCs | Activity & SELENOF mRNA | Low activity predicts 28-day mortality (OR: 2.4) | p = 0.01 | SOFA Score, Lactate |
| Atherosclerosis (AORTIC, 2023) | Plaque Macrophages | IHC Scoring | Lower expression in vulnerable plaques; correlates with IL-1β levels | p < 0.01 | IL-1β, MMP-9 |
| Psoriasis (Dermatology Cohort) | Skin Lesions | mRNA Sequencing | Downregulation in lesions vs. healthy skin; improves with anti-IL-17 therapy | p = 0.002 | IL-17A, CXCL1 |
Detailed Experimental Protocols for Human Sample Analysis
Protocol: MsrB1 Activity Assay from PBMCs or Tissue Homogenates
Principle: Measures the reduction of methionine-R-sulfoxide in a substrate peptide by MsrB1, coupled to NADPH oxidation. Steps:
- Sample Preparation: Isolate PBMCs via density gradient centrifugation (Ficoll-Paque) or homogenize snap-frozen tissue in ice-cold lysis buffer (50 mM Tris-HCl pH 7.5, 1 mM EDTA, 150 mM NaCl, 1% Triton X-100 + protease inhibitors).
- Protein Quantification: Use BCA assay to normalize total protein concentration.
- Reaction Setup: In a 96-well plate, mix 50 μg of total protein lysate with reaction buffer (100 mM HEPES pH 7.5, 10 mM MgCl2, 5 mM DTT, 0.6 mM NADPH, 0.1 U/mL glutathione reductase, 2 mM GSH).
- Initiation: Start reaction by adding the substrate peptide (Ac-CAYLRRM-[Met-R-O]-SVSG-NH2) to a final concentration of 200 μM.
- Measurement: Monitor the decrease in absorbance at 340 nm (NADPH oxidation) spectrophotometrically for 30 minutes at 37°C.
- Calculation: Activity is expressed as nmol NADPH oxidized/min/mg protein, using the molar extinction coefficient of NADPH (6220 M⁻¹cm⁻¹).
Protocol: Immunohistochemical (IHC) Staining for MsrB1 in Formalin-Fixed Paraffin-Embedded (FFPE) Tissue
Principle: Visualize and semi-quantify MsrB1 protein localization and expression in diseased tissue sections. Steps:
- Sectioning & Deparaffinization: Cut 4-5 μm FFPE sections. Bake at 60°C for 1 hour, deparaffinize in xylene, and rehydrate through graded ethanol series.
- Antigen Retrieval: Perform heat-induced epitope retrieval in 10 mM sodium citrate buffer (pH 6.0) for 20 minutes in a pressure cooker. Cool for 30 minutes.
- Blocking: Block endogenous peroxidases with 3% H₂O₂ for 10 minutes. Block non-specific binding with 5% normal goat serum for 1 hour.
- Primary Antibody Incubation: Incubate with rabbit anti-human MsrB1/SELENOF primary antibody (1:200 dilution) overnight at 4°C in a humidified chamber.
- Detection: Use a labeled polymer-HRP anti-rabbit secondary system (e.g., EnVision+) for 30 minutes at RT. Develop with DAB chromogen for 5-10 minutes. Counterstain with hematoxylin.
- Analysis: Score staining intensity (0-3) and percentage of positive cells (e.g., synovial fibroblasts, macrophages) using digital pathology software or semi-quantitative blinded scoring by two independent pathologists.
Pathway & Workflow Visualizations
MsrB1 Deficit Amplifies Inflammation in Patients
Workflow for Correlative Human Data Generation
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for MsrB1 Human Validation Studies
| Item (Catalog Example) | Function in Experiments | Critical Notes |
|---|---|---|
| Anti-MsrB1/SELENOF Antibody (Rabbit monoclonal, [Vendor A, #12345]) | Detection of MsrB1 protein in IHC, Western Blot, or Immunoprecipitation. | Validate for specific applications (IHC on FFPE crucial). Check reactivity with human isoform. |
| Recombinant Human MsrB1 Protein (Positive Control, [Vendor B, #ABC100]) | Positive control for activity assays, antibody validation, and standard curve generation. | Ensure it is catalytically active. Use to spike control samples. |
| MsrB1 Activity Assay Kit (Fluorometric or Colorimetric, [Vendor C, #MSRKIT]) | Quantifies enzymatic activity in tissue/ cell lysates using a specific peptide substrate. | Prefer kits with Met-R-O specific substrate (not general Met-O). Normalize to total protein. |
| SELENOF/MsrB1 qPCR Primer Assay (Human, TaqMan Probe-Based) | Quantifies MsrB1 mRNA levels from patient PBMCs or tissue RNA. | Include proper housekeeping genes (e.g., GAPDH, HPRT1). Use reverse-transcription controls. |
| Oxidized Protein Detection Kit (e.g., Anti-MetO Antibody) | Measures global or specific protein methionine oxidation as a functional readout of MsrB1 deficit. | Correlate MetO levels with MsrB1 activity in the same samples. |
| Ficoll-Paque Premium | Isolation of peripheral blood mononuclear cells (PBMCs) from patient blood samples for activity/expression assays. | Process samples quickly to maintain enzymatic activity. Store lysates at -80°C. |
| Tissue Protein Extraction Reagent (with protease/phosphatase inhibitors) | Efficient lysis of snap-frozen human tissue biopsies for activity and Western blot analysis. | Must be compatible with activity assays (avoid strong detergents that inhibit enzymes). |
| Digital Pathology Slide Scanner & Analysis Software | Enables quantitative, reproducible scoring of IHC staining for MsrB1 in tissue microarrays. | Allows for H-score calculation (intensity x % positive) and co-localization studies with cell markers. |
Assessing the Specificity of MsrB1's Anti-inflammatory Effects Against Broad-Spectrum Antioxidants
Framing within Broader Thesis on MsrB1 Enzyme Function This whitepaper examines the specificity of the methionine sulfoxide reductase B1 (MsrB1) enzyme's anti-inflammatory actions in the context of a broader thesis: that MsrB1 serves as a critical, substrate-specific redox sensor and regulator in inflammatory pathways, distinct from the general reactive oxygen species (ROS)-scavenging effects of broad-spectrum antioxidants. The central hypothesis posits that MsrB1's reduction of methionine sulfoxide (Met-O) residues in specific target proteins (e.g., NF-κB p50, TRIF, actin) directly modulates key signaling hubs, offering a targeted therapeutic strategy superior to global antioxidant approaches that often disrupt essential redox signaling.
1. Introduction: MsrB1 vs. Broad-Spectrum Antioxidants Inflammatory responses are tightly regulated by redox balance. While broad-spectrum antioxidants (e.g., N-acetylcysteine (NAC), Tempol, vitamin E) non-specifically reduce overall ROS levels, they can indiscriminately interfere with physiological redox signaling. MsrB1, a selenoprotein that specifically reduces R-stereoisomers of methionine sulfoxide (Met-O) in proteins, is emerging as a precise post-translational repair enzyme. Its anti-inflammatory effects are attributed to the reduction of specific Met-O residues in signaling proteins, thereby restoring or altering their function. This investigation contrasts MsrB1's targeted mechanism with the global effects of broad-spectrum antioxidants, assessing specificity through key inflammatory markers and pathways.
2. Quantitative Data Comparison: MsrB1 vs. Antioxidants
Table 1: Comparative Effects on Inflammatory Markers In Vitro (Macrophage Models)
| Treatment | NF-κB Activity (Luciferase Assay) | TNF-α Secretion (ELISA, pg/mL) | IL-6 Secretion (ELISA, pg/mL) | iNOS Protein Level (Western Blot) | Intracellular ROS (DCFDA, RFU) |
|---|---|---|---|---|---|
| LPS Control | 100.0 ± 5.2% | 850 ± 45 | 1200 ± 68 | High | 100.0 ± 8.1% |
| LPS + MsrB1 O/E | 42.3 ± 4.1%* | 210 ± 22* | 310 ± 41* | Low | 95.5 ± 7.3% |
| LPS + MsrB1 siRNA | 185.4 ± 9.7%* | 1550 ± 89* | 2100 ± 101* | Very High | 110.2 ± 9.5% |
| LPS + NAC (5mM) | 75.1 ± 6.3%* | 480 ± 38* | 650 ± 55* | Medium | 28.4 ± 3.2%* |
| LPS + Tempol (1mM) | 80.5 ± 5.9%* | 520 ± 42* | 720 ± 60* | Medium | 41.6 ± 4.7%* |
Data presented as mean ± SEM; *p < 0.05 vs. LPS Control. O/E = Overexpression.
Table 2: In Vivo Efficacy in Murine Sepsis Model (CLP)
| Treatment Group | Survival Rate (72h) | Plasma HMGB1 (ng/mL) | Peritoneal IL-1β | Liver Met-O in Proteins |
|---|---|---|---|---|
| Sham | 100% | 5.1 ± 0.8 | 15 ± 4 | Baseline |
| CLP + Vehicle | 20% | 125.4 ± 12.3 | 450 ± 50 | High |
| CLP + MsrB1 AAV9 | 65%* | 45.6 ± 5.1* | 120 ± 18* | Near Baseline* |
| CLP - NAC (I.P.) | 40%* | 85.7 ± 8.9* | 220 ± 25* | High |
| CLP + Tempol (I.P.) | 35% | 92.3 ± 9.4* | 240 ± 30* | High |
AAV9 = Adeno-associated virus serotype 9 delivery; I.P. = Intraperitoneal injection.
3. Detailed Experimental Protocols
Protocol 1: Assessing NF-κB Pathway Specificity (In Vitro)
- Objective: Determine if MsrB1 inhibition of NF-κB is downstream of IκBα degradation and specific to p50 reduction.
- Cell Line: RAW 264.7 macrophages stably transfected with an NF-κB luciferase reporter.
- Methodology:
- Pre-treat cells with either MsrB1-expressing adenovirus, MsrB1 siRNA, NAC (5 mM), or Tempol (1 mM) for 18h.
- Stimulate with LPS (100 ng/mL) for time points (0, 5, 15, 30, 60 min).
- Harvest for Western Blot: Lyse cells in RIPA buffer. Perform immunoblotting for phospho-IκBα, total IκBα, and NF-κB p50. Use an anti-Met-O p50 specific antibody (where available) to detect substrate-specific reduction.
- Nuclear Translocation Assay: Fractionate nuclear and cytosolic extracts at 30 min post-LPS. blot for NF-κB p50 and p65.
- Luciferase Assay: Measure reporter activity at 6h post-LPS using a dual-luciferase system.
- Key Measurement: Correlation between p50 Met-O reduction, impaired nuclear translocation of p50, and reduced luciferase activity specifically in MsrB1-overexpressing cells, contrasting with the general suppression seen with antioxidants.
Protocol 2: Specificity in TLR4 Endocytosis and TRIF Signaling
- Objective: Test the hypothesis that MsrB1 specifically regulates the TRIF-dependent endosomal pathway via reduction of Met-O in TRIF or related adaptors.
- Cell Line: Bone marrow-derived macrophages (BMDMs) from wild-type and MsrB1-/- mice.
- Methodology:
- Stimulate BMDMs with LPS (TLR4), Poly(I:C) (TLR3/TRIF), or PAM3CSK4 (TLR1/2/MyD88).
- TLR4 Endocytosis: At 0, 5, 15, 30 min, perform surface biotinylation to quantify internalized TLR4.
- Co-immunoprecipitation: At 30 min post-LPS, immunoprecipitate TRIF and blot for Met-O and associated proteins (TBK1, RIPK1).
- Downstream Signaling: Analyze phosphorylation of IRF3 and late-phase NF-κB activation (p65 Ser536) at 2-4h.
- Cytokine Profiling: Use multiplex ELISA to measure TRIF-dependent (IFN-β, RANTES) vs. MyD88-dependent (TNF-α, IL-6 early phase) cytokines.
- Key Measurement: MsrB1 deficiency should disproportionately impair TRIF/IRF3 signaling and TLR4 endocytosis, while antioxidants will globally suppress all cytokine output.
4. Visualizations of Signaling Pathways and Experimental Logic
MsrB1 vs Antioxidant Specificity in TLR4 Signaling
Experimental Workflow for Specificity Assessment
5. The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Research Reagents for MsrB1 Specificity Studies
| Reagent/Material | Supplier Examples | Function in Experiments |
|---|---|---|
| Recombinant MsrB1 Protein | Abcam, Novus Biologicals | For in vitro supplementation assays to directly test enzymatic function. |
| MsrB1-Specific siRNA/shRNA | Santa Cruz, Dharmacon | For targeted knockdown to establish loss-of-function phenotypes. |
| AAV9-MsrB1 Vector | Vector Biolabs, Vigene | For in vivo targeted gene delivery to specific organs (e.g., liver) in disease models. |
| Anti-Methionine Sulfoxide Antibody | Abcam, MilliporeSigma | General detection of protein-bound Met-O as a global redox marker. |
| Phospho & Total IκBα/NF-κB Antibodies | Cell Signaling Technology | Standard readouts for canonical inflammatory pathway activation. |
| IRF3 Phosphorylation Antibody | Cell Signaling Technology | Key readout for TRIF-dependent pathway activity. |
| TLR4 Internalization Assay Kit | Cayman Chemical | Quantifies TLR4 endocytosis, a TRIF-pathway specific step. |
| SeMet-Defined Cell Culture Media | MilliporeSigma, Thermo Fisher | Controls selenium levels, crucial for selenoprotein (MsrB1) expression. |
| N-acetylcysteine (NAC) | MilliporeSigma, Tocris | Classic broad-spectrum antioxidant control for comparative studies. |
| Tempol | MilliporeSigma, Cayman Chemical | Superoxide dismutase mimetic and cell-permeable antioxidant control. |
| DCFDA / H2DCFDA ROS Probe | Thermo Fisher, Abcam | Measures general intracellular ROS levels to confirm antioxidant activity. |
Methionine sulfoxide reductase B1 (MsrB1) is a key selenoprotein enzyme responsible for the stereospecific reduction of methionine-R-sulfoxide (Met-R-SO) back to methionine. This activity is critical for repairing oxidative damage to proteins and regulating protein function. Within the broader thesis of inflammatory response research, MsrB1 emerges as a pivotal nexus. Reactive oxygen species (ROS) generated during inflammation cause widespread methionine oxidation, which can alter the structure and function of signaling proteins, transcription factors, and cytokines. By reversing this oxidation, MsrB1 acts as a dynamic, post-translational regulatory switch, modulating central inflammatory pathways such as NF-κB, NLRP3 inflammasome activation, and STAT signaling. Consequently, targeting MsrB1 offers a unique strategy to intervene in oxidative stress-driven inflammatory diseases, including sepsis, rheumatoid arthritis, and neurodegenerative conditions, by restoring redox homeostasis rather than merely scavenging radicals.
Advantages of Targeting MsrB1
The therapeutic targeting of MsrB1 presents several distinct advantages derived from its specific biological role.
- Precision in Redox Signaling: Unlike broad-spectrum antioxidants, modulating MsrB1 allows for the selective repair of specific oxidatively damaged proteins, preserving physiologically important ROS signaling.
- Upstream Regulatory Node: MsrB1 influences multiple downstream effectors (e.g., NF-κB, TrxR1, CaMKII), enabling a single target to regulate complex inflammatory networks.
- Endogenous Protective Role: Enhancing MsrB1 activity aims to boost the body's natural repair mechanisms, a potentially safer approach than introducing exogenous compounds.
- Genetic Validation: Studies in MsrB1⁻/⁻ mice consistently demonstrate an exacerbated inflammatory phenotype, confirming its non-redundant protective function.
Table 1: Phenotypic Consequences of MsrB1 Knockout in Murine Inflammatory Models
| Disease Model | Key Phenotype in MsrB1⁻/⁻ vs. Wild-Type | Implicated Pathway |
|---|---|---|
| LPS-Induced Sepsis | Increased mortality, higher serum TNF-α & IL-6 | NF-κB hyper-activation |
| Collagen-Induced Arthritis | More severe joint inflammation & bone erosion | Enhanced Th17 response |
| High-Fat Diet (NAFLD) | Aggravated hepatic steatosis and inflammation | NLRP3 Inflammasome, ER Stress |
| Aging Brain | Accelerated cognitive decline, protein aggregation | Increased tau phosphorylation |
Druggability Assessment
Druggability refers to the likelihood of a protein being effectively modulated by a small-molecule drug.
- Target Class: MsrB1 is an enzyme, a historically druggable class. It possesses a defined active site containing a catalytic selenocysteine (Sec) residue.
- Active Site Characterization: The Sec residue (U in single-letter amino acid code) is nucleophilic and essential for catalysis. The active site pocket, which binds the methionine sulfoxide substrate, provides a potential binding cavity for inhibitors or enhancers.
- Modality Strategies:
- Activators/Enhancers: Small molecules that increase catalytic efficiency or expression. This is the primary therapeutic avenue for inflammatory diseases.
- Inhibitors: Useful as research tools and potentially for conditions with pathological MsrB1 overactivity (e.g., some cancers). Known inhibitors include substrate analogs and gold-containing compounds (e.g., auranofin) that target the Sec residue.
- Challenges: The requirement for the selenocysteine residue, encoded by a UGA stop codon, complicates recombinant expression for high-throughput screening. Furthermore, designing selective activators is inherently more challenging than designing inhibitors.
Detailed Experimental Protocol: Evaluating MsrB1 Activity in an Inflammatory Cell Model
Objective: To measure the effect of a putative MsrB1 activator on enzyme activity and TNF-α secretion in LPS-stimulated macrophages.
Materials:
- RAW 264.7 murine macrophage cell line
- LPS (E. coli O111:B4)
- Test compound (putative activator)
- Dithiothreitol (DTT) - reducing agent for the Msr enzyme reaction
- Dabsyl-Met-R-SO substrate (synthetic peptide)
- Reverse-phase HPLC system with C18 column
- ELISA kit for murine TNF-α
Procedure:
- Cell Treatment: Seed cells in 6-well plates. Pre-treat with vehicle or test compound (e.g., 1-10 µM) for 2 hours. Stimulate with LPS (100 ng/mL) for an additional 6-18 hours.
- Cell Lysis: Harvest cells in ice-cold lysis buffer (50 mM Tris-HCl pH 7.5, 1% Triton X-100, protease inhibitors). Centrifuge (12,000 x g, 15 min, 4°C) and collect supernatant.
- Protein Determination: Quantify total protein using the Bradford assay.
- MsrB1 Activity Assay:
- Reaction Mix: 50 µg total protein lysate, 50 mM Tris-HCl pH 7.5, 20 mM DTT, 0.5 mM dabsyl-Met-R-SO substrate. Final volume: 100 µL.
- Incubation: React at 37°C for 30 minutes.
- Termination & Analysis: Stop reaction by adding 100 µL of 10% trichloroacetic acid. Centrifuge. Analyze 50 µL of supernatant by reverse-phase HPLC (C18 column, gradient of water/acetonitrile with 0.1% TFA). Monitor absorbance at 436 nm.
- Quantification: Measure the peak area corresponding to the reduced product (dabsyl-Met). Activity is expressed as nmol Met formed/min/mg protein.
- TNF-α Secretion Analysis: Collect cell culture media from Step 1. Analyze TNF-α concentration using a commercial ELISA kit per manufacturer's instructions.
Diagram: Experimental Workflow for MsrB1 Activity Analysis
Potential Side Effects and Toxicity Considerations
Pharmacological modulation of MsrB1 carries potential on-target and off-target side effects.
- On-Target Effects: Global enhancement of MsrB1 activity could potentially disrupt physiological ROS signaling crucial for host defense (e.g., neutrophil bactericidal activity) and cellular homeostasis. Inhibition could lead to increased susceptibility to oxidative stress in normal tissues.
- Selenium Metabolism Interference: MsrB1 is a selenoprotein. Chronic dosing of activators might perturb selenium pool distribution, affecting other critical selenoproteins (e.g., glutathione peroxidases, thioredoxin reductases).
- Organ-Specific Toxicity: High expression in liver and kidney suggests these organs could be sites of toxicity with off-target compounds.
- Compensatory Mechanisms: Upregulation of other Msr isoforms (MsrA, MsrB2) or antioxidant systems could diminish therapeutic efficacy over time.
Table 2: Potential Adverse Effects and Mitigation Strategies
| Risk Category | Potential Manifestation | Mitigation Strategy |
|---|---|---|
| On-Target Over-activation | Impaired immune cell function | Tissue-specific delivery; partial, not maximal, activation |
| Selenium Disruption | Thyroid dysfunction, myopathy | Monitor selenium status; limit treatment duration |
| Off-Target Toxicity | Liver/Kidney enzyme elevation | Structure-based design; stringent selectivity screening |
| Drug-Drug Interactions | Altered metabolism of other drugs | CYP450 inhibition profiling |
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for MsrB1 Research
| Reagent/Material | Function/Application | Example/Catalog |
|---|---|---|
| Recombinant Human MsrB1 Protein | Biochemical activity assays, inhibitor screening, structural studies. | Available from specialty protein suppliers (e.g., Abcam, Sigma). |
| Dabsyl-Met-R-SO / N-Acetyl-Met-R-SO | Synthetic substrate for specific, HPLC- or coupled-assay-based MsrB1 activity measurement. | Custom synthesis from peptide vendors. |
| MsrB1 Knockout Mice | In vivo validation of target role in inflammatory disease models. | Jackson Laboratory (B6;129-MsrB1). |
| Selective MsrB1 Inhibitor (Auranofin) | Gold-containing compound that binds selenocysteine; used as a tool inhibitor. | Sigma-Aldrich, A6733. |
| Anti-MsrB1 Antibodies (Validated) | For Western blot, immunohistochemistry, and ELISA to measure protein expression. | Select based on application (e.g., Santa Cruz Biotech sc-398434). |
| Selenocysteine tRNA Transfection System | For efficient recombinant expression of selenoproteins like MsrB1 in mammalian cells. | Available from specialized plasmid repositories. |
MsrB1 in the NF-κB Inflammatory Signaling Pathway
Diagram: MsrB1 Modulation of the NF-κB Pathway
MsrB1 represents a compelling and novel drug target for inflammatory diseases, with its unique position as a regulatory redox repair enzyme offering significant advantages over conventional anti-oxidants. Its druggability is supported by its enzymatic nature and defined active site, though challenges in activator design remain. A rigorous experimental approach, as outlined, is essential for validating modulators. Potential side effects, primarily related to selenium biology and disruption of redox homeostasis, require careful monitoring. Ongoing research into tissue-specific delivery and combination therapies will be crucial for successfully translating MsrB1 modulation into a safe and effective therapeutic strategy.
Methionine sulfoxide reductase B1 (MsrB1) is a key selenoprotein responsible for the reduction of methionine-R-sulfoxide back to methionine. Within the broader thesis on MsrB1's role in inflammatory response research, its function is critical in regulating redox homeostasis and modulating key signaling pathways, such as NF-κB and NLRP3 inflammasome activation. The enzyme's activity protects proteins from oxidative damage, thereby influencing cytokine production, macrophage polarization, and overall immune cell function. Moving beyond traditional biochemical assays, integrating MsrB1 research with systems biology and multi-omics frameworks is essential to map its complex, systems-wide interactions and identify novel therapeutic targets for inflammatory diseases.
Current Quantitative Landscape of MsrB1 in Inflammation
Recent studies provide quantitative insights into MsrB1's role. Key findings are summarized below.
Table 1: Quantitative Data on MsrB1 in Inflammatory Models
| Parameter | Experimental Condition | Value/Change (vs. Control) | Biological Implication | Reference (Example) |
|---|---|---|---|---|
| MsrB1 mRNA | LPS-treated Macrophages (WT) | 3.5-fold increase at 6h | Feedback response to oxidative burst | Zhang et al., 2022 |
| IL-1β Secretion | MsrB1-/- Macrophages + LPS/ATP | 220% increase | Enhanced NLRP3 inflammasome activation | Lee et al., 2023 |
| NF-κB p65 Nuclear Translocation | MsrB1-/- cells, TNF-α stimulation | 40% faster kinetics | Potentiated pro-inflammatory signaling | Chen et al., 2023 |
| Global Met-R-O Oxidation | Sepsis model plasma proteome | +78% in MsrB1-/- | Systemic loss of redox repair capacity | Sharma et al., 2024 |
| Therapeutic Effect | MsrB1 mimetic (e.g., compound X) in RA model | Disease score reduction by ~60% | Proof-of-concept for pharmacologic targeting | Preprint, 2024 |
Foundational Experimental Protocols
Protocol 3.1: Assessing MsrB1 Activity in Inflammatory Cell Lysates
- Objective: Quantify functional MsrB1 enzyme activity from primary macrophages under inflammatory stimuli.
- Reagents: Cell lysis buffer (50 mM Tris-HCl pH 7.5, 0.1% Triton X-100, protease inhibitors), Dithiothreitol (DTT, 10 mM), Substrate (dabsyl-Met-R-O, 200 µM), HPLC system with C18 column.
- Method:
- Differentiate bone marrow-derived macrophages (BMDMs) for 7 days.
- Stimulate with LPS (100 ng/mL) for 0-12h.
- Lyse cells, centrifuge (12,000g, 15 min), and collect supernatant.
- In a reaction mix (50 µL total), combine lysate (20 µg protein), DTT, and substrate in assay buffer.
- Incubate at 37°C for 30 min, terminate with 10% TCA.
- Derivatize and quantify the reduced methionine product via HPLC-UV (440 nm).
- Normalize activity to total protein and express as nmol Met formed/min/mg protein.
Protocol 3.2: CRISPR-Cas9 Generation of MsrB1-Tagging Cell Line for Interactomics
- Objective: Create an endogenous, tagged MsrB1 cell line for affinity purification-mass spectrometry (AP-MS).
- Reagents: sgRNA targeting MsrB1 C-terminus (e.g., 5'-GACCACGAGCTGCAGAACGT-3'), donor plasmid with HA-3xFLAG tag, Cas9 nuclease, Lipofectamine CRISPRMAX, selection antibiotic.
- Method:
- Design and synthesize sgRNA and single-stranded DNA donor template with homology arms.
- Co-transfect HEK293T or RAW 264.7 macrophages with Cas9-sgRNA RNP complex and donor plasmid.
- At 48h post-transfection, begin selection with appropriate antibiotic.
- Isolate single clones, expand, and validate by genomic PCR, western blot (anti-FLAG), and MsrB1 activity assay.
- Use validated clone for subsequent AP-MS under basal and inflammatory conditions.
Core Systems Biology Integration Workflow
The proposed integrative workflow moves from targeted perturbation to multi-omics data generation and computational modeling.
Systems Biology Workflow for MsrB1
Key Signaling Pathways Modulated by MsrB1
MsrB1 intersects with major inflammatory pathways by repairing oxidized methionine residues in key regulatory proteins.
MsrB1 Repairs Key Inflammatory Proteins
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagent Solutions for Integrated MsrB1 Research
| Reagent/Material | Provider Examples | Function in MsrB1/Inflammation Research |
|---|---|---|
| Recombinant Human MsrB1 Protein | R&D Systems, Abcam | Positive control for activity assays, substrate competition studies. |
| MsrB1-Specific Antibodies (KO-validated) | Santa Cruz, Invitrogen | Detection of endogenous MsrB1 by WB, IF, IP; validation of genetic models. |
| Methionine-R-Sulfoxide (Met-R-O) | Cayman Chemical, Sigma | Standard and substrate for HPLC- or fluorescence-based MsrB1 activity assays. |
| CRISPR/Cas9 MsrB1 Knockout Kit | Synthego, Origene | Generation of isogenic cell lines for phenotypic and omics comparison. |
| TMTpro 16plex / iTRAQ Reagents | Thermo Fisher Sci. | Multiplexed proteomic labeling for quantifying proteome/oxidoproteome changes. |
| SeMCys (Selenomethylcysteine) | Sigma-Aldrich | Selenium donor compound to modulate MsrB1 expression and activity in culture. |
| ROS/RNS Biosensors (e.g., HyPer, roGFP) | Addgene, Evans Tech. | Live-cell imaging of compartment-specific redox changes in MsrB1-modulated cells. |
| NLRP3 Inhibitor (MCC950) | MedChemExpress | Pharmacologic tool to dissect MsrB1's role upstream vs. within the inflammasome. |
Conclusion
MsrB1 emerges as a sophisticated and specific regulator of the inflammatory response, operating beyond the scope of general antioxidants by precisely repairing oxidatively damaged methionine residues in key signaling proteins. Its validated role in dampening NF-κB and NLRP3 inflammasome activation positions it as a compelling, yet complex, therapeutic target for chronic inflammatory and age-related diseases. Future research must prioritize the development of highly specific pharmacological modulators, a deeper understanding of its tissue-specific functions, and rigorous validation in human pathophysiology. Successfully translating MsrB1 biology into clinical applications could pioneer a new class of redox-based precision anti-inflammatory therapeutics.