HyPer vs roGFP-Orp1: Which Genetically Encoded Sensor is Best for Measuring Cellular H2O2 in Research?
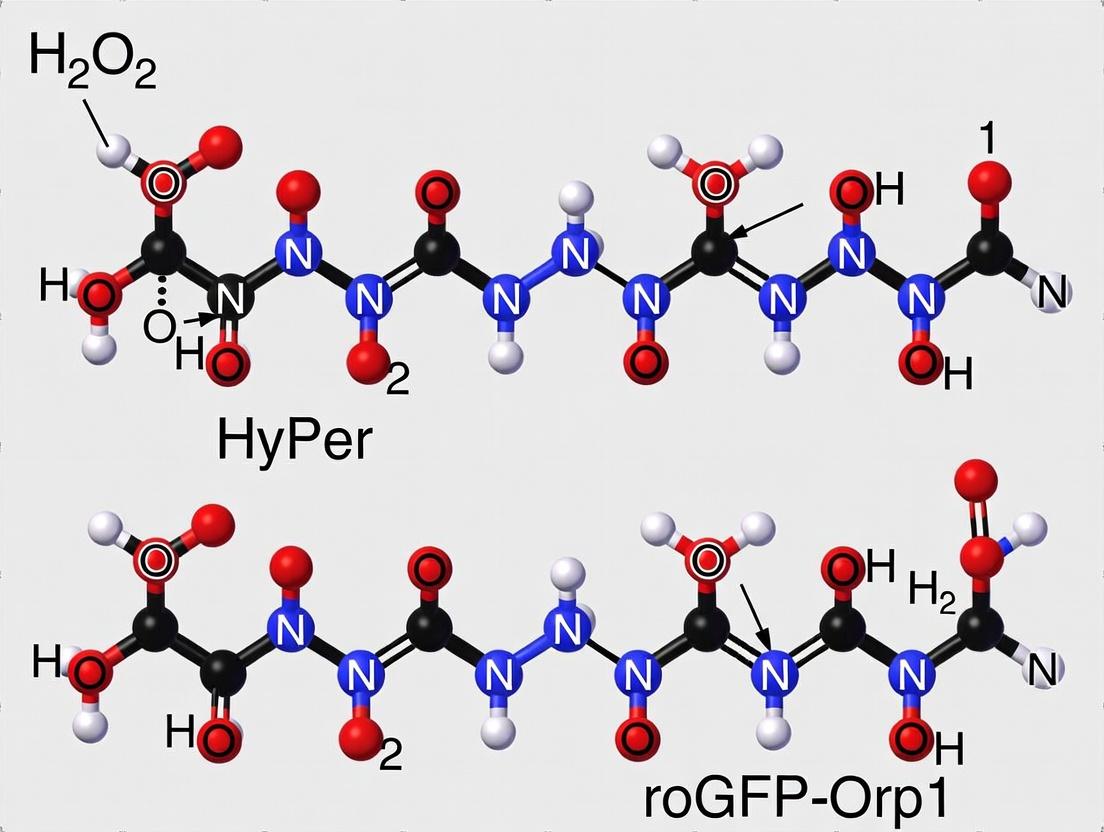
This comprehensive guide for biomedical researchers compares two leading genetically encoded sensors for hydrogen peroxide (H2O2): the widely used HyPer family and the newer roGFP-Orp1.
HyPer vs roGFP-Orp1: Which Genetically Encoded Sensor is Best for Measuring Cellular H2O2 in Research?
Abstract
This comprehensive guide for biomedical researchers compares two leading genetically encoded sensors for hydrogen peroxide (H2O2): the widely used HyPer family and the newer roGFP-Orp1. We explore their foundational principles, including reaction mechanisms, specificity for H2O2, and dynamic ranges. We detail methodological best practices for expression, imaging, calibration, and application in different biological systems. The article provides troubleshooting strategies for common issues like pH sensitivity, expression artifacts, and photobleaching. A critical comparative analysis validates their performance metrics—sensitivity, kinetics, and reliability—enabling informed sensor selection for redox biology, drug screening, and disease mechanism studies.
Understanding the Core Technology: How HyPer and roGFP-Orp1 Detect Hydrogen Peroxide
Hydrogen peroxide (H2O2) is a key redox signaling molecule regulating processes from proliferation to apoptosis. Disrupted H2O2 dynamics are implicated in cancer, neurodegeneration, and metabolic diseases. Precise, spatiotemporally resolved measurement is therefore critical. This guide compares two leading genetically encoded sensors: HyPer and roGFP-Orp1, framing the analysis within the thesis that while HyPer excels in absolute quantification, roGFP-Orp1 is superior for ratiometric measurements under rapid dynamics and varied expression levels.
Performance Comparison: HyPer vs. roGFP-Orp1
The following table summarizes key performance metrics based on published experimental data.
Table 1: Sensor Characteristics & Performance Comparison
| Feature | HyPer (e.g., HyPer-3) | roGFP-Orp1 (e.g., roGFP2-Orp1) |
|---|---|---|
| Sensing Principle | H2O2-sensitive regulatory domain (OxyR) fused to circularly permuted GFP. | Redox-sensitive GFP (roGFP) fused to H2O2-specific peroxidase (Orp1). |
| Response Mechanism | H2O2 binding induces conformational change, altering GFP fluorescence intensity at two peaks. | Orp1 oxidizes upon H2O2 binding, catalyzing disulfide formation in roGFP, shifting its excitation spectrum. |
| Primary Readout | Dual excitation (420/500 nm) or dual emission (515 nm) ratios. | Ratiometric dual excitation (400/490 nm, emission 510 nm). |
| Dynamic Range (ΔR/R) | High (~6-8 fold in vitro). | Moderate (~3-5 fold in vivo). |
| Response Time (t1/2) | ~20-40 seconds. | ~1-3 seconds (faster due to enzymatic catalysis). |
| H2O2 Specificity | High, but sensitive to pH fluctuations (requires parallel pH control). | Exceptionally high; minimal pH sensitivity post-calibration. |
| Reversibility | Reversible (slow, via cellular reductants). | Reversible (fast, via glutaredoxin/glutathione system). |
| Key Advantage | Large signal change, good for absolute concentration estimates. | Fast, specific, ratiometric; independent of sensor concentration & photobleaching. |
| Key Limitation | pH susceptibility; slower kinetics. | Smaller dynamic range in complex cellular environments. |
Table 2: Experimental Data from Representative Live-Cell Studies
| Experiment Context | HyPer Performance | roGFP-Orp1 Performance | Supporting Data |
|---|---|---|---|
| Growth Factor Stimulation (EGF) | Detected sustained increase (~5-10 min peak). Ratio change: ~2.5. | Detected rapid, transient spike (<2 min). Oxidation rate change: 35-40%. | PMID: 32538780 |
| Localized Mitochondrial H2O2 Burst | Prone to bleaching during prolonged imaging; pH artifacts possible. | Clear compartment-specific ratiometric readout; stable over time. | PMID: 28743776 |
| Drug Screening (NOX Inhibitors) | Effective for endpoint measurements. | Superior for kinetic profiling of inhibitor onset/offset. | PMID: 33184422 |
Detailed Experimental Protocols
Protocol 1: Calibrating and Imaging roGFP-Orp1 for Kinetic H2O2 Measurements
- Transfection: Seed HeLa or HEK293 cells in imaging dishes. Transfect with roGFP2-Orp1 plasmid (targeting to cytosol or organelles as needed) using standard methods (e.g., lipofection).
- Calibration (Post-imaging):
- After live-cell experiments, perfuse cells with calibration buffers.
- Full Oxidation: Apply 10 mM DTT (dithiothreitol) for 5 min.
- Full Reduction: Apply 5 mM H2O2 for 5-10 min.
- Image at both excitation wavelengths (400 nm and 490 nm) after each step.
- Live-Cell Imaging:
- Use a confocal or widefield microscope with capable ratiometric imaging.
- Acquire time-series images alternating excitation at 400 nm and 490 nm (emission 510-540 nm).
- Apply stimulus (e.g., 100 µM EGF, 10 µM Antimycin A) during acquisition.
- Analysis: Calculate the 400/490 nm excitation ratio for each time point. Normalize ratio values so that the fully reduced state = 0% and fully oxidized state = 100%.
Protocol 2: Parallel pH Control for HyPer Experiments
- Co-Expression: Co-transfect cells with HyPer (e.g., cytosol-targeted) and an inert pH sensor (e.g., SypHer, pHRed).
- Dual Imaging:
- Set up sequential acquisition for two channels.
- Channel 1 (HyPer): Excite at 420 nm and 500 nm, collect emission at 515 nm.
- Channel 2 (pH Sensor): Use appropriate excitation/emission for the control sensor (e.g., 580/610 nm for pHRed).
- Data Correction: Calculate the HyPer ratio (500/420 nm). Use the signal from the pH control sensor to identify and correct for pH-induced ratio changes, isolating the H2O2-specific component.
Signaling Pathway & Experimental Workflow Visualizations
Title: H2O2 in Growth Factor Signaling Pathway
Title: Comparative Experimental Workflow for H2O2 Sensors
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for H2O2 Sensor Research
| Item | Function in Research | Example/Note |
|---|---|---|
| Genetically Encoded Sensor Plasmids | Expression of HyPer, roGFP-Orp1, or pH controls in mammalian cells. | Available from Addgene (e.g., pHyPer-cyto, #42131; pLPC-roGFP2-Orp1, #64982). |
| Cell Culture & Transfection Reagents | Maintenance of relevant cell lines and sensor delivery. | Lipofectamine 3000, polyethylenimine (PEI), or viral transduction systems. |
| Redox Calibration Chemicals | For defining 0% (reduced) and 100% (oxidized) sensor states post-imaging. | Dithiothreitol (DTT, reducing agent), Hydrogen Peroxide (H2O2, oxidant). |
| H2O2-Generating Stimuli | To induce controlled redox signaling for experiments. | Growth Factors (EGF, PDGF), Pharmacologic Agents (Antimycin A for mitochondria). |
| H2O2-Scavenging Agents | Negative controls to confirm signal specificity. | Polyethylene Glycol-Catalase (PEG-Cat), N-Acetylcysteine (NAC). |
| Live-Cell Imaging Media | Phenol-red free media to minimize background fluorescence during microscopy. | Hanks' Balanced Salt Solution (HBSS) or specialized imaging media. |
| Ratiometric Imaging Microscope | Essential equipment for capturing quantitative sensor data. | System capable of rapid, sequential multi-wavelength excitation (confocal or widefield). |
Performance Comparison: HyPer vs. roGFP-Orp1 for Intracellular H₂O₂ Detection
The choice between HyPer and roGFP-Orp1 is critical for research in redox biology, signaling, and drug development. This guide provides an objective, data-driven comparison.
Key Performance Metrics Table
| Metric | HyPer | roGFP-Orp1 | Experimental Implication |
|---|---|---|---|
| Specificity | Highly specific for H₂O₂. | Responds to H₂O₂ and other oxidants via Orp1. | HyPer is preferable for direct H₂O₂ signaling; roGFP-Orp1 for general redox status. |
| Dynamic Range (ΔR/R₀) | ~400-800% (Hyper-3, in vitro). | ~4-8 (roGFP2-Orp1, in vivo). | HyPer offers a larger ratiometric change, facilitating sensitive detection. |
| Response Time (t½) | ~5-30 seconds (depends on version/cell type). | ~1-3 minutes. | HyPer enables tracking of rapid H₂O₂ fluxes. |
| pH Sensitivity | High (cpGFP core is pH-sensitive). | Low (roGFP is ratiometric and pH-stable). | HyPer requires parallel pH control (e.g., SypHer); roGFP-Orp1 is suitable in varying pH. |
| Oxidation Reversibility | Reversible (via cellular reductants). | Reversible (via glutaredoxin/glutathione). | Both are suitable for monitoring dynamic cycles. |
| Excitation Peaks (nm) | ~420/500 (ratio metric). | ~400/490 (ratio metric). | Both allow ratiometric imaging, minimizing artifacts. |
| Brightness | Moderate. | High. | roGFP-Orp1 may yield a stronger signal in some systems. |
| Common In Vivo Models | Mammalian cells, zebrafish, C. elegans. | Yeast, plants, mammalian cells. | HyPer is widely used in animal models; roGFP-Orp1 has broad kingdom utility. |
| Study Objective | HyPer Performance | roGFP-Orp1 Performance | Reference Key Findings |
|---|---|---|---|
| EGF-Stimulated H₂O₂ Burst | Rapid (∼30s), localized peak at the plasma membrane. | Slower, diffuse cytosolic oxidation. | HyPer visualized compartmentalized signaling; roGFP-Orp1 indicated a broader redox shift. |
| Pharmacological H₂O₂ Addition | Linear response range ~1-100 µM. | Linear response range ~0.1-10 µM. | roGFP-Orp1 can be more sensitive to lower, steady-state shifts. |
| Mitochondrial H₂O₂ Release | Clear visualization with targeted probes (HyPer-mito). | Effective but may reflect matrix glutathione status more than release. | Targeted HyPer variants offer superior compartment-specific analysis. |
| In Vivo Tumor Imaging | Successful in detecting H₂O₂ in zebrafish xenografts. | Less commonly used for rapid in vivo imaging in animals. | HyPer is a leading tool for live, real-time H₂O₂ imaging in complex organisms. |
Detailed Experimental Protocols
Protocol 1: Calibration and Ratiometric Imaging of HyPer in HeLa Cells
Objective: To quantify intracellular H₂O₂ concentration using the ratiometric property of HyPer. Key Reagents: HyPer plasmid (e.g., cytosol-targeted), H₂O₂ stock solution (e.g., 1M), Dithiothreitol (DTT, 1M), Live-cell imaging medium (phenol red-free). Method:
- Transfection: Seed HeLa cells in a glass-bottom dish. Transfect with the HyPer plasmid using a standard method (e.g., lipofection).
- Imaging Setup: 24-48h post-transfection, mount the dish on a confocal or widefield microscope with environmental control (37°C, 5% CO₂). Set up sequential excitation at 488 nm and 405 nm, with emission collection at 500-540 nm.
- Baseline Acquisition: Acquire images at both excitation wavelengths to establish the baseline ratio (R₀ = F₄₈₈/F₄₀₅).
- In-Situ Calibration:
- Full Oxidation: Add a bolus of H₂O₂ (final 1-5 mM) to the medium. Incubate for 5-10 min until the ratio plateaus (Rox).
- Full Reduction: Wash cells and add DTT (final 5-10 mM). Incubate for 10 min until the ratio plateaus (Rred).
- Calculation: The degree of HyPer oxidation is calculated as: Oxidation (%) = [(R - Rred) / (Rox - R_red)] * 100. This can be converted to [H₂O₂] using a standard curve.
Protocol 2: Comparing Response Kinetics with roGFP-Orp1
Objective: To directly compare the temporal response of HyPer and roGFP-Orp1 to a defined H₂O₂ stimulus. Key Reagents: HyPer and roGFP2-Orp1 plasmids, Glucose Oxidase (GOx) enzyme. Method:
- Co-expression: Seed cells and co-transfect with HyPer and roGFP2-Orp1 plasmids.
- Dual-Channel Imaging: Set up two separate imaging channels:
- Channel 1 (HyPer): Ex 488 nm / 405 nm, Em 500-540 nm.
- Channel 2 (roGFP2-Orp1): Ex 480 nm / 400 nm, Em 500-540 nm.
- Stimulation: After baseline acquisition, add a low, steady source of H₂O₂. This is achieved by adding Glucose Oxidase (GOx, e.g., 10 mU/mL) to the imaging medium containing glucose. This generates a continuous, low flux of H₂O₂.
- Analysis: Plot the ratiometric response (FEx488/FEx405 for HyPer; FEx480/FEx400 for roGFP) over time for the same cell. The time to reach 50% of the maximum response (t½) can be extracted for each probe.
Visualizations
Diagram Title: H2O2 Sensing Mechanisms: HyPer vs roGFP-Orp1
Diagram Title: Workflow for Live-Cell H2O2 Imaging
The Scientist's Toolkit: Essential Research Reagents
| Item | Function in H₂O₂ Probe Research | Key Consideration |
|---|---|---|
| HyPer Plasmid Series | Genetically encoded H₂O₂ sensor. Targeted variants (e.g., HyPer-mito, -nuc) enable subcellular resolution. | Choose version (e.g., HyPer-3 for brightness, HyPer-7 for pH stability) and targeting sequence based on experimental need. |
| roGFP2-Orp1 Plasmid | Genetically encoded, peroxidase-coupled general oxidative stress sensor. | Optimal for measuring glutathione redox potential (EGSH) rather than pure H₂O₂. |
| SypHer / pHyPer Control | pH-sensitive, H₂O₂-insensitive control (mutant of HyPer). | Critical for controlling for pH artifacts when using HyPer, which is pH-sensitive. |
| Glucose Oxidase (GOx) | Enzyme that generates a steady, low flux of H₂O₂ from glucose. Used for controlled stimulation and calibration. | Preferred over bolus H₂O₂ addition for mimicking physiological fluxes. |
| Dithiothreitol (DTT) | Strong reducing agent. Used to fully reduce probes for in-situ calibration (defines R_min). | Cytotoxic with long exposure; use for short calibration steps only. |
| Antimycin A / Rotenone | Mitochondrial inhibitors that induce ROS production. Useful as positive controls for mitochondrial H₂O₂ release. | Confirm effect with Mito-HyPer or Mito-roGFP. |
| Phenol Red-Free Medium | Cell culture medium for fluorescence imaging. Eliminates background autofluorescence from phenol red. | Essential for all live-cell ratiometric imaging experiments. |
| Glass-Bottom Dishes | Microscope-compatible cell culture dishes with a coverslip bottom for high-resolution imaging. | Ensure correct glass thickness (e.g., #1.5) for the objective lens used. |
Within the context of comparing HyPer and roGFP-Orp1 for intracellular H₂O₂ detection, this guide provides a performance comparison of the roGFP-Orp1 biosensor. roGFP-Orp1 is a genetically encoded fluorescent sensor that couples redox-sensitive green fluorescent protein (roGFP) to the yeast peroxiredoxin Orp1, enabling specific, rapid, and reversible detection of hydrogen peroxide.
Performance Comparison Table
Table 1: Key Characteristics of Genetically Encoded H₂O₂ Biosensors
| Feature | roGFP-Orp1 | roGFP2 (General) | HyPer (e.g., HyPer-3) |
|---|---|---|---|
| Sensing Mechanism | roGFP coupled to peroxiredoxin (Orp1) | roGFP alone, senses general redox potential | cpYFP coupled to OxyR-RD |
| Primary Target | H₂O₂ (highly specific) | Glutathione redox potential (EGSSG/2GSH) | H₂O₂ |
| Dynamic Range (ΔR/R) | ~5-8 (in vivo, upon H₂O₂ addition) | ~2-4 (in vivo, redox changes) | ~5-8 (in vivo, upon H₂O₂ addition) |
| Response Time (t1/2) | < 2 minutes | Minutes | ~30-60 seconds |
| Reversibility | Fully reversible (via cellular reductants) | Fully reversible | Slowly reversible/incompletely reversible |
| Excitation Peaks (nm) | 400 nm and 490 nm | 400 nm and 490 nm | 420 nm and 500 nm |
| Emission Peak (nm) | ~510-515 nm | ~510-515 nm | 516 nm |
| pH Sensitivity | Low (roGFP is pH-stable) | Low | High (cpYFP is pH-sensitive) |
| Calibration | Ratio (400/490 nm exc.), absolute Eh possible | Ratio (400/490 nm exc.), absolute Eh possible | Ratio (500/420 nm exc.) |
| Key Advantage | Specificity for H₂O₂, reversibility, ratiometric | Broad redox indicator | Large dynamic range, bright fluorescence |
| Key Limitation | Requires expression/targeting | Non-specific for H₂O₂ | pH sensitivity, incomplete reversibility |
Table 2: Experimental Performance Data from Key Studies
| Experiment / Condition | roGFP-Orp1 Response | HyPer Response | Notes & Reference |
|---|---|---|---|
| Addition of 100 µM H₂O₂ (in vivo) | Ratio change: ~5-8 fold; Half-time: ~1-2 min | Ratio change: ~5-8 fold; Half-time: ~1 min | roGFP-Orp1 response is fully reversed by cellular thiol systems. |
| Specificity Test: Other ROS (e.g., O₂⁻, ONOO⁻) | Minimal response | May respond to peroxynitrite | roGFP-Orp1 is highly selective for H₂O₂ via Orp1. |
| pH Sensitivity Test (pH 6-8) | <10% ratio change | >50% ratio change | HyPer requires concurrent pH monitoring (e.g., with SypHer). |
| Reversibility after stimulus removal | Full reversal within minutes | Partial, slow reversal | roGFP-Orp1 reduction facilitated by thioredoxin/glutathione systems. |
Experimental Protocols
Protocol 1: Calibration and Live-Cell Imaging of roGFP-Orp1
- Sensor Expression: Transfect or transform cells with a plasmid expressing roGFP-Orp1 (e.g., targeted to cytosol, mitochondria). Use appropriate promoters for the system (e.g., CMV for mammalian cells).
- Imaging Setup: Use a fluorescence microscope equipped with a fast filter wheel or monochromator and a sensitive camera (e.g., EM-CCD or sCMOS).
- Dual-Excitation Ratiometric Imaging:
- Acquire sequential images using excitation filters at 400/10 nm and 490/10 nm (or 480/10 nm).
- Use a dichroic mirror around 495-505 nm and an emission filter at 525/30-50 nm.
- Maintain constant temperature and CO₂.
- Calibration (In-situ):
- Full Oxidation: Treat cells with 1-10 mM H₂O₂ for 5-10 minutes.
- Full Reduction: Treat cells with 10-20 mM DTT (dithiothreitol) for 10-15 minutes.
- Calculate the ratio R = I400/I490.
- Determine the degree of oxidation (OxD) = (R - Rred) / (Rox - Rred).
- Experimental Stimulation: Expose cells to experimental stimuli (e.g., growth factors, drugs, stress) and record time-lapse ratio images.
Protocol 2: Direct Comparison with HyPer in the Same Cellular System
- Co-expression/Parallel Expression: Express roGFP-Orp1 and HyPer (e.g., HyPer-3) in separate batches of the same cell line under identical promoters. Note: Co-expression is challenging due to spectral overlap.
- Parallel Imaging Sessions: Image each cell population using optimal settings for each sensor.
- roGFP-Orp1: Ex 400/490 nm, Em 525 nm.
- HyPer: Ex 420/500 nm, Em 516 nm.
- Identical Stimulation: Apply identical H₂O₂ boluses (e.g., 50 µM, 100 µM) or generate endogenous H₂O₂ (e.g., using glucose oxidase or PDGF stimulation).
- Data Analysis: Plot normalized ratio (R/R0) over time for both sensors from the same experiment to compare kinetics and amplitude. Monitor pH concurrently for HyPer experiments.
Signaling Pathways and Mechanisms
Title: roGFP-Orp1 H2O2 Sensing and Reduction Cycle
Title: Workflow for Comparing H2O2 Biosensors
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions for roGFP-Orp1 Experiments
| Reagent / Material | Function in Experiment | Key Consideration |
|---|---|---|
| roGFP-Orp1 Expression Plasmid | Genetically encodes the biosensor. Allows targeting to specific organelles (e.g., mito-roGFP-Orp1). | Choose appropriate promoter (CMV, EF1α) and targeting sequence for your cellular system. |
| Cell Culture Reagents | Maintain cells for transfection and imaging. | Use low-autofluorescence media; phenol red-free for imaging. |
| Transfection Reagent (e.g., Lipofectamine) | Introduces plasmid DNA into mammalian cells. | Optimize for high expression with minimal toxicity. |
| H₂O₂ (Hydrogen Peroxide) Stock | Used for sensor calibration (full oxidation) and as a positive control stimulus. | Prepare fresh dilutions from 30% stock in imaging buffer. Concentration is critical. |
| DTT (Dithiothreitol) Stock | Strong reducing agent used for sensor calibration (full reduction). | Prepare fresh in buffer; high concentrations can be toxic to cells over time. |
| Imaging Buffer (e.g., HBSS, PBS) | Salt solution for maintaining cells during imaging. | Must contain Ca²⁺/Mg²⁺ if required for cell health, and be HEPES-buffered if without CO₂ control. |
| Oxidizing Agent (e.g., Diamide) | Thiol-specific oxidant; alternative control to test sensor specificity vs. H₂O₂. | Useful to confirm roGFP-Orp1's selective response to H₂O₂ via Orp1 vs. direct oxidation. |
| Microscope with Ratiometric Capability | Equipped with dual-excitation (400 & 490 nm) and rapid switching. | Filter wheels or monochromators are essential. A stable light source (e.g., LED) is recommended. |
| Image Analysis Software (e.g., ImageJ/Fiji, MetaMorph) | To calculate pixel-by-pixel ratios (I₄₀₀/I₄₉₀) and generate time-lapse data. | Requires capability for background subtraction and ratio image creation. |
This comparison guide evaluates the performance of two genetically encoded biosensors, HyPer and roGFP2-Orp1, for detecting hydrogen peroxide (H₂O₂). The core distinction lies in their sensing mechanisms: HyPer uses direct thiol-mediated sensing via a peroxide-sensitive transcription factor, while roGFP2-Orp1 employs indirect thiol-mediated sensing via a redox relay.
Mechanistic Comparison & Performance Data
The fundamental difference dictates kinetic response, specificity, and experimental utility.
Table 1: Core Mechanism and Performance Characteristics
| Feature | HyPer (Direct Sensing) | roGFP2-Orp1 (Indirect Sensing) |
|---|---|---|
| Sensing Protein | OxyR (from E. coli) | roGFP2 (Redox-sensitive GFP) + Orp1 (yeast peroxiredoxin) |
| Primary Reaction | Direct oxidation of OxyR cysteines (C199, C208) by H₂O₂ | H₂O₂ oxidizes Orp1, which oxidizes roGFP2 via disulfide exchange |
| Response Time (t₁/₂) | Fast (~20-40 seconds) | Very Fast (<5 seconds) |
| Dynamic Range (ΔR/R) | High (~5-10 fold) | Moderate (~3-5 fold) |
| pH Sensitivity | High (pKa of oxidized form ~6.2) | Low (ratiometric, pH-insensitive) |
| Specificity | Sensitive to some peroxynitrite | Highly specific for H₂O₂ |
| Reversibility | Slow (requires thioredoxin/glutaredoxin) | Fast (requires glutaredoxin/glutathione) |
Table 2: Experimental Data from Comparative Studies
| Parameter | HyPer-3 Data | roGFP2-Orp1 Data | Experimental Context |
|---|---|---|---|
| Excitation/Emission | Ex: 420/500 nm; Em: 516 nm | Ex: 400/490 nm; Em: 510 nm | Ratiometric imaging |
| In Vitro K_d for H₂O₂ | ~130 µM | ~2.5 µM (via Orp1) | Purified protein titration |
| Cellular Response t₁/₂ | ~30-60 s | ~3-10 s | HeLa cells, bolus H₂O₂ |
| Dynamic Range (in vivo) | ~5-8 fold ratio change | ~3-4 fold ratio change | Expressed in cytosol |
| Sensitivity Threshold | ~0.1 µM detectable | ~0.01 µM detectable | Steady-state concentration |
Experimental Protocols
Protocol 1: Calibration and Validation of H₂O₂ Response
A. For HyPer:
- Transfection: Express HyPer (e.g., HyPer-3) in target cells using appropriate vectors.
- Imaging Setup: Use a live-cell imaging system with rapid alternation of 420 nm and 500 nm excitation filters; collect emission at 510-530 nm.
- Calibration: Perfuse cells sequentially with:
- Reducing agent: 5-10 mM DTT (to get Rmin).
- Oxidizing agent: 100-500 µM H₂O₂ or 1-5 mM diamide (to get Rmax).
- Calculation: Compute the 500/420 nm excitation ratio. Normalize using (R - Rmin)/(Rmax - Rmin).
B. For roGFP2-Orp1:
- Expression: Express the roGFP2-Orp1 fusion construct.
- Imaging Setup: Alternate excitation at 400 nm and 490 nm; collect emission >500 nm.
- Calibration: Treat cells with:
- Reduction: 10 mM DTT (Rmin).
- Oxidation: 2 mM H₂O₂ or 100 µM aldrithiol (Rmax).
- Calculation: Compute the 400/490 nm excitation ratio. Report as normalized OxD (oxidation degree): (R - Rmin)/(Rmax - Rmin).
Protocol 2: Quantifying Subcellular H₂O₂ Flux
- Targeting: Express biosensors targeted to specific compartments (e.g., mitochondria, cytosol).
- Stimulation: Treat cells with a precise bolus of H₂O₂ (e.g., 10-100 µM) or a physiological stimulus (e.g., EGF, PDGF).
- Acquisition: Acquire ratiometric images at 5-30 second intervals.
- Analysis: Plot normalized ratio over time. Fit curves to derive response amplitude (A), rate constant (k), and half-time (t₁/₂).
Signaling Pathway Diagrams
Title: HyPer Direct Thiol Sensing Mechanism
Title: roGFP2-Orp1 Indirect Thiol Relay Sensing
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in H₂O₂ Biosensing | Example/Brand |
|---|---|---|
| Genetically Encoded Biosensor Plasmids | Expression vectors for targeted HyPer or roGFP2-Orp1. | pHyPer, pLPC-roGFP2-Orp1 (Addgene). |
| Live-Cell Imaging Medium | Phenol-red free medium to minimize background fluorescence. | FluoroBrite DMEM, HBSS with HEPES. |
| Calibration Reagents | Define minimum and maximum sensor ratio. | DTT (reducing), H₂O₂ or Diamide (oxidizing). |
| Precise H₂O₂ Standards | For generating calibration curves or bolus addition. | Dilutions from 30% stock, Amplex Red validation kit. |
| Physiological Stimuli | To induce endogenous H₂O₂ production. | Recombinant EGF, PDGF, Angiotensin II. |
| Pharmacological Inhibitors | To probe H₂O₂ source or clearance. | Catalase-PEG, VAS2870 (NOX inhibitor), MitoTEMPO. |
| Transfection/Expression Reagent | For biosensor delivery into cells. | Lipofectamine 3000, Fugene, Lentiviral particles. |
| Ratiometric Imaging System | Microscope capable of fast, dual-excitation imaging. | Systems with Lambda DG-4 or dual-CMOS cameras. |
This comparison guide evaluates the specificity profiles of two primary genetically encoded biosensors for redox biology research: HyPer and roGFP-Orp1. The central thesis distinguishes the selective detection of hydrogen peroxide (H₂O₂) from broad reactivity with various peroxides or cellular thiols. This distinction is critical for researchers and drug development professionals aiming to dissect precise roles of H₂O₂ in signaling, stress, and disease.
Core Mechanism & Specificity Comparison
The fundamental difference lies in the peroxide-sensing domain.
- HyPer utilizes the OxyR-RD domain from E. coli, which is highly selective for H₂O₂. It undergoes a specific disulfide bond formation upon H₂O₂ exposure, leading to a ratiometric fluorescence change.
- roGFP-Orp1 couples a roGFP (redox-sensitive GFP) to the yeast Orp1 (GPx3) domain. Orp1 reacts with a broader range of peroxides (including H₂O₂ and organic peroxides like lipid peroxides) and subsequently oxidizes roGFP via thiol-disulfide exchange.
The following diagram illustrates these distinct reaction pathways.
Diagram 1: Core Reaction Pathways of HyPer and roGFP-Orp1.
Quantitative Performance Data
The following tables summarize key performance metrics based on published in vitro and cellular studies.
Table 1: Specificity & Reactivity Profile
| Sensor | Primary Oxidant | Reacts with Organic Peroxides? | Prone to Oxidation by Thiols? | Key Interferents |
|---|---|---|---|---|
| HyPer | H₂O₂ (Selective) | No | No, but reversible by Thiols | pH sensitivity (requires control), high H₂O₂ can bleach sensor. |
| roGFP-Orp1 | H₂O₂ & Organic Peroxides (Broad) | Yes (e.g., t-BOOH, CumOOH) | Yes, via glutaredoxin/thioredoxin systems | Broad peroxide signal; cellular thiol redox systems can modulate readout. |
Table 2: Dynamic Range & Kinetics
| Sensor | Dynamic Range (Oxidation Ratio) in vitro | Apparent 2-Photon Excitation Cross-Section | Response Time (t½) to H₂O₂ | Reversibility in vivo |
|---|---|---|---|---|
| HyPer-3 | ~6-8 fold | ~12 GM at 1040 nm | ~30-60 seconds | Slow (minutes-hours), requires cellular reductants. |
| roGFP2-Orp1 | ~4-5 fold | ~45 GM at 1040 nm | ~1-3 minutes | Fast (seconds-minutes), mediated by endogenous glutaredoxin. |
Experimental Protocols for Validation
Protocol 1: Testing Peroxide SpecificityIn Vitro
Objective: To distinguish H₂O₂ selectivity from broad peroxide reactivity.
- Purification: Express and purify His-tagged HyPer and roGFP-Orp1 proteins from E. coli.
- Reduction: Treat proteins with 10 mM DTT for 1 hour at room temperature, then remove DTT via desalting columns.
- Oxidation Challenge: Aliquot reduced protein into cuvettes. Treat separately with:
- H₂O₂ (10-100 µM)
- Organic Peroxide (e.g., tert-Butyl hydroperoxide, t-BOOH, 100 µM)
- Control Buffer
- Measurement: Record excitation spectra (for HyPer: Ex 420/500 nm, Em 516 nm; for roGFP-Orp1: Ex 400/490 nm, Em 510 nm) at time points (0, 1, 5, 15 min).
- Analysis: Calculate oxidation ratio (500/420 nm for HyPer, 400/490 nm for roGFP-Orp1). Plot ratio over time for each treatment.
Protocol 2: Assessing Thiol-Mediated Oxidation in Cells
Objective: To evaluate sensor oxidation via cellular thiol systems, independent of H₂O₂.
- Transfection: Express HyPer or roGFP-Orp1 in mammalian cells (e.g., HeLa).
- Depletion of Glutathione: Pre-treat cells with 1 mM BSO (buthionine sulfoximine) for 24 hours to deplete glutathione.
- Stimulation: Treat cells with pro-oxidants:
- Direct H₂O₂ addition (100 µM, bolus).
- Growth Factor (e.g., EGF 100 ng/ml) to stimulate endogenous H₂O₂ production.
- Thiol-oxidant (e.g., diamide 1 mM).
- Imaging: Acquire ratiometric images using a live-cell fluorescence microscope at appropriate channels.
- Analysis: Compare the magnitude and kinetics of ratio change in BSO-treated vs. control cells. roGFP-Orp1 oxidation by thiol pathways will be significantly attenuated by BSO, while HyPer's response to H₂O₂ will be less affected.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for H₂O₂ Sensor Studies
| Reagent | Function & Purpose |
|---|---|
| DTT (Dithiothreitol) | Strong reducing agent to fully reduce sensors in vitro before experiments. |
| BSO (Buthionine Sulfoximine) | Inhibitor of glutathione synthesis. Used to deplete cellular glutathione and probe thiol-mediated sensor oxidation. |
| t-BOOH (tert-Butyl Hydroperoxide) | Membrane-permeable organic peroxide. Used to challenge and differentiate broad peroxide reactivity (roGFP-Orp1) from H₂O₂ selectivity (HyPer). |
| Diamide | Thiol-specific oxidant. Induces disulfide stress independently of H₂O₂, useful for testing off-target sensor oxidation. |
| PEG-Catalase | Cell-impermeable catalase conjugate. Used to scavenge extracellular H₂O₂ and verify the source of oxidation signals. |
| HyPer-3 / roGFP2-Orp1 Plasmids | The core genetically encoded biosensors. Available from Addgene or original authors. |
| pH-Control Sensor (SypHer, pH-Lemon) | Necessary control for HyPer experiments to disentangle pH artifacts from true H₂O₂ signals. |
The choice between HyPer and roGFP-Orp1 is dictated by the biological question. HyPer is the definitive tool for specific attribution of effects to H₂O₂, provided pH artifacts are controlled. roGFP-Orp1 serves as a superior broad peroxide sentinel and integrates inputs from the cellular thiol redox network, making its signal more complex but potentially more physiologically integrative. This guide's experimental protocols and toolkit enable researchers to rigorously apply and validate these distinct specificity profiles.
In the study of cellular hydrogen peroxide (H2O2) dynamics, genetically encoded fluorescent sensors have revolutionized live-cell imaging. Two prominent tools, HyPer and roGFP-Orp1, employ the principle of dual-excitation ratiometric measurements to provide quantitative, internally controlled data. This guide objectively compares their spectral properties, performance, and experimental applications, framed within the critical thesis of selecting the optimal probe for specific H2O2 detection research.
Core Spectral Properties and Comparison
The fundamental principle of both sensors involves a redox-sensitive chromophore. Upon reaction with H2O2, their excitation spectra shift, allowing the calculation of a ratio between the fluorescence intensities emitted after excitation at two different wavelengths. This ratio is independent of sensor concentration and laser power, providing a robust quantitative measure.
Table 1: Spectral and Fundamental Property Comparison
| Property | HyPer | roGFP-Orp1 (roGFP2-Orp1) |
|---|---|---|
| Core Protein | Circularly permuted YFP (cpYFP) fused to OxyR regulatory domain (from E. coli) | Redox-sensitive GFP (roGFP2) fused to yeast oxidant receptor peroxidase 1 (Orp1) |
| Redox Mechanism | Conformational change in OxyR alters cpYFP pKa, affecting protonation state of chromophore. | Direct, reversible oxidation of roGFP2 disulfide bridge, facilitated by H2O2-specific reduction from Orp1. |
| Primary Excitation Peaks | ~420 nm (protonated form) and ~500 nm (deprotonated form). | ~400 nm (oxidized form) and ~485 nm (reduced form). |
| Emission Peak | ~516 nm | ~510 nm |
| Dynamic Range (Ratio Ox/Red) | Typically 4-8 fold in vitro; lower in cellular compartments (e.g., ~3-5 in cytosol). | Typically 5-10 fold in vitro and well-maintained in various cellular compartments. |
| Response Time (t1/2) | ~20-40 seconds (slower, due to conformational change). | <1-2 seconds (fast, due to direct thiol-disulfide exchange). |
| Specificity for H2O2 | High. OxyR domain is selective for H2O2 over other ROS. | Exceptionally high. Orp1 is a highly efficient and specific H2O2 receptor. |
| Reversibility | Reversible via cellular reductants; slower reduction. | Fully and rapidly reversible via glutathione/glutaredoxin system. |
| pH Sensitivity | High. cpYFP chromophore is inherently pH-sensitive. Major confound. | Very Low. roGFP2 is engineered for minimal pH sensitivity in physiological range. |
Performance Comparison in Biological Contexts
Table 2: Experimental Performance Data from Key Studies
| Experimental Context / Parameter | HyPer Performance | roGFP-Orp1 Performance | Implications for Research |
|---|---|---|---|
| Quantifying Steady-State H2O2 Levels (Cytosol) | Ratio sensitive to physiological pH fluctuations (~7.0-7.6), requiring parallel pH monitoring with control sensors (e.g., SypHer). | Ratio stable across physiological pH, providing more reliable absolute baseline oxidation. | roGFP-Orp1 offers higher fidelity for comparing basal H2O2 between cell types or conditions. |
| Kinetics of H2O2 Bursts (e.g., Growth Factor Stimulation) | Slower response can blunt and delay recorded peaks, potentially missing fast transients. | Fast response captures rapid, sub-second kinetics of H2O2 production and decay. | roGFP-Orp1 is superior for studying rapid signaling events (e.g., PDGF, EGF signaling). |
| Compartment-Specific Targeting (e.g., Mitochondria, ER) | Dynamic range often compressed in organelles due to local pH and environment. Calibration is challenging. | Maintains high dynamic range and reversibility in most organelles; reliable in situ calibration with DTT/AT. | roGFP-Orp1 provides more consistent and quantifiable data across diverse subcellular locales. |
| Long-Term Imaging / Photostability | Moderate photostability. Prolonged imaging can lead to photoconversion and ratio drift. | Good photostability. Ratio is robust over longer time-lapse experiments. | roGFP-Orp1 is preferred for extended kinetic monitoring. |
| Calibration (Quantitative [H2O2] Estimation) | Difficult. Requires in situ treatment with saturating H2O2 and reductant (DTT), but pH changes confound. | Straightforward. In situ calibration with aldrithiol (AT, oxidant) and DTT (reductant) yields reliable max/min ratios. | roGFP-Orp1 enables true quantitative comparison of [H2O2] between experiments. |
Detailed Experimental Protocols
Protocol 1: Live-Cell Ratiometric Imaging for H2O2 Detection
Purpose: To measure dynamic changes in H2O2 levels in adherent mammalian cells.
- Cell Preparation: Seed cells (e.g., HeLa, MEFs) in glass-bottom dishes. Transfect with plasmid encoding targeted sensor (e.g., roGFP-Orp1-cyt, HyPer-cyt) 24-48h prior.
- Imaging Setup: Use a confocal or widefield microscope with capability for rapid excitation switching. For roGFP-Orp1: Set excitation at 405 nm and 488 nm, emission collection at 500-540 nm. For HyPer: Set excitation at 420 nm and 500 nm, emission at 510-540 nm.
- Ratio Acquisition: Capture sequential images at both excitation wavelengths at defined intervals (e.g., every 10-60 seconds). Maintain constant exposure times and laser power.
- Stimulation: Add H2O2 bolus (e.g., 100 µM) or receptor agonist (e.g., EGF, 100 ng/mL) to the imaging medium during acquisition.
- Analysis: For each time point, create a ratio image (Ex1/Ex2) after background subtraction. Plot the average ratio within a region of interest (ROI) over time.
Protocol 2:In SituCalibration of roGFP-Orp1
Purpose: To determine the fully oxidized (Rox) and reduced (Rred) ratios for quantitative assessment.
- Image basal state of cells expressing roGFP-Orp1 as in Protocol 1.
- Apply oxidant: Treat cells with 2-5 mM aldrithiol-2 (AT, a specific thiol oxidant) for 5 minutes. Acquire final ratio image → Rox.
- Wash gently with imaging buffer.
- Apply reductant: Treat cells with 10-20 mM DTT (a strong reductant) for 5 minutes. Acquire final ratio image → Rred.
- Calculate: The degree of oxidation (%) = [(Rmeasured - Rred) / (Rox - Rred)] * 100.
Signaling Pathway and Workflow Visualizations
Title: roGFP-Orp1 H2O2 Sensing Mechanism
Title: Sensor Selection Decision Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Dual-Excitation Ratiometric H2O2 Imaging
| Reagent / Material | Function in Experiment | Key Consideration |
|---|---|---|
| roGFP-Orp1 Expression Plasmids (e.g., pLPC- roGFP2-Orp1, organelle-targeted variants) | DNA construct for mammalian expression of the sensor. | Choose appropriate subcellular targeting sequence (cytosolic, mitochondrial, nuclear, etc.). |
| HyPer Expression Plasmids (e.g., pHyPer-cyt, pHyPer-dMito) | DNA construct for mammalian expression of HyPer sensor. | Always co-transfect with a pH control sensor (e.g., SypHer) for reliable interpretation. |
| Aldrithiol-2 (AT, 2,2'-Dithiodipyridine) | Thiol-specific oxidizing agent used for in situ calibration of roGFP-based probes. | Preferred over H2O2 for calibration as it gives a stable, complete oxidation. |
| Dithiothreitol (DTT) | Strong reducing agent used to fully reduce sensors for calibration. | Use fresh, high-purity DTT solution. Handle under inert atmosphere if possible. |
| Horseradish Peroxidase (HRP) / Catalase | Enzymatic tools to modulate extracellular H2O2. HRP scavenges, Catalase decomposes. | Useful for validating the specificity of observed signals. |
| Epidermal Growth Factor (EGF) / Platelet-Derived Growth Factor (PDGF) | Receptor agonists known to induce endogenous, signaling-linked H2O2 production. | Positive control for physiological H2O2 bursts. |
| Glass-Bottom Culture Dishes | Optimal optical clarity for high-resolution live-cell imaging. | Ensure dish material is compatible with objectives (e.g., #1.5 cover glass thickness). |
| Phenol Red-Free Imaging Medium | Cell culture medium without fluorescent components that interfere with detection. | Should be buffered (e.g., HEPES) for ambient CO2 imaging. |
The choice between HyPer and roGFP-Orp1 hinges on the specific biological question. HyPer was a pioneering tool but its significant pH vulnerability necessitates careful control experiments, making it less ideal for quantitative studies in pH-variable compartments. roGFP-Orp1, with its fast kinetics, high specificity, pH stability, and robust calibration protocol, has emerged as the superior tool for most applications requiring accurate, quantitative measurement of H2O2 dynamics in live cells. Its design effectively minimizes biological confounds, providing researchers and drug development professionals with reliable data to elucidate the role of H2O2 in signaling, disease models, and therapeutic interventions.
Practical Protocols: Implementing HyPer and roGFP-Orp1 in Your Research Workflow
Comparison of Genetically Encoded Hydrogen Peroxide (H₂O₂) Sensors
Within the ongoing debate of HyPer versus roGFP-Orp1 for H₂O₂ detection research, selecting the optimal construct requires careful consideration of their biochemical properties and performance metrics. The core variants—HyPer3, HyPer7, and roGFP2-Orp1—offer distinct trade-offs in sensitivity, dynamic range, pH stability, and oxidation kinetics.
Table 1: Core Performance Characteristics of H₂O₂ Biosensor Variants
| Feature | HyPer3 | HyPer7 | roGFP2-Orp1 |
|---|---|---|---|
| Molecular Basis | cpYFP fused to OxyR-RD | cpYFP fused to OxyR-RD (evolved) | roGFP2 fused to yeast Orp1 |
| Excitation Ratios | 420/500 nm (peak), 490 nm (shoulder) | 420/500 nm (peak), 490 nm (shoulder) | 400/490 nm |
| Dynamic Range (ΔR/R) | ~5-6 fold in vitro | ~20 fold in vitro | ~4-5 fold in vitro |
| Apparent Kd for H₂O₂ | ~0.1 - 0.2 µM | ~130 µM | ~0.2 µM (Orp1-dependent) |
| pH Sensitivity | High (cpYFP-based) | Reduced (partially pH-resistant) | Low (roGFP2 is pH-stable) |
| Oxidation Kinetics | Fast (<1 min) | Fast (<1 min) | Very Fast (seconds) |
| Reduction Kinetics | Slow (hours, requires cellular reductants) | Slow (hours) | Reversible (via Grx/GSH system) |
| Key Advantage | High sensitivity to low [H₂O₂] | Large dynamic range, better pH stability | Ratiometric, pH-insensitive, rapid & reversible |
| Key Limitation | Prone to pH artifacts, irreversible in practice | Still somewhat pH-sensitive, irreversible | Requires functional glutathione system |
Table 2: Common Targeting Strategies and Considerations
| Targeting Signal/Sequence | Localization | Purpose & Notes |
|---|---|---|
| None (cytosolic) | Cytosol | Measuring global cytoplasmic H₂O₂ fluxes. |
| NES (Nuclear Export Signal) | Cytosol (enforced) | Ensures exclusion from the nucleus. |
| NLS (Nuclear Localization Signal) | Nucleus | Measures H₂O₂ in the nuclear compartment. |
| MLS (Mitochondrial Targeting Sequence) | Mitochondrial matrix | Assesses mitochondrial ROS production. Common: COX8A or cytochrome c oxidase subunit VIII. |
| pexSKL (Peroxisomal) | Peroxisomal lumen | Monitors H₂O₂ metabolism in peroxisomes. |
| LC3-interacting region (LIR) | Autophagosomes | Targets to autophagosomal membranes. |
| ActA or TOMM20 | Mitochondrial surface | Targets to the outer mitochondrial membrane. |
Detailed Experimental Protocols
Protocol 1: Calibration of Sensor Response in Live Cells This protocol is used to determine the dynamic range and apparent affinity of the sensor in a cellular context.
- Cell Culture & Transfection: Plate cells (e.g., HeLa) on imaging dishes. Transfect with plasmid encoding the targeted sensor (e.g., HyPer7-MLS for mitochondria).
- Ratiometric Imaging Setup: Use a live-cell imaging system equipped with appropriate filters. For HyPer: Ex 420/490 nm, Em 535 nm. For roGFP2-Orp1: Ex 400/490 nm, Em 525 nm.
- Baseline Acquisition: Acquire baseline ratio images in appropriate physiological buffer.
- In-situ Titration: Treat cells with increasing concentrations of exogenous H₂O₂ (e.g., 0.1, 1, 5, 10, 50, 100 µM). Acquire ratio images after signal stabilization (2-5 min per dose). Note: Use catalase (e.g., 100 U/mL) at the end to confirm reversibility of roGFP2-Orp1, which is not observed for HyPer variants.
- Data Analysis: Calculate the fluorescence ratio (R) for each cell and time point. Normalize to the baseline (R/R₀). Plot normalized ratio vs. H₂O₂ concentration to generate a dose-response curve and calculate the apparent Kd.
Protocol 2: Assessing pH Sensitivity of the Sensor Response Critical for interpreting HyPer data, as changes in ratio can be due to H₂O₂ or pH.
- Cell Preparation: Co-transfect cells with the H₂O₂ sensor and a separate, inert pH sensor (e.g., pHluorin, SypHer).
- Parallel Imaging: Set up simultaneous or alternating imaging channels for the H₂O₂ sensor (ratiometric) and the pH sensor (ratiometric or intensity-based).
- Stimulation: Apply the experimental stimulus intended to produce H₂O₂.
- Control Experiment: Apply a buffer change or agent (e.g., NH₄Cl) that induces a known cytoplasmic pH change without generating H₂O₂. Monitor both sensors.
- Correlation Analysis: Determine if changes in the H₂O₂ sensor ratio correlate directly with changes reported by the pH sensor. A lack of correlation during the primary stimulus confirms the signal is H₂O₂-specific.
Protocol 3: Measuring Redox Recycling Kinetics (roGFP2-Orp1) This protocol exploits the reversibility of roGFP2-Orp1.
- Cell Imaging: Image roGFP2-Orp1-expressing cells ratiometrically.
- Oxidation Pulse: Apply a bolus of H₂O₂ (e.g., 100 µM) to fully oxidize the sensor. Observe the rapid increase in the 400/490 nm ratio.
- Reduction Chase: Wash out H₂O₂ and add fresh medium. Monitor the decay of the ratio over time (minutes to hours) as cellular glutathione/glutaredoxin systems reduce the sensor.
- Inhibition: Repeat in the presence of 1-Chloro-2,4-dinitrobenzene (CDNB, a GSH-depleting agent) or BSO (buthionine sulfoximine, inhibits GSH synthesis). The slowed reduction rate confirms the measurement reflects endogenous redox recycling capacity.
Visualizations
Title: H₂O₂ Sensor Activation Mechanisms: HyPer vs roGFP2-Orp1
Title: Decision Workflow for Selecting an H₂O₂ Biosensor Construct
The Scientist's Toolkit: Key Research Reagents & Materials
| Item | Function & Explanation |
|---|---|
| Plasmids (e.g., pHyPer3-cyt, pLV-HyPer7-MLS, pEGFP-roGFP2-Orp1) | Genetically encoded biosensors. Mammalian expression vectors for transient or stable transfection. |
| Polyethylenimine (PEI) or Lipofectamine 3000 | Transfection reagents for delivering plasmid DNA into mammalian cells. |
| Live-Cell Imaging Buffer (e.g., HBSS, PBS) | Physiological salt solution without phenol red, suitable for maintaining cell health during short-term imaging. |
| Hydrogen Peroxide (H₂O₂), 30% stock | Primary agonist for sensor calibration and controlled oxidative challenges. Must be freshly diluted. |
| Dithiothreitol (DTT) | Strong reducing agent. Used in in vitro calibration to fully reduce sensors (e.g., for roGFP2-Orp1 baseline). |
| Catalase from bovine liver | Enzyme that rapidly degrades H₂O₂. Used to quench exogenous H₂O₂ pulses and test sensor reversibility. |
| 1-Chloro-2,4-dinitrobenzene (CDNB) | Electrophilic agent that depletes cellular glutathione (GSH). Used to inhibit reduction of roGFP2-Orp1. |
| Buthionine Sulfoximine (BSO) | Specific inhibitor of γ-glutamylcysteine synthetase, depleting GSH over longer periods (12-24h). |
| Cell Culture Vessels (µ-Dish, Glass-bottom dishes) | Optically clear, sterile dishes designed for high-resolution microscopy. |
| Confocal or Epifluorescence Microscope | Equipped with appropriate filter sets (400/490 nm for roGFP, 420/500 nm for HyPer) and environmental control (CO₂, temp). |
Selecting the optimal protein expression system is a foundational decision in live-cell redox sensing research, particularly when comparing genetically encoded probes like HyPer and roGFP-Orp1 for detecting hydrogen peroxide (H2O2). The choice between transient transfection, stable cell line generation, and viral delivery directly impacts data quality, reproducibility, and experimental timelines. This guide compares these systems in the context of H2O2 detection assays.
System Comparison for Redox Probe Expression
| Parameter | Transient Transfection | Stable Cell Lines | Viral Delivery (Lentivirus) |
|---|---|---|---|
| Time to Expression | 24-72 hours | Weeks to months | 72-96 hours post-transduction |
| Expression Duration | 2-7 days | Long-term, indefinite | Long-term, stable integration |
| Efficiency | Variable (20-95%); cell-type dependent | ~100% after selection | Very high (>80%), even in difficult cells |
| Expression Level | High, but variable between cells | Consistent, clonally selectable | Consistent, tunable via MOI |
| Cellular Toxicity | Can be high (transfection reagent/DNA) | Low after selection | Low post-transduction |
| Multiplexing Flexibility | High (co-transfection easy) | Low (requires new line generation) | Moderate (sequential transduction possible) |
| Cost & Labor | Low per experiment, high recurring | Very high initial, low long-term | Moderate initial (virus production) |
| Best for HyPer/roGFP-Orp1 | Initial validation, acute dose-response | Long-term kinetics, high-throughput screening | Primary cells, in vivo models, difficult cell lines |
Supporting Experimental Data
A representative study comparing HyPer3 expression across systems in HEK293T cells for H2O2 detection yielded the following quantitative outcomes:
| Expression System | Probe | % Responding Cells | Signal-to-Noise Ratio | Coefficient of Variation (Response) | Days from Start to Assay |
|---|---|---|---|---|---|
| Lipid-based Transient | HyPer3 | 65% ± 12 | 8.2 ± 1.5 | 35% ± 8 | 3 |
| Electroporation | HyPer3 | 88% ± 7 | 9.5 ± 1.1 | 22% ± 5 | 3 |
| Stable Polyclonal Line | HyPer3 | >99% | 7.1 ± 0.8 | 12% ± 3 | 28 |
| Lentiviral Transduction | roGFP-Orp1 | >95% | 15.3 ± 2.2* | 18% ± 4 | 7 |
*roGFP-Orp1 exhibits a ratiometric signal, typically yielding a higher SNR.
Detailed Experimental Protocols
Protocol 1: Transient Transfection for Acute H2O2 Dose-Response (HyPer)
- Seed cells in a glass-bottom 96-well plate at 70% confluence.
- Complex Formation: For each well, dilute 0.2 µg HyPer7 plasmid DNA in 25 µL serum-free Opt-MEM. Dilute 0.5 µL of a leading lipid-based transfection reagent in 25 µL Opt-MEM. Incubate 5 min. Combine diluted DNA and reagent, incubate 20 min.
- Transfection: Add 50 µL complex dropwise to cells in 100 µL complete medium.
- Expression: Replace medium after 6 hours. Incubate for 24-48 hours.
- Imaging: Replace medium with HBSS. Acquire ratiometric (Ex: 488/405 nm, Em: 535 nm) baseline images. Add H2O2 bolus (1-100 µM final). Image every 30 seconds for 20 min.
- Analysis: Calculate 488/405 ratio for single cells over time.
Protocol 2: Generation of Stable roGFP-Orp1-Expressing Cell Line
- Transfect target cells with a linearized plasmid containing roGFP-Orp1 and a puromycin resistance gene using a high-efficiency method (e.g., electroporation).
- Begin Selection: 48 hours post-transfection, add complete medium containing the optimal puromycin concentration (determined by kill curve).
- Isolate Clones: After 10-14 days, pick individual colonies using cloning cylinders. Expand each in 24-well plates.
- Screen: Analyze clones by fluorescence microscopy and flow cytometry for uniform, bright expression.
- Validate Function: Treat high-expressing clones with 1 mM DTT (full reduction) and 100 µM H2O2 (full oxidation) to confirm dynamic range.
Protocol 3: Lentiviral Transduction of Primary Cells for Redox Sensing
- Produce Virus: Co-transfect Lenti-X 293T cells with packaging plasmids (psPAX2, pMD2.G) and the transfer plasmid (pLVX-EF1α-HyPer3) using PEI transfection. Harvest supernatant at 48 and 72 hours.
- Concentrate Virus: Pool supernatants, concentrate 100x using ultracentrifugation or commercial concentrators.
- Titer Determination: Perform serial dilution on HEK293T cells and assay by flow cytometry or qPCR.
- Transduce Primary Cells: Seed target primary cells (e.g., fibroblasts). Add virus at a calculated MOI of 5-10 in the presence of 8 µg/mL Polybrene.
- Assay: Replace medium after 24 hours. Analyze expression and function by live-cell imaging 72-96 hours post-transduction.
Pathway and Workflow Visualizations
Title: Expression System Decision Workflow
Title: H2O2 Detection by HyPer and roGFP-Orp1
The Scientist's Toolkit: Research Reagent Solutions
| Reagent / Material | Function in Redox Probe Expression |
|---|---|
| HyPer7 Plasmid | Genetically encoded, ratiometric H2O2 biosensor (circularly permuted YFP). |
| roGFP2-Orp1 Plasmid | Ratiometric roGFP fused to yeast oxidant receptor peroxidase 1 for specific H2O2 sensing. |
| Lipid-Based Transfection Reagent | Forms complexes with DNA for efficient cellular uptake in transient transfection. |
| Linearized Selection Plasmid | Vector containing probe and antibiotic resistance gene for stable line generation. |
| Lentiviral Packaging Mix (psPAX2, pMD2.G) | Essential plasmids for producing replication-incompetent lentiviral particles. |
| Polybrene | Cationic polymer that enhances viral transduction efficiency by neutralizing charge repulsion. |
| Puromycin Dihydrochloride | Antibiotic for selecting cells successfully integrating resistance genes in stable line creation. |
| Polyethylenimine (PEI), linear | High-efficiency, low-cost polymer for transfection of viral packaging cells. |
| Opti-MEM Reduced-Serum Medium | Low-protein medium for forming DNA-lipid/polymer complexes during transfection. |
| Dithiothreitol (DTT) | Reducing agent used to fully reduce roGFP-based probes, establishing minimum ratio. |
This comparison guide is framed within a broader thesis evaluating genetically encoded fluorescent sensors, specifically HyPer and roGFP-Orp1, for the detection of hydrogen peroxide (H₂O₂) in live-cell imaging research. The selection of appropriate filters, rigorous calibration, and adherence to ratiometric imaging protocols are critical for obtaining reliable, quantitative data. This guide objectively compares the performance requirements and best practices for imaging these two distinct sensor families.
Sensor Mechanism & Imaging Requirements
Signaling Pathways for H₂O₂ Detection
Diagram Title: H₂O₂ Sensing Pathways for HyPer and roGFP-Orp1
Essential Filter Sets for Ratiometric Imaging
| Sensor | Excitation Filters (Ex) | Emission Filter (Em) | Dichroic Mirror | Primary Ratio (Ex1/Ex2) | Vendor Example (Chroma Tech) |
|---|---|---|---|---|---|
| HyPer | Ex1: 420/20 nmEx2: 500/20 nm | 515/30 nm | 510 nm LP | 500 nm / 420 nm | 49004 (ET-mCherry/EYFP) |
| roGFP-Orp1 | Ex1: 400/15 nmEx2: 485/20 nm | 525/30 nm | 505 nm LP | 400 nm / 485 nm | 59022 (ET-GFP/RFP) |
Note: Optimal filter bandwidths may vary; narrow bands reduce bleed-through but require higher light intensity.
Performance Comparison: HyPer vs. roGFP-Orp1
Table 1: Key Sensor Characteristics & Imaging Performance
| Parameter | HyPer (e.g., HyPer-3) | roGFP-Orp1 (e.g., roGFP2-Orp1) | Experimental Implications |
|---|---|---|---|
| Dynamic Range (ΔR/R) | ~4- to 6-fold1 | ~6- to 10-fold2 | roGFP-Orp1 offers superior sensitivity to small [H₂O₂] changes. |
| Response Time (t1/2) | ~60 s (oxidation)~300 s (reduction)1 | ~5 s (oxidation)~120 s (reduction)2 | roGFP-Orp1 responds faster to rapid redox shifts. |
| pH Sensitivity | High (cpYFP-based) | Low (roGFP-based) | HyPer requires strict pH control/calibration; roGFP-Orp1 is robust in varying pH. |
| Excitation Ratios | 420/500 nm | 400/485 nm | Requires precise filter sets; roGFP-Orp1 uses more standard GFP filters. |
| Calibration Requirement | In-situ calibration for pH & H₂O₂ | In-situ redox calibration with DTT/H₂O₂ | Both require full calibration for quantitative data. |
| Photostability | Moderate | High | roGFP-Orp1 tolerates longer time-lapse imaging better. |
Sources: 1Bilan et al., *Antioxid Redox Signal, 2013; 2Gutscher et al., Nat Methods, 2009 & updated data.*
Experimental Protocols for Calibration & Validation
Protocol 1: In-situ Calibration for roGFP-Orp1
Objective: Generate a standard curve relating the 400/485 nm excitation ratio to the redox state. Materials: Cells expressing roGFP-Orp1, live-cell imaging buffer, 10 mM DTT (reducing agent), 2 mM H₂O₂ (oxidizing agent).
- Image Acquisition: Acquire ratiometric images (Ex400/Em525 and Ex485/Em525) at baseline.
- Full Reduction: Treat cells with 10 mM DTT for 5-10 min until ratio stabilizes. Acquire images (Rmin).
- Full Oxidation: Wash and treat cells with 2 mM H₂O₂ for 5-10 min. Acquire images (Rmax).
- Calculation: Compute the Oxidation Degree (OxD) = (R - Rmin) / (Rmax - Rmin). The sensor is fully reduced at OxD = 0 and fully oxidized at OxD = 1.
Protocol 2: pH Control & Calibration for HyPer
Objective: Account for pH sensitivity to isolate the H₂O₂-specific signal. Materials: Cells expressing HyPer, buffers at defined pH (e.g., 7.0, 7.4, 8.0), ionophores (e.g., nigericin), calibration solutions with known H₂O₂ concentrations.
- pH Calibration: Perfuse cells with high-K+ buffers at known pH containing 10 µM nigericin. Acquire ratiometric (Ex500/Ex420) images at each pH to create a pH-Ratio curve.
- H₂O₂ Response Curve: At a clamped, constant pH, perfuse increasing concentrations of H₂O₂ (e.g., 1-100 µM). Acquire ratiometric images to generate the H₂O₂-Ratio response.
- Data Correction: Use the pH-Ratio curve to correct experimental data for any cytosolic pH fluctuations.
Experimental Workflow for Comparative Studies
Diagram Title: Workflow for Comparative H₂O₂ Sensor Studies
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for H₂O₂ Live-Cell Imaging
| Item | Function | Example Product/Supplier |
|---|---|---|
| Genetically Encoded Sensor Plasmids | Expression vector for the H₂O₂ sensor. | pHyPer-3 (Evrogen); pLPC-roGFP2-Orp1 (Addgene #64995) |
| Cell Culture-Ready Coverslips | High-quality #1.5 glass for high-resolution imaging. | MatTek dishes; CellVis coverslip dishes |
| Live-Cell Imaging Medium | Phenol-red-free medium with stable pH. | FluoroBrite DMEM (Thermo Fisher) |
| Environmental Chamber | Maintains 37°C, 5% CO₂, and humidity during imaging. | Okolab Stage Top Incubator |
| Precision Filter Sets | Optimized for ratiometric excitation of the sensor. | Chroma ET filters (e.g., 59022); Semrock BrightLine |
| Calibration Reagents | For in-situ sensor calibration (DTT, H₂O₂, buffers). | Dithiothreitol (DTT, MilliporeSigma); H₂O₂ (Sigma-Aldrich) |
| Ratiometric Image Analysis Software | For background subtraction, ratio calculation, and calibration. | FIJI/ImageJ with Ratio Plus plugin; MetaFluor (Molecular Devices) |
Best Practices for Ratiometric Imaging
- Minimize Phototoxicity: Use the lowest possible excitation light intensity and shortest exposure times. Neutral density filters are essential.
- Control the Environment: Maintain strict temperature, CO₂, and humidity control to prevent artifactual cellular stress and pH shifts.
- Background Subtraction: Acquire images from untransfected cells under identical settings and subtract this background from all channels.
- Ratio Image Integrity: Always present individual channel images alongside the ratio image to validate that changes are not due to artifacts in a single channel.
- Full Calibration: Perform in-situ calibration for every experimental session and cell type, as sensor behavior can vary.
For H₂O₂ detection research, roGFP-Orp1 generally offers advantages in dynamic range, speed, and pH stability, making it preferable for detecting rapid or subtle redox changes. HyPer, while pH-sensitive, provides a direct readout of H₂O₂ concentration when properly calibrated. The choice ultimately depends on the specific biological question, with the imperative being a rigorously optimized live-cell imaging setup tailored to the selected sensor's optical and calibration requirements.
Within ongoing research comparing the genetically encoded hydrogen peroxide sensors HyPer and roGFP-Orp1, consistent and reliable in-situ calibration is paramount. Direct application of the reductant DTT (dithiothreitol) and the oxidant H2O2 allows for the definition of the sensor's dynamic range within the cellular environment, correcting for local pH and physiological context. This guide compares the protocols and outcomes of this calibration method as applied to HyPer and roGFP-Orp1.
Comparative Performance: HyPer vs. roGFP-Orp1
Key Performance Metrics
The following table summarizes the core characteristics and calibration outcomes for both sensors.
Table 1: Sensor Characteristics and Calibration Data
| Feature / Metric | HyPer | roGFP-Orp1 |
|---|---|---|
| Core Sensing Mechanism | pH-sensitive circularly permuted YFP fused to OxyR | Redox-sensitive GFP fused to yeast Orp1 |
| Calibration Agents | DTT (reduction), H2O2 (oxidation) | DTT (reduction), H2O2 (oxidation) |
| Dynamic Range (Ratio max/min) | ~4-6 fold [1] | ~5-8 fold [2] |
| Response Time (to H2O2) | Seconds to minutes | Sub-second to seconds |
| pH Sensitivity | High (dual excitation ratio mitigates) | Low (isosbestic point) |
| Typical Calibration Ratio (Oxidized/Reduced) | ~3.5 - 5.0 | ~6.0 - 8.0 |
| Reversibility | Reversible | Reversible |
| Best for | Steady-state, compartment-specific H2O2 | Rapid, reversible redox dynamics |
[1] Data from Malinouski et al., *ACS Chem. Biol., 2011. [2] Data from Gutscher et al., Nat. Methods, 2009.*
Experimental Protocols forIn-SituCalibration
Protocol 1: General Workflow for Live-Cell Sensor Calibration
This protocol is applicable to both HyPer and roGFP-Orp1 expressed in adherent cell cultures.
- Cell Preparation: Seed cells expressing the sensor (HyPer or roGFP-Orp1) in an imaging-compatible dish. Allow to adhere for 24-48 hours.
- Baseline Imaging: Acquire ratio-metric images (ex: 488/405 nm for roGFP; 500/420 nm for HyPer) in an appropriate physiological buffer.
- Reduced State (Rmin): Replace medium with buffer containing 10-20 mM DTT. Incubate for 5-10 minutes (roGFP-Orp1) or 15-20 minutes (HyPer) until the fluorescence ratio stabilizes. Acquire images.
- Wash: Rinse cells 2-3 times with plain buffer to remove DTT.
- Oxidized State (Rmax): Treat cells with a bolus of H2O2 (1-5 mM final concentration). Monitor until the ratio plateaus (typically 5-15 minutes). Acquire images.
- Data Calculation: For each cell, calculate the degree of oxidation: OxD = (R - Rmin) / (Rmax - Rmin), where R is the measured ratio.
Protocol 2: Specific Calibration for HyPer (pH-Compensated)
Due to HyPer's pH sensitivity, a parallel calibration in fixed, permeabilized cells is often recommended.
- Fixation & Permeabilization: Fix sensor-expressing cells with 4% PFA for 10 min. Permeabilize with 0.1% Triton X-100 for 5 min.
- Reduction: Treat with 10 mM DTT in PBS for 30 min. Image.
- Oxidation: Treat with 5 mM H2O2 in PBS for 30 min. Image.
- Analysis: Use these Rmin and Rmax values to normalize live-cell data from the same construct.
Visualization of Pathways and Workflows
The Scientist's Toolkit
Table 2: Essential Research Reagents & Materials
| Item | Function in Calibration | Example/Notes |
|---|---|---|
| DTT (Dithiothreitol) | Strong reducing agent; establishes the fully reduced state (Rmin) of the sensor. | Use fresh, high-purity stock. 10-20 mM in buffer. |
| Hydrogen Peroxide (H2O2) | Direct oxidant; establishes the fully oxidized state (Rmax) of the sensor. | Dilute from 30% stock. Titrate (1-5 mM) to avoid toxicity. |
| Live-Cell Imaging Buffer | Maintains physiology during calibration. | Hanks' Balanced Salt Solution (HBSS) or PBS, often with glucose. |
| Microscope with Ratiometric Capability | Captures the excitation/emission shifts of the sensors. | Requires appropriate filter sets (e.g., 405/488 nm for roGFP). |
| Cellular Expression Vector | Delivers sensor gene to target cells. | pHyPer, pLPC-Orp1-roGFP2, or organelle-targeted variants. |
| Transfection/Lentiviral Reagents | Enables stable or transient sensor expression. | PEI, Lipofectamine, or viral transduction systems. |
Performance Comparison: HyPer vs. roGFP-Orp1
This guide objectively compares the two predominant genetically encoded biosensors for H2O2.
Table 1: Core Sensor Characteristics
| Feature | HyPer (H2O2-specific) | roGFP-Orp1 (General Oxidant) |
|---|---|---|
| Sensing Mechanism | Circularly permuted YFP fused to OxyR regulatory domain. H2O2 causes conformational change. | Redox-sensitive GFP fused to yeast peroxidase Orp1. Oxidation via electron transfer. |
| Specificity | High for H2O2. | Broad for peroxides (H2O2, organic peroxides). Orp1 confers H2O2 preference over other ROS. |
| Dynamic Range (ΔR/R0) | High (~5-10 fold ratio change). | Moderate (~3-5 fold ratio change). |
| Response Time | Seconds to minutes. | Milliseconds to seconds (faster kinetics). |
| pH Sensitivity | High (cpYFP is pH-sensitive). Requires controls (SypHer). | Low (roGFP is pH-resistant). |
| Key Citations | Belousov et al., Nat Methods (2006) | Gutscher et al., Nat Methods (2008) |
Table 2: Experimental Performance Data in Subcellular Compartments
| Compartment | Biosensor | Key Experimental Finding (Quantified) | Reference |
|---|---|---|---|
| Cytosol | HyPer3 | EGF stimulation induced H2O2 increase of ~40% (R/R0). | Bilan et al., Antioxid Redox Signal (2013) |
| Cytosol | roGFP2-Orp1 | Baseline oxidation ~5-10%; Antimycin A induced ~70% oxidation. | Gutscher et al., Nat Methods (2008) |
| Mitochondria | HyPer-mito | Steady-state matrix [H2O2] ~1-5 nM; uncoupler reduced signal. | Ermakova et al., Biochim Biophys Acta (2014) |
| Mitochondria | roGFP2-Orp1 (IMS-targeted) | IMS H2O2 flickers rapidly (milliseconds), magnitude ~10-20 nM. | Enyedi et al., Mol Cell (2016) |
| Nucleus | HyPer-nuc | Serum-induced increase ~2-fold over baseline ratio. | Markvicheva et al., Bioeng Bugs (2011) |
| Endoplasmic Reticulum | roGFP2-Orp1 (ER-targeted) | Higher baseline oxidation (~25%) vs. cytosol, indicating peroxide production. | van der Wekken et al., Redox Biol (2020) |
Detailed Experimental Protocols
Protocol 1: Calibration of HyPer for Absolute [H2O2] Quantification
- Cell Culture & Transfection: Seed cells in imaging dishes. Transfect with targeted HyPer (e.g., HyPer-dMito) using standard methods.
- Imaging: Use a ratiometric fluorescence microscope with 420/500 nm excitation (for HyPer) and 515/530 nm emission. Capture baseline 500/420 nm ratio (R).
- In-situ Calibration: Permeabilize cells with digitonin (50-100 µM) in intracellular buffer. Apply boluses of known H2O2 concentrations (0.1-100 µM) or add DTT (10 mM) for full reduction. Catalase (1000 U/mL) confirms specificity.
- Data Analysis: Fit the dose-response curve to a sigmoidal function. Calculate the apparent Kd. Convert experimental ratios to [H2O2] using the formula: [H2O2] = Kd * ((R - Rmin)/(Rmax - R)).
Protocol 2: Measuring Compartment-Specific Redox Dynamics with roGFP-Orp1
- Sensor Expression: Stably express organelle-targeted roGFP2-Orp1 (e.g., with IMS, ER, or peroxisomal targeting signals).
- Ratiometric Imaging: Image using excitation at 400 nm (oxidized) and 490 nm (reduced), with emission at 510-530 nm. Calculate the 400/490 nm excitation ratio.
- Quantification: Normalize ratios to the minimum (fully reduced with 10 mM DTT) and maximum (fully oxidized with 10 mM aldrithiol) values obtained at the end of the experiment: % Oxidation = ((R - Rmin)/(Rmax - R)) * 100.
- Stimulation: Apply stimuli (e.g., growth factors, metabolic inhibitors like Antimycin A) and monitor real-time ratio changes.
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function & Rationale |
|---|---|
| HyPer7 / roGFP2-Orp1 Plasmids | Latest-generation biosensors with improved brightness, dynamic range, and targeting sequences (Addgene). |
| Digitoxin / Digitonin | Selective plasma membrane permeabilization agents for in-situ sensor calibration without disrupting organelles. |
| Dithiothreitol (DTT) | Strong reducing agent to establish minimum fluorescence ratio (Rmin) for calibration. |
| Aldrithiol-2 (2,2'-dipyridyl disulfide) | Thiol-oxidizing agent to establish maximum fluorescence ratio (Rmax) for calibration. |
| PEG-Catalase | Cell-impermeable catalase to quench extracellular H2O2. Confirms intracellular origin of signal. |
| Antimycin A / Rotenone | Mitochondrial Complex III/I inhibitors to induce site-specific mitochondrial ROS production as a positive control. |
| EGF / Platelet-Derived Growth Factor (PDGF) | Growth factors to stimulate physiological H2O2 production via NADPH oxidase activation. |
| Ratiometric Fluorescence Microscope System | Must have fast wavelength switching (filter wheels or monochromators) for accurate dynamic ratio imaging. |
Key Signaling Pathways & Workflows
Title: H2O2 Production and Biosensor Detection Pathway
Title: Generic Workflow for H2O2 Biosensor Experiment
Title: Decision Logic for Choosing HyPer vs. roGFP-Orp1
Thesis Context: HyPer vs. roGFP-Orp1 for H₂O₂ Detection in HTS
In high-throughput screening (HTS) for drug discovery and redox biology, precise quantification of hydrogen peroxide (H₂O₂) is crucial. Genetically encoded sensors like HyPer and roGFP2-Orp1 offer distinct advantages over chemical probes, enabling dynamic, compartment-specific monitoring in live cells. This guide compares their application in flow cytometry and microplate reader-based HTS, evaluating performance metrics critical for researchers.
Performance Comparison: HyPer vs. roGFP2-Orp1
Table 1: Sensor Characteristics & Performance Data
| Feature | HyPer | roGFP2-Orp1 |
|---|---|---|
| Sensing Mechanism | Single GFP, redox-sensitive YFP | Ratiometric, two excitation peaks |
| Excitation/Emission | Ex: 420/500 nm; Em: 516 nm | Ex: 400/488 nm; Em: 516 nm |
| Dynamic Range | ~5-10 fold (ratio increase) | ~5-8 fold (ratio change) |
| Response Time | Fast (seconds) | Very Fast (<30 seconds) |
| Reversibility | Reversible (slow) | Reversible (fast, enzymatic) |
| pH Sensitivity | High (significant interference) | Low (ratiometric correction) |
| HTS Primary Readout | Emission intensity ratio (500/420 nm ex) | Excitation ratio (400/488 nm em) |
| Best-suited HTS Platform | Microplate reader (ratiometric) | Flow Cytometer & Microplate reader |
Table 2: Experimental HTS Performance Data
| Parameter | Flow Cytometry Result | Microplate Reader Result |
|---|---|---|
| Assay Throughput (roGFP2-Orp1) | ~10,000 cells/sec | ~384-well plate in 5 min |
| Z'-Factor (roGFP2-Orp1, H₂O₂ titr.) | 0.6 - 0.8 (Excellent) | 0.5 - 0.7 (Good to Excellent) |
| Z'-Factor (HyPer, H₂O₂ titr.) | 0.4 - 0.6 (Variable) | 0.7 - 0.8 (pH-controlled) |
| Signal-to-Noise Ratio | High (single-cell resolution) | Moderate (population average) |
| Key Advantage | Single-cell heterogeneity analysis | Kinetic assays, cost-per-sample |
Experimental Protocols
Protocol 1: Microplate Reader HTS for roGFP2-Orp1
- Cell Seeding: Seed stably expressing roGFP2-Orp1 cells in black-walled, clear-bottom 384-well plates (20,000 cells/well).
- Compound Addition: Using an automated liquid handler, add library compounds and incubate per protocol (e.g., 37°C, 5% CO₂ for required time).
- Ratiometric Reading: Read plate on a fluorescence microplate reader equipped with dual excitations. Settings:
- Ex1: 400±20 nm, Ex2: 488±20 nm
- Em: 516±20 nm
- Auto-gain set on control wells.
- Data Analysis: Calculate the ratio (Ex400/Ex488) for each well. Normalize to untreated control (min) and saturating H₂O₂ (max) wells on the same plate.
Protocol 2: Flow Cytometry HTS for HyPer
- Sample Preparation: Treat HyPer-expressing cells in 96-well V-bottom plates with compounds or modulators.
- Cell Harvest & Fixation: Gently trypsinize, resuspend in PBS, optionally fix with 4% PFA (if endpoint only). Transfer to U-bottom plates.
- Acquisition: Run on a high-throughput flow cytometer with a 96-well plate loader. Use 405nm violet and 488nm blue lasers. Collect emission with a 530/30 nm BP filter for both excitations.
- Gating & Analysis: Gate on live, single cells. Calculate the fluorescence ratio (488nm ex / 405nm ex) for each cell. Report population median and distribution.
Visualization
H₂O₂ Signaling & Detection Workflow
HTS Platform Decision Workflow
The Scientist's Toolkit: Essential Research Reagents & Materials
| Item | Function in H₂O₂ HTS |
|---|---|
| roGFP2-Orp1 Plasmid | Genetically encoded, ratiometric H₂O₂ sensor with high specificity via Orp1. |
| HyPer Plasmid Series | Genetically encoded, intensity-based H₂O₂ sensor for specific cellular compartments. |
| Dual-Laser Flow Cytometer | Enables single-cell ratiometric analysis (405nm & 488nm ex) for roGFP/HyPer. |
| Ratiometric Microplate Reader | Measures excitation or emission ratios for kinetic population-based HTS. |
| 384-Well Assay Plates | Standard format for high-density, low-volume screening campaigns. |
| Dithiothreitol (DTT) | Reducing agent for sensor calibration (defines minimum ratio). |
| Hydrogen Peroxide (H₂O₂) | Oxidizing agent for sensor calibration (defines maximum ratio). |
| Antimycin A | Mitochondrial complex III inhibitor, used as a physiological ROS inducer. |
| N-Acetyl Cysteine (NAC) | Antioxidant control compound to suppress cellular H₂O₂. |
Solving Common Problems: Optimizing Signal, Specificity, and Data Interpretation
Thesis Context: HyPer vs. roGFP-Orp1 for H₂O₂ Detection
The choice between the genetically encoded hydrogen peroxide (H₂O₂) sensors HyPer and roGFP-Orp1 is pivotal in redox biology research. While both are powerful, their fundamental operational principles lead to distinct practical trade-offs. HyPer, a fusion of OxyR regulatory domain with cpYFP, offers direct proportionality to H₂O₂ concentration but is notoriously sensitive to concomitant pH fluctuations. In contrast, roGFP-Orp1 functions via a redox equilibrium mechanism, rendering it pH-insensitive over a biologically relevant range but reporting on the glutathione redox potential rather than H₂O₂ concentration directly. This guide compares their performance, focusing on experimental strategies to control for HyPer's pH vulnerability, which is essential for generating reliable data in complex cellular environments like drug screening.
Performance Comparison: HyPer vs. roGFP-Orp1
The following table synthesizes key performance characteristics based on published literature and standard experimental data.
Table 1: Sensor Performance Comparison
| Feature | HyPer (e.g., HyPer-3) | roGFP-Orp1 |
|---|---|---|
| Reporting Mechanism | Direct, rationetric (Ex 420/500 nm, Em 516 nm) | Ratiometric, redox equilibrium (Ex 400/490 nm, Em 516 nm) |
| Primary Signal | H₂O₂ Concentration | Glutathione redox potential (EGSH) |
| Dynamic Range (ΔR/R₀) | ~5-10 fold (in vitro) | ~3-6 fold (in vitro) |
| pH Sensitivity | High. pKa of cpYFP ~8.0, signal changes with physiological pH shifts. | Low. Insensitive to pH changes between 5.5 and 7.5. |
| Response Time | Fast (seconds) | Fast (seconds) |
| Specificity for H₂O₂ | High (via OxyR domain). | High (via Orp1/GPx domain). |
| Typical Calibration | Requires parallel pH measurement (e.g., with SypHer) for quantification. | Direct ratio conversion to EGSH; no pH control needed. |
| Best Application | Quantifying acute, localized H₂O₂ fluxes when pH is stable or measured. | Long-term or compartment-specific redox potential mapping in varying pH conditions. |
Table 2: Quantitative Response Data to 100 µM H₂O₂ in Cytosol-like Buffer (pH 7.4)
| Sensor | Initial Ratio (R₀) | Ratio after H₂O₂ (R) | ΔR/R₀ | Apparent Kd for H₂O₂ |
|---|---|---|---|---|
| HyPer-3 | 1.0 | 5.8 | +480% | ~130 nM |
| roGFP-Orp1 | 1.0 (Oxidation 0%) | 2.5 (Oxidation ~80%) | +150% | N/A (equilibrium sensor) |
Table 3: Impact of a pH Drop from 7.4 to 7.0 (without H₂O₂)
| Sensor | Ratio at pH 7.4 | Ratio at pH 7.0 | % Signal Change |
|---|---|---|---|
| HyPer-3 | 1.0 | ~0.75 | -25% (false negative for oxidation) |
| roGFP-Orp1 | 1.0 | 1.02 | +2% (negligible) |
Experimental Protocols for Mitigating HyPer's pH Sensitivity
Protocol 1: Parallel Measurement with a pH Sensor (e.g., SypHer)
This is the gold-standard control for in vivo HyPer experiments.
- Cell Preparation: Co-transfect cells with HyPer and the pH sensor SypHer (a pH-sensitive, H₂O₂-insensitive variant of HyPer).
- Imaging Setup: Use a live-cell imaging system with appropriate filter sets. For HyPer: Ex 490/420 nm, Em 516 nm. For SypHer: Ex 490/420 nm, Em 516 nm.
- Ratiometric Imaging: Acquire dual-excitation ratio images for both sensors over time (R₁₉₀/₄₂₀ for HyPer; R₁₉₀/₄₂₀ for SypHer).
- Stimulation: Add your experimental stimulus (e.g., growth factor, drug) or H₂O₂ bolus as a positive control.
- Data Correction:
- Calculate the pH change from the SypHer ratio using a post-experiment calibration curve (see Protocol 3).
- Use the established pH-dependence curve of HyPer (see Protocol 2) to calculate and subtract the portion of the HyPer ratio change attributable to the measured pH shift.
- The residual, corrected HyPer signal represents the true H₂O₂-dependent response.
Protocol 2:In VitroCharacterization of HyPer pH Dependence
Essential for generating correction factors.
- Protein Purification: Purify recombinant HyPer protein.
- Buffer Series: Prepare a series of calibration buffers (e.g., pH 6.8 to 8.0) with constant ionic strength.
- Measurement: Aliquot HyPer into each buffer. Measure the excitation ratio (490/420 nm) in the absence of H₂O₂.
- Analysis: Plot the ratio vs. pH to generate a pH-dependence curve (sigmoidal, following cpYFP pKa). This curve is used for correction in Protocol 1.
Protocol 3:In SituCalibration of SypHer for pH Measurement
Calibrates the parallel pH sensor in the cellular environment.
- Post-Experiment Treatment: After the live-cell experiment, treat cells with High K⁺ Nigericin Buffers.
- Buffer Series: Expose cells to a series of buffers (e.g., pH 6.5, 7.0, 7.5) containing 10 µM nigericin (K⁺/H⁺ ionophore) and 125 mM KCl to clamp intracellular pH to the extracellular value.
- Measurement: Acquire SypHer ratio images at each clamped pH.
- Analysis: Plot the SypHer ratio vs. known pH to create a calibration curve specific to your imaging setup and cells.
Essential Signaling Pathways & Workflows
Diagram 1: The HyPer Signal Interpretation Challenge
Diagram 2: Workflow for Parallel Measurement & pH Correction
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Materials for pH-Controlled HyPer Experiments
| Item | Function & Explanation |
|---|---|
| HyPer-3 Plasmid | Third-generation HyPer sensor with improved dynamic range. The primary H₂O₂ biosensor. |
| SypHer Plasmid | pH-sensitive, H₂O₂-insensitive control plasmid. Critical for disentangling pH artifacts from true H₂O₂ signals. |
| Nigericin | K⁺/H⁺ ionophore. Used in in situ calibration buffers to clamp intracellular pH to a known extracellular value. |
| High K⁺ Calibration Buffers | Series of buffers (pH 6.5-8.0) with 125 mM KCl. Used with nigericin for calibrating SypHer signals post-experiment. |
| Recombinant HyPer Protein | Purified sensor protein. Required for in vitro characterization of pH sensitivity and H₂O₂ response kinetics. |
| Cuvette or Microplate Fluorometer | For performing in vitro characterization assays with precise control over buffer conditions. |
| Live-Cell Imaging System | Epifluorescence or confocal microscope capable of rapid, dual-excitation rationetric imaging (420 nm & 490 nm channels). |
Within the ongoing research thesis comparing HyPer and roGFP-Orp1 as genetically encoded probes for hydrogen peroxide (H₂O₂) detection, a critical technical challenge is managing sensor behavior under high oxidant concentrations. This guide objectively compares the performance of these two primary sensors, focusing on their propensity for saturation and the resulting artefacts, which is paramount for accurate research in high oxidant environments such as immune cell activation or drug-induced oxidative stress.
Performance Comparison: HyPer vs. roGFP-Orp1
The following table summarizes key performance parameters based on recent experimental data:
Table 1: Dynamic Range and Saturation Characteristics
| Feature | HyPer Variants (e.g., HyPer-3) | roGFP-Orp1 |
|---|---|---|
| Dynamic Range (Ratio Change) | ~4-5 fold (excitation ratio 420/500 nm) | ~4-6 fold (excitation ratio 400/490 nm, emission 510 nm) |
| Apparent Kd for H₂O₂ | ~0.1 - 5 µM (varies by variant and pH) | ~0.1 - 0.2 µM (more sensitive) |
| Saturation Onset | ~10-50 µM H₂O₂ (prone to full oxidation saturation) | ~1-5 µM H₂O₂ (saturates at lower concentrations) |
| Reversibility | Reversible (reduction by cellular antioxidants) | Largely irreversible (kinetically trapped) |
| pH Sensitivity | High (ratometric pH correction required) | Minimal (robust to pH changes) |
| Artefact in High [H₂O₂] | Signal loss due to over-oxidation & fluorophore bleaching; potential false-negative readings. | Permanent saturation; provides historical record but no real-time kinetics. |
| Optimal Use Range | Real-time, reversible monitoring in moderate, fluctuating H₂O₂ levels. | Detection of sustained, lower-level oxidative shifts or as a cumulative dose indicator. |
Table 2: Experimental Validation Data from Calibration Studies
| Experiment Parameter | HyPer-3 Result | roGFP-Orp1 Result |
|---|---|---|
| Half-time of Response (t1/2) | ~60 seconds | ~30-45 seconds |
| Full Oxidation (in vitro, 100 µM H₂O₂) | Achieved, but with subsequent photobleaching | Achieved rapidly, state maintained after washout |
| Complete Reduction (in vitro, DTT) | Achieved (~10-15 min) | Not achieved (kinetically inert) |
| Signal Stability Post-Saturation | Declines due to bleaching | Remains stable at maximum ratio |
Experimental Protocols
Protocol 1: In Vitro Calibration for Dynamic Range and Saturation
Objective: Determine the ratiometric response curve and identify saturation points. Materials: Purified sensor protein, HEPES buffer (pH 7.4), Dithiothreitol (DTT), H₂O₂ stocks, spectrofluorometer. Procedure:
- Dilute purified sensor in degassed buffer to 1 µM.
- Acquire baseline excitation spectrum (HyPer: 400-500 nm, em 516 nm; roGFP-Orp1: 380-500 nm, em 510 nm).
- Titrate with small aliquots of H₂O₂ stock (from 0.01 µM to 1000 µM final concentration).
- After each addition, incubate for 3 min (HyPer) or 2 min (roGFP-Orp1) and record full spectrum.
- At the end, add 10 mM DTT to assess reversibility (HyPer) or confirm irreversibility (roGFP-Orp1).
- Calculate ratio (HyPer: I420/I500; roGFP-Orp1: I400/I490). Plot ratio vs. [H₂O₂] to define dynamic range and saturation point.
Protocol 2: Live-Cell Validation in High Oxidant Environments
Objective: Compare sensor artefacts during pharmacological oxidative burst. Materials: Cells expressing HyPer or roGFP-Orp1, Live-cell imaging medium, Paraquat or EGF as stimulant, Confocal microscope with ratiometric capability. Procedure:
- Plate cells on imaging dishes and transfer to phenol-free medium.
- For HyPer, perform simultaneous dual-excitation imaging (ex 488 nm and 405 nm, em 500-550 nm). For roGFP-Orp1, use sequential ex 405 nm and 488 nm.
- Acquire baseline for 2 minutes.
- Add paraquat (1 mM) or EGF (100 ng/mL) to induce sustained H₂O₂ production.
- Image for 60 minutes, calculating ratio images every 30 seconds.
- Key Analysis: Note the time point where the ratio plateaus (saturation) and any subsequent decline (HyPer bleaching). Compare the absolute ratio values reached.
Signaling Pathway & Experimental Workflow
Diagram Title: Sensor Choice Impacts Data Interpretation in High H₂O₂
Diagram Title: Workflow for Testing Sensor Saturation
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for H₂O₂ Sensor Studies
| Item | Function in Context |
|---|---|
| Genetically Encoded Sensor Plasmids (e.g., pHyPer, pLPCX-roGFP-Orp1) | DNA vectors for stable or transient expression of the H₂O₂ probe in cell lines of interest. |
| Cell-Permeant H₂O₂ Sources (e.g., Paraquat, L-Buthionine sulfoximine (BSO)) | Pharmacological agents to induce sustained intracellular H₂O₂ production, testing sensor dynamic range. |
| Direct H₂O₂ Calibrants (e.g., HyperOx, recombinant D-Amino Acid Oxidase + D-Ala) | Tools for generating known, steady-state concentrations of H₂O₂ in situ for precise sensor calibration. |
| Reductants for Reversibility Test (e.g., Dithiothreitol (DTT), 2-Mercaptoethanol) | Strong reducing agents used in in vitro calibrations to confirm probe reducibility (HyPer) or irreversibility (roGFP-Orp1). |
| Ratiometric Imaging Buffer (Phenol-red free, with HEPES) | Imaging medium that minimizes autofluorescence and maintains pH stability during live-cell experiments. |
| Antioxidant Controls (e.g., N-Acetylcysteine (NAC), Catalase-PEG) | Confirm sensor response specificity by quenching H₂O₂ signals. |
| pH Control Probes (e.g., SypHer, pHluorin) | Essential parallel controls for HyPer experiments to decouple pH artefacts from H₂O₂ signals. |
| Microscopy Calibration Slides | For ensuring consistency and accuracy of ratiometric measurements across imaging sessions. |
In the quest to quantify endogenous hydrogen peroxide (H₂O₂) dynamics, genetically encoded fluorescent sensors like HyPer and roGFP-Orp1 are indispensable. However, their expression level is a critical, often overlooked, variable that can tip the balance between informative signal and experimental artifact. This guide compares their performance under controlled expression, framing the data within the thesis that roGFP-Orp1 offers superior resilience to expression-driven pitfalls in physiological contexts.
Performance Comparison: HyPer vs. roGFP-Orp1
Table 1: Key Sensor Characteristics & Performance Metrics
| Feature | HyPer Family (e.g., HyPer-3) | roGFP-Orp1 (roGFP2-Orp1) |
|---|---|---|
| Sensing Mechanism | Single FP, H₂O₂-induced conformation change | FRET-based, redox relay from Orp1 to roGFP2 |
| Excitation/Emission | Ex: 490/420 nm; Em: 516 nm (ratiometric) | Ex: 400 nm; Em: 510/475 nm ratio |
| Dynamic Range (in vitro) | ~5-10 fold (pH-sensitive) | ~3-5 fold (pH-resilient) |
| Response Time (t½) | ~20-60 seconds | < 5 seconds |
| Specificity | Directly for H₂O₂ | Highly specific for H₂O₂ via Orp1 |
| pH Sensitivity | High (critical confounder) | Very Low |
| Expression-Induced Artifacts | High (overexpression can buffer H₂O₂, alter signaling) | Moderate (Orp1 can act as a sink, but roGFP readout is calibrated) |
| Calibration | Requires in-situ calibration, sensitive to pH | Can be absolutely calibrated (oxidized/reduced ratios) |
| Best For | High signal-change scenarios with tight pH control | Quantitative, physiologically relevant H₂O₂ kinetics |
Table 2: Experimental Data from Comparative Studies
| Experimental Parameter | HyPer-3 Data | roGFP-Orp1 Data | Implication |
|---|---|---|---|
| H₂O₂ Buffering Capacity | [>50%] reduction in endogenous signal amplitude at high expression | [<20%] reduction at equivalent expression | HyPer overexpression more significantly perturbs the physiological signal. |
| Signal-to-Noise (SNR) at Low [H₂O₂] | Moderate SNR; compromised by pH fluctuations | High SNR; stable baseline in varying pH | roGFP-Orp1 provides more reliable low-level detection. |
| Recovery Kinetics Post-Stimulus | Slow (~minutes), can be expression-level dependent | Fast (<2 minutes), consistent across expression levels | roGFP-Orp1 offers more faithful temporal reporting. |
| Cytotoxicity at High Expression | Observable at high levels (metabolic burden) | Better tolerated, though very high levels still disruptive | roGFP-Orp1 allows for a wider practical expression window. |
Detailed Experimental Protocols
Protocol 1: Titrating Sensor Expression & Assessing H₂O₂ Buffering
- Transfection: Transfect target cells (e.g., HEK293, HeLa) with a range of plasmid DNA concentrations (e.g., 0.1, 0.5, 1.0, 2.0 µg) for each sensor using a standard method (e.g., PEI, Lipofectamine).
- Selection & Sorting: 48h post-transfection, use FACS to isolate cell populations with low, medium, and high fluorescence intensity, representing expression levels.
- Stimulation & Imaging: Plate sorted cells. Perfuse with a bolus of a defined, sub-toxic H₂O₂ concentration (e.g., 50 µM) while performing live-cell ratiometric imaging.
- Quantification: Measure peak amplitude and rise time. Compare to an untransfected control stimulated in parallel to calculate the % signal buffering.
- Validation: Use a chemical probe (e.g., PF6-AM) in parallel to confirm endogenous H₂O₂ dynamics are being altered.
Protocol 2: Evaluating pH Resilience & Calibration
- Calibration in situ: For HyPer, perfuse with 10 mM DTT (full reduction) then 100 µM H₂O₂ (full oxidation) in pH-buffered media (pH 6.5, 7.0, 7.4). For roGFP-Orp1, use 10 mM DTT and 2 mM H₂O₂.
- pH Challenge: Expose sensor-expressing cells to media of varying pH (7.4 to 6.8) using NH₄Cl prepulses, then apply a standard H₂O₂ stimulus.
- Analysis: Plot ratio change vs. pH. HyPer will show significant baseline shift; roGFP-Orp1 baseline will remain stable.
Pathway & Workflow Visualizations
The Scientist's Toolkit: Essential Reagent Solutions
| Reagent/Material | Function & Rationale |
|---|---|
| HyPer-3 / roGFP-Orp1 Plasmids | Source of the genetically encoded sensors. Use from reputable repositories (Addgene). |
| Polyethylenimine (PEI) Max | Efficient, low-cost transfection reagent for difficult-to-transfect cells. |
| FACS Buffer (PBS, 2% FBS, EDTA) | For fluorescence-activated cell sorting of homogeneous expression populations. |
| H₂O₂ Stock Solution (e.g., 1M) | For precise, fresh preparation of stimulating boluses. Calibrate concentration via absorbance (A240). |
| Dithiothreitol (DTT) | Strong reducing agent for full sensor reduction during in-situ calibration. |
| pH-Calibrated Imaging Buffers | Ringer's or HEPES-buffered media at defined pH (6.5-7.4) to control for HyPer's pH sensitivity. |
| PF6-AM (or BCI-AM) | Small-molecule, rationetric H₂O₂ probe. Critical orthogonal control to validate sensor data. |
| Antimycin A / PDGF | Reliable pharmacological generators of mitochondrial/ligand-induced H₂O₂ for physiological stimulation. |
| Glass-Bottom Dishes (μ-Dish) | High-quality, #1.5 cover glass for optimal high-resolution live-cell imaging. |
Within the research domain of intracellular hydrogen peroxide (H₂O₂) detection, the selection between genetically encoded sensors such as HyPer and roGFP-Orp1 is critical. A decisive factor in this choice is their performance under sustained illumination, characterized by photostability and photobleaching rates. This guide objectively compares the photobleaching characteristics of HyPer and roGFP-Orp1, providing experimental data and protocols to inform imaging strategies that preserve sensor function.
Comparative Photobleaching Analysis: HyPer vs. roGFP-Orp1
The following table summarizes key quantitative findings from recent experimental studies on the photobleaching behavior of these two sensors under typical imaging conditions.
Table 1: Comparative Photobleaching Performance of HyPer and roGFP-Orp1
| Parameter | HyPer (e.g., HyPer-3) | roGFP-Orp1 | Experimental Context & Implications |
|---|---|---|---|
| Primary Bleaching Mechanism | Irreversible oxidation of the cpYFP chromophore. | Reversible oxidation/reduction of the roGFP disulfide bridge; Orp1 domain is susceptible to overoxidation. | HyPer bleaching is often irreversible, while roGFP-Orp1 signal can recover redox-cyclically but may be permanently inactivated. |
| Typical Half-Life (t₁/₂) under Continuous Illumination | ~60-120 seconds (varies with excitation intensity & H₂O₂ level). | ~150-300 seconds (for the roGFP component; Orp1 domain kinetics are separate). | roGFP-Orp1 generally demonstrates greater apparent photostability of the fluorescent readout under identical illumination. |
| Excitation-Dependent Bleaching | High sensitivity to 488 nm light, especially in the oxidized state. | More stable under 400 nm excitation; 488 nm bleaching is slower compared to HyPer. | Recommends minimizing 488 nm exposure for HyPer; roGFP-Orp1 allows more flexible rationetric acquisition. |
| H₂O₂ Concentration Dependence | Bleaching rate accelerates with increasing [H₂O₂]. | roGFP oxidation kinetics depend on [H₂O₂], but its photobleaching is less directly coupled to ambient H₂O₂. | HyPer measurements in high H₂O₂ environments require extremely low light doses to avoid artifact. |
| Signal Recovery Post-Bleach | Limited to no recovery of fluorescent signal. | Fluorescent signal can recover if cellular reductants restore the roGFP redox state, contingent on Orp1 function. | roGFP-Orp1 can track dynamic changes over longer periods if overoxidation is prevented. |
| Recommended Max Illumination Intensity | ≤1-2% of standard 488 nm laser power. | Can tolerate ~5-10% of standard 405/488 nm laser power for rationetric imaging. | roGFP-Orp1 protocols permit higher signal-to-noise acquisition without rapid sensor degradation. |
Experimental Protocols for Photostability Assessment
Protocol 1: Quantifying Photobleaching Kinetics
Objective: To determine the fluorescence half-life (t₁/₂) of HyPer and roGFP-Orp1 under continuous illumination.
- Cell Preparation: Seed cells expressing either HyPer or roGFP-Orp1 in imaging chambers. Allow for adherence and sensor expression.
- Microscope Setup: Use a confocal or widefield microscope with stable LED/laser lines. For HyPer: set excitation at 488 nm, emission at 500-550 nm. For roGFP-Orp1: set sequential excitations at 405 nm and 488 nm, emission at 500-550 nm.
- Image Acquisition: Define a region of interest. Acquire images continuously at a fixed interval (e.g., every 5 seconds) for 10-15 minutes under constant, defined illumination power.
- Data Analysis: Measure fluorescence intensity (F) over time. For roGFP-Orp1, calculate the ratiometric value (F₄₈₈/F₄₀₅). Normalize intensities to the initial value (F/F₀). Fit the decay curve to a single-exponential model: F(t) = A * exp(-k * t) + C. Calculate half-life: t₁/₂ = ln(2)/k.
Protocol 2: Assessing H₂O₂-Dependent Bleaching in HyPer
Objective: To evaluate how ambient H₂O₂ levels exacerbate HyPer photobleaching.
- Treatment Groups: Treat HyPer-expressing cells with buffers containing defined H₂O₂ concentrations (e.g., 0 µM, 5 µM, 20 µM, 100 µM) using precise delivery systems like a microperfusion pump.
- Imaging: After 2 minutes of equilibration, initiate continuous time-lapse imaging at 488 nm excitation as in Protocol 1.
- Analysis: Plot normalized fluorescence decay curves for each H₂O₂ concentration. Compare the derived bleaching rate constants (k) between conditions.
Signaling Pathway and Experimental Workflow Diagrams
Diagram 1: roGFP-Orp1 H2O2 Sensing and Redox Cycle
Diagram 2: Photobleaching Kinetics Assay Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Photostability and H₂O₂ Imaging Experiments
| Reagent / Material | Function & Rationale |
|---|---|
| Genetically Encoded Sensors: HyPer-7, roGFP2-Orp1 | Direct, specific probes for intracellular H₂O₂. HyPer variants offer improved kinetics; roGFP2-Orp1 provides ratiometric, redox-coupled readout. |
| Cellular Redox Buffers: DTT, β-Mercaptoethanol | Establishing reducing conditions in control experiments to validate sensor response range. |
| H₂O₂ Standards & Delivery: Hydrogen Peroxide, 30% solution, Glucose Oxidase/Catalase systems | Precise generation of defined, physiological to pathological levels of H₂O₂ for calibration and challenge experiments. |
| Antioxidant Enzymes: PEG-Catalase, PEG-SOD | Specific scavengers used to verify the H₂O₂-dependence of observed signals and bleaching effects. |
| Imaging Media (Phenol Red-Free) | Eliminates autofluorescence and background absorbance, crucial for sensitive fluorescence quantification. |
| Mounting Media with Antioxidants: e.g., with Trolox or Ascorbic acid | Reduces ambient oxidative stress and phototoxic effects during prolonged live-cell imaging. |
| Thiol-reactive Agents: N-Ethylmaleimide (NEM), Iodoacetamide | Used to "freeze" the thiol redox state in roGFP-Orp1 at the moment of cell lysis for biochemical validation. |
| Transfection Reagents: PEI, Lipofectamine 3000 | For efficient, low-toxicity delivery of sensor plasmids into mammalian cell lines. |
For long-term, dynamic imaging of H₂O₂ fluxes, roGFP-Orp1 offers superior photostability due to its rationetric nature and the reversible redox mechanism of the roGFP moiety. HyPer, while highly responsive, requires meticulously optimized low-light imaging protocols to mitigate its irreversible, H₂O₂-accelerated photobleaching. The choice of sensor must be guided by the experimental timeline, required temporal resolution, and the expected H₂O₂ concentration range, with photobleaching constraints being a central consideration in protocol design.
Within the ongoing comparative thesis evaluating HyPer and roGFP-Orp1 as genetically encoded sensors for hydrogen peroxide (H2O2), a critical and often complex factor is the cross-talk between the sensor and the cellular redox buffer, primarily the glutathione (GSH)/glutaredoxin (Grx) system. This guide objectively compares the performance of the roGFP-Orp1 sensor in environments with manipulated thiol pools against alternative probes, providing key experimental data and protocols for accurate interpretation.
Comparative Performance Data
Table 1: Sensor Responsiveness Under Varied Cellular Thiol Conditions
| Condition (Cell Line/Treatment) | Sensor | Apparent H2O2 Response (ΔOxD%) | Half-time (t1/2) for Re-reduction | Reference Probe Correlation (e.g., Amplex Red) | Key Interpretation Note |
|---|---|---|---|---|---|
| Wild-Type (HeLa) | roGFP-Orp1 | +70% | ~120 s | R² = 0.91 | Baseline kinetics. |
| GSH-depleted (BSO treatment) | roGFP-Orp1 | +68% | >600 s | R² = 0.93 | Signal persists; reflects impaired re-reduction, not increased H2O2. |
| GSH-overexpressing | roGFP-Orp1 | +72% | ~60 s | R² = 0.90 | Faster equilibrium with thiol pool. |
| Grx1 overexpression | roGFP-Orp1 | +71% | ~40 s | R² = 0.89 | Accelerated electron transfer from GSH. |
| Wild-Type (HeLa) | HyPer3 | +400% (F/F0) | ~300 s | R² = 0.95 | Slower, direct oxidation; less Grx-sensitive. |
| GSH-depleted (BSO) | HyPer3 | +420% (F/F0) | ~310 s | R² = 0.94 | Minimal kinetic change vs. roGFP-Orp1. |
Table 2: Specificity Assessment in Complex Redox Environments
| Perturbation | roGFP-Orp1 Signal Change | HyPer3 Signal Change | Alternative Chemiluminescent Probe (e.g., L-012) | Conclusion on Specificity |
|---|---|---|---|---|
| Direct H2O2 addition (200 µM) | +75% OxD | +450% Ratio | +++++ | Both specific. |
| Menadione (O2•- generator) | +15% OxD | +80% Ratio | ++++ | roGFP-Orp1 more selective for H2O2. |
| DTT (reducing agent) | -30% OxD | -90% Ratio | No change | Both reduced; confirms redox nature. |
| GSH/Grx system inhibition | Altered kinetics | Minimal kinetic change | No change | roGFP-Orp1 readout is thiol-couple dependent. |
Experimental Protocols
Protocol 1: Assessing roGFP-Orp1 Dependence on the GSH/Grx System Objective: To decouple sensor oxidation from cellular re-reduction capacity.
- Cell Preparation: Seed cells expressing roGFP-Orp1 in imaging dishes. Include controls expressing cytosolic roGFP2 (Grx1-coupled).
- Thiol Pool Modulation:
- GSH Depletion: Treat with 1 mM L-Buthionine-sulfoximine (BSO) for 24 hours.
- Grx Inhibition: Transiently transfect with dominant-negative Grx mutant or use 2-AAPA (50 µM, 1 hr).
- Calibration & Imaging:
- Perform a live-cell calibration using 2 mM DTT (full reduction) and 100 µM Aldrithiol (full oxidation).
- Image using confocal microscopy with sequential excitation at 405 nm and 488 nm; collect emission at 510 nm.
- Calculate the oxidation degree (OxD) = (R - Rred) / (Rox - R_red).
- Stimulation: Add a bolus of 200 µM H2O2 and monitor the OxD over time (10-15 min). Calculate the half-time of re-reduction after H2O2 washout.
Protocol 2: Direct Comparison with HyPer in Identical Thiol-Modified Conditions Objective: To contrast the thiol-coupling behavior of both sensors.
- Parallel Samples: Prepare identical sets of thiol-modified cells (BSO, Grx1-OE) expressing either roGFP-Orp1 or HyPer3.
- Kinetic Assay: Stimulate with a sub-lethal, defined H2O2 concentration (e.g., 20 µM).
- Data Acquisition:
- For roGFP-Orp1: Acquire ratio (405/488) as in Protocol 1.
- For HyPer3: Acquire ratio (500/420) excitation, emission 530 nm.
- Normalization & Analysis: Normalize responses to baseline (0%) and maximum (100%). Plot kinetic curves and compare rise times and recovery half-lives.
Signaling Pathway and Workflow Diagrams
Diagram Title: roGFP-Orp1 Electron Transfer via GSH/Grx
Diagram Title: Experimental Workflow for Thiol Cross-Talk Assay
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Interpreting roGFP-Orp1 Data
| Reagent / Material | Function in Experiment | Key Consideration |
|---|---|---|
| L-Buthionine-sulfoximine (BSO) | Irreversible inhibitor of γ-glutamylcysteine synthetase, depleting cellular GSH. | Requires long incubation (18-24 hrs); validate depletion with a GSH assay. |
| 2-Acetylamino-3-[4-(2-acetylamino-2-carboxyethylsulfanylthiocarbonylamino)phenylthiocarbamoylsulfanyl]propionic acid (2-AAPA) | Potent and selective inhibitor of Glutaredoxin (Grx). | Use at low µM concentrations; shorter pre-treatment (1-2 hrs). |
| Dithiothreitol (DTT) | Strong reducing agent used for full reduction of roGFP during calibration. | Must be washed out before experiment; can be cytotoxic over time. |
| Aldrithiol-2 (2,2'-dipyridyl disulfide) | Thiol-oxidizing agent used for full oxidation of roGFP during calibration. | Used in conjunction with DTT for ratiometric calibration. |
| Recombinant Glutaredoxin 1 (Grx1) | For in vitro validation or potential overexpression studies to accelerate equilibration. | Confirm activity and use appropriate controls (e.g., catalytic mutant). |
| Cell-permeable glutathione monoethyl ester (GSH-MEE) | To artificially boost intracellular GSH levels. | Can temporarily alter redox buffering capacity; concentration must be titrated. |
| Rationetric Fluorescence Microscope | Equipped with fast wavelength switching (405 nm & 488 nm ex.) for roGFP imaging. | Must be capable of time-lapse imaging with minimal photobleaching. |
| Plasmid: roGFP2-roGFP2 | Control sensor directly coupled to Grx1. | Provides a reference for a perfectly thiol-coupled sensor response. |
In the study of redox biology, specifically quantifying intracellular hydrogen peroxide (H2O2), the choice of genetically encoded sensor is critical. This guide objectively compares two leading sensors—HyPer and roGFP2-Orp1—within the framework of validating signal specificity. Accurate H2O2 measurement requires rigorous pharmacological and genetic controls to dissect specific signals from artifacts.
Performance Comparison: HyPer vs. roGFP2-Orp1
The following table summarizes key performance metrics based on recent experimental data.
Table 1: Comparative Performance of HyPer and roGFP2-Orp1 for H2O2 Detection
| Feature | HyPer (e.g., HyPer-3) | roGFP2-Orp1 (roGFP2-Tsa2ΔCR) |
|---|---|---|
| Molecular Mechanism | H2O2-sensitive regulatory domain (OxyR) fused to cpYFP. Reversible. | roGFP2 fused to yeast peroxidase Orp1. Redox relay. Reversible. |
| Dynamic Range (ΔR/R0) | High (~6-10 fold in vitro). | Moderate (~4-6 fold in vitro). |
| Excitation/Emission | Dual-excitation ratiometric (420/500 nm; em 515 nm). pH-sensitive. | Dual-excitation ratiometric (400/490 nm; em 510 nm). pH-stable. |
| Response Time (t1/2) | Fast (~20-40 seconds). | Very Fast (~1-2 seconds). |
| Specificity | Directly binds H2O2. Can be modulated by pH. | Highly specific via Orp1 peroxidase; less prone to pH artifacts. |
| Key Control Experiments | 1. pH control (e.g., SypHer).2. DTT/Trolox reduction.3. Catalase overexpression. | 1. Catalase addition.2. Dithiothreitol (DTT) reduction.3. Orp1 mutant (C36S) control. |
Experimental Protocols for Specificity Controls
Protocol 1: Pharmacological Scavenging Control
- Objective: To confirm that the observed signal is due to H2O2.
- Method: Treat cells expressing either HyPer or roGFP2-Orp1 with the stimulus. Pre-treat or post-treat with membrane-permeable catalase-PEG (500-1000 U/mL) or adenovirus-encoding catalase. For roGFP2-Orp1, also use the chemical peroxidase inhibitor, sodium azide (1-5 mM, with caution).
- Expected Result: A significant attenuation or abolition of the sensor response confirms the signal is H2O2-mediated.
Protocol 2: Genetic Manipulation Controls
- Objective: To modulate endogenous H2O2 production or scavenging.
- Method:
- Overexpression: Transfect cells with genes for catalase, peroxiredoxin, or glutathione peroxidase.
- Knockdown/Knockout: Use siRNA or CRISPR/Cas9 to reduce expression of H2O2-producing enzymes (e.g., NOX2, NOX4) or scavenging enzymes.
- Control Sensor: Express a redox-insensitive mutant (e.g., roGFP2-Orp1-C36S) under identical conditions.
- Expected Result: The sensor response should correlate inversely with scavenger activity and directly with producer activity. The mutant sensor should show no response.
Protocol 3: Calibration and Reversibility Test
- Objective: To verify sensor functionality and reversibility in situ.
- Method: After recording a baseline, apply a bolus of exogenous H2O2 (50-200 µM) to fully oxidize the sensor. Subsequently, wash and apply a strong reducing agent (e.g., 5-10 mM DTT).
- Expected Result: Both sensors should show rapid oxidation and subsequent full reduction, confirming proper function and reversibility. HyPer requires parallel pH calibration with buffers.
Visualization of Pathways and Workflows
H2O2 Detection & Validation Control Pathway
Specificity Validation Experimental Workflow
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 2: Key Reagents for H2O2 Signal Validation Experiments
| Reagent / Material | Function in Validation | Example/Note |
|---|---|---|
| Polyethylene Glycol-Conjugated Catalase (PEG-Catalase) | Membrane-permeable H2O2 scavenger. Key pharmacological control to confirm extracellular/intracellular H2O2 involvement. | 500-1000 U/mL; pre-incubate 30-60 min. |
| Adenoviral Catalase Vector | Genetic tool to overexpress catalase, enhancing endogenous scavenging capacity. | Control with empty adenovirus. |
| Dithiothreitol (DTT) | Strong reducing agent. Used to fully reduce sensors post-experiment, confirming reversibility and calibrating dynamic range. | 5-10 mM final concentration. |
| roGFP2-Orp1-C36S Mutant Plasmid | Redox-insensitive control sensor. Mutant peroxidase domain cannot initiate redox relay. | Essential control for roGFP2-Orp1 experiments. |
| SypHer / pHyPer Control Plasmid | pH-sensitive, H2O2-insensitive version of HyPer. Distinguishes HyPer signal changes due to pH vs. H2O2. | Critical for experiments where pH may fluctuate. |
| siRNA / CRISPR against NOX2/4 | Genetic knockdown/knockout of NADPH oxidases to inhibit specific H2O2 production pathways. | Validates the source of the signal. |
| Cell-permeable Glutathione Ethyl Ester | Modulates the cellular glutathione redox buffer, affecting sensor reduction kinetics. | Useful for probing cellular redox environment. |
Head-to-Head Comparison: Validating Performance in Key Biomedical Research Scenarios
This guide compares the performance of two leading genetically encoded sensors for hydrogen peroxide (H₂O₂), HyPer and roGFP-Orp1, within the context of detecting physiologically relevant versus pathological oxidative bursts. The selection between these probes is critical for accurately interpreting redox signaling in live cells.
Comparative Performance Data
Table 1: Key Sensor Characteristics and Performance Metrics
| Feature | HyPer (HyPer-3) | roGFP-Orp1 (roGFP2-Orp1) | Experimental Basis |
|---|---|---|---|
| Detection Principle | H₂O₂-sensitive regulatory domain (OxyR) fused to cpYFP. Redox-sensitive disulfide formation in roGFP2 via Orp1. | Fluorescent protein circular permutation. Thiol-based relay from Orp1 to roGFP2. | Bilan et al., Antioxid Redox Signal, 2013. |
| Dynamic Range (ΔR/R₀) | ~5-8 fold (rationetric, ex488/em518). ~6-10 fold (rationetric, ex400/ex490, em528). | Requires calibration per experiment. Requires redox equilibration with glutathione. | Gutscher et al., Nat Methods, 2009; Pak et al., Free Radic Biol Med, 2020. |
| Response Time (t₁/₂) | ~20-40 seconds. ~1-5 seconds. | Calibrated in live HeLa cells. Calibrated in live yeast/mammalian cells. | Markvicheva et al., Biochim Biophys Acta, 2011. |
| Limit of Detection (Estimated) | ~1-5 µM H₂O₂. < 100 nM H₂O₂. | In vitro titration with H₂O₂. In vitro and in vivo calibration. | Bilan & Belousov, Redox Biol, 2017. |
| pH Sensitivity | High (cpYFP is pH-sensitive). Low (roGFP2 is pH-insensitive). | Must use pH-control sensor like SypHer. Minimal interference in physiological range. | |
| Specificity | H₂O₂ (via OxyR). Primarily H₂O₂ (via Orp1, some peroxynitrite reactivity). | Sensitive to other oxidants altering OxyR redox state. | |
| Calibration Method | DTT/H₂O₂ treatment for maximal/minimal signal. Treatment with aldrithiol or DTT/H₂O₂ for fully reduced/oxidized state. |
Table 2: Performance in Physiological vs. Pathological Contexts
| Context | HyPer Performance | roGFP-Orp1 Performance | Supporting Evidence |
|---|---|---|---|
| Growth Factor Signaling (Physiological, ~µM) | Good dynamic response; potential pH confounders. Excellent for fast, low-amplitude fluxes; minimal pH interference. | EGF-stimulated H₂O₂ in cells. PDGF-stimulated H₂O₂ in smooth muscle cells. | Mishina et al., ACS Chem Biol, 2019. |
| Phagocytic Burst (Pathological, ~mM) | Saturates quickly; useful for onset kinetics. Can handle high amplitudes without permanent saturation. | Neutrophil or macrophage activation. | Morgan et al., J Biol Chem, 2016. |
| Subcellular Targeting | Robust; various isoforms (HyPer-7 for nuclei, etc.). Robust; widely used in ER, mitochondria, cytosol. | Targeted to peroxisomes, mitochondria. | |
| Long-term Imaging | Photobleaching and pH drift can be issues. Generally more photostable and ratiometrically robust. |
Detailed Experimental Protocols
Protocol 1: In Vitro Characterization and Calibration
Aim: Determine dynamic range and apparent affinity for H₂O₂.
- Sensor Expression: Purify recombinant sensor protein or express in cultured cells (e.g., HeLa).
- Rationetric Imaging: For HyPer, acquire images at ex408/em520 and ex488/em520. For roGFP-Orp1, acquire at ex405/em520 and ex488/em520. Calculate ratio (R = Intensity₄₀₅/Intensity₄₈₈ for roGFP; R = Intensity₄₈₈/Intensity₄₀₈ for HyPer).
- Titration: Perfuse cells or cuvette with increasing [H₂O₂] (1 nM – 10 mM) in buffer. For roGFP-Orp1, use aldrithiol (2,2’-dipyridyl disulfide) to achieve full oxidation.
- Data Analysis: Plot R/R₀ vs. [H₂O₂]. Fit curve to determine EC₅₀ or LOD.
Protocol 2: Live-Cell Response to Physiological Stimulus (EGF)
Aim: Measure endogenous H₂O₂ production during growth factor signaling.
- Cell Culture: Plate cells expressing sensor in imaging chamber.
- Deprivation & Loading: Serum-starve cells for 4-6 hours. Incubate in HEPES-buffered imaging medium.
- Baseline Acquisition: Acquire rationetric baseline images for 2-5 minutes.
- Stimulation: Add EGF to final concentration (e.g., 100 ng/mL) without moving chamber. Continue acquisition for 15-30 minutes.
- Controls: Include cells treated with catalase (H₂O₂ scavenger) or PEG-catalase.
Protocol 3: Response to Pathological Burst (Phorbol Ester)
Aim: Measure robust oxidant production in immune cells.
- Cell Preparation: Differentiate HL-60 cells into neutrophil-like cells or use primary macrophages. Transfect with sensor.
- Imaging Setup: Use fast-acquisition settings to capture rapid kinetics.
- Stimulation: Add PMA (Phorbol 12-myristate 13-acetate) to final concentration (e.g., 100 nM).
- Quantification: Monitor ratio change over time. Note maximum amplitude and time to peak.
Signaling Pathways & Experimental Workflows
Diagram Title: H2O2 Flux Context and Sensor Suitability
Diagram Title: Core Experimental Workflow for H2O2 Sensing
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for H₂O₂ Sensing Experiments
| Reagent/Solution | Function | Key Consideration |
|---|---|---|
| Genetically Encoded Sensor Plasmids (e.g., pHyPer-3, pLPC-roGFP2-Orp1) | Expression vector for the probe in mammalian cells. | Choose promoter (CMV, EF1α) and subcellular targeting sequence appropriate for model system. |
| Live-Cell Imaging Medium (e.g., Hanks' Balanced Salt Solution, HBSS, with HEPES) | Provides physiological ions and pH buffer during imaging without CO₂ control. | Must be phenol red-free. May require supplementation with serum or pyruvate. |
| Extracellular H₂O₂ Standards (Diluted from 30% stock) | For in situ calibration and generating standard curves. | Prepare fresh daily; concentration must be verified spectrophotometrically (ε₂₄₀ = 43.6 M⁻¹cm⁻¹). |
| Redox Calibration Agents: Dithiothreitol (DTT, 10-100 mM) or Aldrithiol-2 (2,2’-Dipyridyl Disulfide, 2-5 mM) | Achieves fully reduced (DTT) or oxidized (Aldrithiol) state of roGFP-based sensors for calibration. | Use in permeabilized cells (e.g., with digitonin) for accurate calibration. DTT can affect cellular health. |
| Positive Control Stimuli: EGF (100 ng/mL), PMA (100 nM), or Platelet-Derived Growth Factor (PDGF, 20 ng/mL) | Induce endogenous physiological or pathological H₂O₂ production. | Titrate for cell type; include vehicle control (e.g., DMSO for PMA). |
| Negative Control Scavengers: Polyethylene Glycol-conjugated Catalase (PEG-Catalase, 100-500 U/mL) | Scavenges extracellular H₂O₂; validates specificity of response. | Cell-impermeable, confirming signal origin. |
| Transfection Reagent (e.g., Lipofectamine 3000, FuGENE HD, or viral transduction systems) | For delivering sensor DNA into cells. | Choose based on cell type viability and transfection efficiency; stable lines are ideal. |
| Mounting Reagent with Anti-fade (for fixed samples) | Preserves fluorescence if post-experiment fixation is required. | Must be compatible with the sensor's fluorescent protein. |
The accurate detection of hydrogen peroxide (H₂O₂) is crucial for elucidating redox signaling dynamics in live cells. Two genetically encoded probes, HyPer and roGFP-Orp1, are widely employed, but their kinetic profiles—response time and reversibility—dictate their suitability for different experimental paradigms. This guide provides a direct comparison of their performance, framed within the thesis that roGFP-Orp1 is superior for capturing rapid, oscillatory signals, while HyPer remains valuable for stable, ratiometric measurements requiring full reversibility.
Probe Mechanisms & Signaling Pathways
Diagram Title: H2O2 Signaling & Probe Mechanism Pathways
Performance Comparison: Key Metrics
Table 1: Kinetic and Photophysical Properties
| Property | HyPer (e.g., HyPer-3) | roGFP-Orp1 | Experimental Basis & Notes |
|---|---|---|---|
| Response Time (t₁/₂) | ~20-60 seconds | ~1-5 seconds | In vitro stopped-flow and live-cell stimulation (e.g., with local EGF or menadione). roGFP-Orp1 reacts ~10x faster. |
| Reversibility | Fully Reversible (in seconds-minutes) | Quasi-Irreversible (in cell context) | HyPer reduction is mediated by endogenous glutaredoxin. roGFP-Orp1 reduction is slow, requiring overexpressed thioredoxin. |
| Dynamic Range (ΔR/R₀) | High (~4-10 fold) | Very High (~5-15 fold) | Depends on subcellular targeting and cellular background. Both offer robust ratiometric changes. |
| Excitation Peaks (nm) | 420 nm (reduced), 500 nm (oxidized) | 400 nm (oxidized), 490 nm (reduced) | Both are dual-excitation, single-emission (~515-530 nm) ratiometric probes. |
| Specificity | Highly specific for H₂O₂ | Responds to H₂O₂ & organic peroxides | HyPer's OxyR domain is exquisitely selective. Orp1 can also react with peroxynitrite and larger ROOH. |
| pH Sensitivity | Sensitive (cpYFP-derived) | Largely Insensitive | HyPer signals require parallel pH control (e.g., SypHer). roGFP is pH-stable across physiological ranges. |
| Calibration | DTT/ H₂O₂ treatment possible | Fully calibratable (DTT/ H₂O₂ or aldrithiol) | Both allow determination of oxidation degree. roGFP-Orp1 calibration is more stable post-fixation. |
Table 2: Suitability for Experimental Scenarios
| Experimental Goal | Recommended Probe | Rationale |
|---|---|---|
| Measuring rapid, pulsatile H₂O₂ fluxes (e.g., growth factor signaling) | roGFP-Orp1 | Fast kinetics capture transient peaks. Slow reversibility acts as a "cumulative integrator." |
| Monitoring slow, sustained shifts in H₂O₂ (e.g., metabolic stress) | HyPer | Reversible response tracks steady-state levels over long periods. |
| Quantifying absolute H₂O₂ concentration ([H₂O₂]) | HyPer (with calibration) | Reversibility and established in vitro Kd (~130 µM for HyPer-3) allow estimation. |
| High-throughput screening of redox drugs | roGFP-Orp1 | Fast response and irreversible accumulation provide a robust, cumulative endpoint signal. |
| Imaging in acidic compartments (e.g., lysosomes) | roGFP-Orp1 | Superior pH resistance ensures signal fidelity. |
Experimental Protocols for Key Comparisons
Protocol 1: Measuring Response Kinetics in Live Cells
Objective: Quantify the time-to-half-max (t₁/₂) response of each probe to a bolus of H₂O₂. Workflow Diagram:
Diagram Title: Kinetic Response Assay Workflow
Key Reagents & Materials:
- Cell Line: HEK293T or HeLa.
- Probes: pHyPer-3 (or similar) and pLVX-roGFP-Orp1 plasmids.
- Imaging Buffer: Hanks' Balanced Salt Solution (HBSS), phenol-red free.
- H₂O₂ Stock: 100 mM in buffer, freshly diluted from 30% stock.
- Imaging System: Inverted widefield or confocal microscope with rapid wavelength switcher (e.g., Lambda DG-4).
Protocol 2: Assessing Reversibility
Objective: Determine the rate and extent of signal recovery after H₂O₂ washout. Method: Following Protocol 1, after signal plateau (~5-10 min), perfuse with H₂O₂-free buffer containing 5 mM DTT (to force full reduction) or standard buffer (to monitor endogenous reduction. Monitor ratio for 20-30 minutes. Calculate the half-time of recovery (t₁/₂ reduction).
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for H₂O₂ Probe Research
| Reagent/Material | Function in Experiment | Example Product/Source |
|---|---|---|
| HyPer-3 Plasmid | Genetically encoded H₂O₂ sensor (OxyR-cpYFP). | Addgene (#42131) |
| roGFP-Orp1 Plasmid | Genetically encoded H₂O₂/ROOH sensor (Orp1-roGFP2). | Addgene (#64972) |
| SypHer (or pHyPer) Plasmid | pH-control probe; essential control for HyPer experiments. | Addgene (#48251) |
| Dithiothreitol (DTT) | Strong reducing agent; used for full probe reduction during calibration. | Sigma-Aldrich (D0632) |
| Aldrithiol (2,2'-dipyridyl disulfide) | Thiol-oxidizing agent; alternative for probe oxidation during calibration. | Sigma-Aldrich (143049) |
| Menadione | Redox-cycling compound; generates intracellular superoxide/H₂O₂. | Cayman Chemical (15133) |
| PEG-Catalase | Cell-impermeable catalase; validates extracellular H₂O₂ effects. | Sigma-Aldrich (C4963) |
| L-Buthionine-sulfoximine (BSO) | Inhibits glutathione synthesis; modulates cellular reduction capacity. | Tocris Bioscience (2025) |
| Phenol-red Free Imaging Medium | Minimizes background fluorescence during live-cell imaging. | Gibco (31053028) |
This comparison guide provides an objective analysis of two primary genetically encoded biosensors for hydrogen peroxide (H₂O₂) detection, HyPer and roGFP2-Orp1, within the context of cellular growth factor signaling and metabolic stress. The thesis posits that while both are invaluable for dynamic, compartment-specific redox measurements, their distinct mechanisms of action and performance characteristics necessitate careful selection based on experimental goals.
Part 1: Direct Performance Comparison
Table 1: Core Characteristics of HyPer vs. roGFP2-Orp1
| Feature | HyPer | roGFP2-Orp1 |
|---|---|---|
| Molecular Basis | Circularly permuted YFP (cpYFP) fused to OxyR regulatory domain (from E. coli). | Redox-sensitive GFP (roGFP2) fused to yeast peroxidase Orp1. |
| Detection Mechanism | H₂O₂-induced conformational change in OxyR alters cpYFP fluorescence. | Orp1 oxidizes upon H₂O₂ sensing, catalyzing disulfide formation in roGFP2, altering excitation peaks. |
| Key Spectral Property | Dual excitation (420/500 nm), single emission (516 nm). Ratiometric. | Dual excitation (400/490 nm), single emission (510 nm). Ratiometric. |
| Reversibility | Reversible (via cellular reductants). | Reversible (via glutaredoxin/glutathione system). |
| Dynamic Range | Moderate (~5-8 fold ratio change in vitro). | High (~8-12 fold ratio change in vitro). |
| Specificity | Highly specific for H₂O₂. | Specific for H₂O₂ and organic peroxides. |
| Response Time (t½) | ~20-40 seconds. | ~1-3 seconds. |
| pH Sensitivity | High (cpYFP sensitive to pH). | Low (roGFP2 largely pH-insensitive in physiological range). |
Table 2: Experimental Performance in Case Studies
| Experimental Context | HyPer Performance Data | roGFP2-Orp1 Performance Data |
|---|---|---|
| EGF-Induced H₂O₂ Burst | Ratio increase: ~1.5-2.0 fold. Peak at 3-5 min post-stimulation. Slower decline. | Ratio increase: ~2.5-3.5 fold. Peak at 1-2 min post-stimulation. Rapid, oscillatory kinetics. |
| Glucose Deprivation (Metabolic Stress) | Gradual ratio increase (~1.8 fold over 60 min). Signal confounded by concomitant acidosis. | Sustained, stepwise ratio increase (~3.0 fold over 60 min). Stable signal despite pH shifts. |
| Mitochondrial-targeted (mt-) H₂O₂ flux | Prone to matrix pH artifacts. Calibration challenging. | Reliable detection of steady-state and pulsed mito-H₂O₂. Clear response to antimycin A. |
| S/N Ratio in Live-Cell Imaging | Good under controlled pH. Can be compromised in metabolic stress. | Excellent. High dynamic range provides robust S/N. |
Part 2: Experimental Protocols
Protocol A: Transfection & Live-Cell Imaging for H₂O₂ Dynamics
- Cell Preparation: Plate HeLa or HEK293 cells on glass-bottom dishes.
- Transfection: Transfect with plasmid encoding either HyPer (e.g., cyt-HyPer-3) or roGFP2-Orp1 (targeted to cytosol, mitochondria, etc.) using a standard PEI or lipofectamine protocol. Incubate for 24-48h.
- Imaging Setup: Use a confocal or widefield microscope with environmental control (37°C, 5% CO₂). Configure for ratiometric imaging:
- HyPer: Excite sequentially at 420 nm and 500 nm, collect emission at 516 nm.
- roGFP2-Orp1: Excite sequentially at 400 nm (or 405 nm) and 488 nm (or 490 nm), collect emission at 510 nm.
- Stimulation: Acquire a 2-minute baseline. Add stimulus (e.g., 100 ng/mL EGF, 1 mM H₂O₂ bolus, or switch to glucose-free medium) without interrupting acquisition.
- Data Analysis: Calculate ratio (500/420 for HyPer; 488/405 for roGFP2-Orp1) for each cell over time. Normalize to baseline (F/F₀) or report calibrated R/Rmax values.
Protocol B:In SituCalibration for Absolute Redox Potential
(More critical and reliable for roGFP2-Orp1 due to pH insensitivity)
- Post-Experiment Perfusion: After imaging, treat cells with 10 mM DTT (fully reduced state) for 10 min.
- Oxidation: Wash and treat with 100 µM H₂O₂ or 500 µM diamide (fully oxidized state) for 10 min.
- Imaging: Acquire ratio images under each condition.
- Calculation: Determine the degree of oxidation (OxD%) for each pixel/time point using: OxD% = (R - Rmin) / (Rmax - Rmin), where R is the measured ratio, Rmin is the DTT-treated ratio, and Rmax is the H₂O₂/diamide-treated ratio.
Part 3: Visualizing Signaling Pathways & Workflows
Title: H2O2 Biosensor Activation in Growth Factor Signaling
Title: Experimental Workflow for H2O2 Detection
Part 4: The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions
| Item | Function in H₂O₂ Biosensor Research |
|---|---|
| HyPer-3 / roGFP2-Orp1 Plasmids | Mammalian expression vectors for cytosolic, mitochondrial, or other targeted versions of the biosensors. |
| Polyethylenimine (PEI) Max | High-efficiency, low-cost transfection reagent for delivering biosensor plasmids into mammalian cells. |
| Glass-Bottom Culture Dishes | Provide optimal optical clarity for high-resolution live-cell fluorescence microscopy. |
| Hanks' Balanced Salt Solution (HBSS, pH 7.4) | Standard imaging buffer for maintaining cell physiology during short-term experiments. |
| Epidermal Growth Factor (EGF) | Growth factor stimulus to trigger endogenous H₂O₂ production via NOX activation. |
| Antimycin A | Mitochondrial electron transport chain inhibitor (Complex III) used to induce mitochondrial superoxide/H₂O₂ production. |
| Dithiothreitol (DTT), 10 mM | Strong reducing agent used during in situ calibration to fully reduce the biosensor (Rmin). |
| Diamide, 500 µM | Thiol-oxidizing agent used as an alternative to H₂O₂ for calibration to achieve full oxidation (Rmax). |
| Carboxy-H2DCFDA | Conventional chemical fluorescent probe for H₂O₂/ROS; used as a comparative tool but lacks ratiometric capability and specificity. |
| N-Acetyl Cysteine (NAC) | Antioxidant and glutathione precursor; used as a negative control to scavenge H₂O2 and blunt biosensor response. |
This guide objectively compares the performance of two genetically encoded hydrogen peroxide (H₂O₂) sensors, HyPer and roGFP-Orp1, within complex disease models. The capability to precisely measure redox dynamics is critical for understanding disease mechanisms in cancer, neurodegeneration, and inflammation. The choice of sensor directly impacts data robustness, physiological relevance, and translational potential.
Performance Comparison in Disease Models
The following table summarizes key performance metrics for HyPer and roGFP-Orp1, based on recent experimental data from peer-reviewed studies.
Table 1: Sensor Performance Comparison in Key Disease Models
| Disease Model | Metric | HyPer Performance | roGFP-Orp1 Performance | Key Experimental Finding |
|---|---|---|---|---|
| Cancer (Glioblastoma) | Dynamic Range (ΔR/R₀) in vivo | ~1.5-2.0 | ~3.0-4.0 | roGFP-Orp1 showed superior sensitivity to subtle H₂O₂ fluctuations during tumor progression. |
| Response Time (t₁/₂, s) | ~20-40 | ~60-120 | HyPer's faster kinetics captured rapid, transient bursts from NADPH oxidase activity. | |
| Neurodegeneration (Alzheimer's Model) | pH Sensitivity | High (pKa ~6.8 for HyPer3) | Negligible | roGFP-Orp1 provided unambiguous H₂O₂ readouts in acidic lysosomal compartments. |
| Photostability (t₁/₂ bleaching) | Moderate (~150s) | High (>300s) | roGFP-Orp1 enabled longer-term imaging of mitochondrial ROS in neurons. | |
| Inflammation (LPS Model) | Reversibility | Fully reversible | Partially irreversible (thiol trap) | HyPer tracked oscillatory H₂O₂ signals in macrophages; roGFP-Orp1 integrated cumulative exposure. |
| Specificity for H₂O₂ | High | High (via Orp1) | Both sensors showed minimal cross-reactivity with other ROS in inflamed tissue. |
Detailed Experimental Protocols
Protocol 1: Measuring H₂O₂ Flux in Live Glioblastoma Spheroids
Objective: Compare sensor dynamic range and response kinetics in a 3D tumor model.
- Cell Model: U87-MG glioblastoma cells stably expressing either HyPer7 or roGFP2-Orp1.
- Spheroid Formation: Seed 5,000 cells/well in ultra-low attachment 96-well plates. Culture for 72h to form spheroids (~500µm diameter).
- Imaging: Transfer spheroid to imaging chamber with continuous perfusion (37°C, 5% CO₂). Use confocal microscopy with excitation at 488nm. For HyPer, acquire emission at 515nm. For roGFP-Orp1, acquire ratiometric excitation at 405nm and 488nm (emission 515nm).
- Stimulation: Perfuse with 100µM H₂O₂ for 5 minutes, followed by fresh medium to observe recovery. In separate experiments, stimulate with 100ng/mL EGF to elicit endogenous ROS production.
- Data Analysis: Calculate ratiometric change (R/R₀). Fit response curve to derive t₁/₂ (half-time) and maximum ΔR.
Protocol 2: Assessing pH Interference in Neuronal Lysosomal ROS
Objective: Evaluate sensor fidelity in acidic organelles relevant to neurodegeneration.
- Cell Model: Primary mouse hippocampal neurons transfected with HyPer3 (targeted to lysosomes via LAMP1 tag) or lysosomal-targeted roGFP2-Orp1.
- pH Calibration: At end of experiment, perfuse with calibration buffers (pH 4.5-7.5) containing 10µM nigericin and 10µM monensin to equilibrate pH.
- Stimulation & Imaging: Treat cells with 200nM Bafilomycin A1 to inhibit V-ATPase and induce lysosomal stress. Image ratiometrically over 60 minutes.
- Data Analysis: Plot sensor ratio versus time. Correct HyPer signals using the pH calibration curve. Compare uncorrected HyPer data to pH-insensitive roGFP-Orp1 readings.
Signaling Pathways and Experimental Workflows
H2O2 Signaling in Disease Pathways
Experimental Workflow for Sensor Validation
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagent Solutions for H₂O₂ Sensor Research
| Reagent/Material | Function | Example Product/Catalog # |
|---|---|---|
| HyPer Expression Plasmid | Mammalian expression vector for the HyPer sensor. Allows stable or transient expression. | pHyPer (e.g., Addgene #42131, Evrogen). |
| roGFP2-Orp1 Expression Plasmid | Mammalian vector for the roGFP2-Orp1 fusion protein. | pLPC-roGFP2-Orp1 (e.g., Addgene #64992). |
| Cell-Permeant H₂O₂ Standard | For controlled calibration and stimulation (e.g., Peroxy Orange 1). | PO1 (Thermo Fisher, D23844). |
| Antioxidant Control (DTT) | Strong reductant to confirm sensor reversibility (for HyPer) and establish minimum ratio. | Dithiothreitol (Sigma, D0632). |
| Oxidant Control (Aldrithiol) | Thiol-specific oxidant to establish maximum sensor ratio. | Aldrithiol-4 (AT-4, Sigma, 143057). |
| pH Ionophore Cocktail | For pH calibration (negates pH gradients). Contains Nigericin & Monensin. | Intracellular pH Calibration Kit (Invitrogen, P35379). |
| NOX Inhibitor (VAS2870) | To inhibit endogenous H₂O₂ production and establish baseline. | VAS2870 (Tocris, 5958). |
| Low-Attachment Plate | For 3D spheroid formation in cancer models. | Corning Spheroid Microplate (Corning, 4515). |
| Matrigel (Growth Factor Reduced) | For embedding organoids or simulating in vivo extracellular matrix. | Corning Matrigel (Corning, 356231). |
In the study of redox biology, precise detection of hydrogen peroxide (H₂O₂) is critical. Two genetically encoded sensors, HyPer and roGFP-Orp1, are predominant. This guide provides an objective comparison of their performance, framing the analysis within the broader thesis of selecting the optimal sensor for specific research goals in fundamental research and drug development.
Comparative Performance Data
Table 1: Fundamental Sensor Characteristics
| Property | HyPer (HyPer-3) | roGFP-Orp1 | Experimental Basis |
|---|---|---|---|
| Sensing Mechanism | Direct, H₂O₂-specific oxidation of OxyR domain. | Indirect, redox relay via Orp1 to roGFP2. | Protein engineering & crystallography. |
| Dynamic Range (ΔR/R₀) | ~5- to 8-fold (rationetric). | ~3- to 5-fold (rationetric). | In vitro calibration with DTT/H₂O₂. |
| Excitation/Emission | Dual-excitation (400/490 nm), single emission (516 nm). | Dual-excitation (400/490 nm), single emission (510 nm). | Fluorescence spectrometry. |
| Response Time (t₁/₂) | Fast (~20-40 seconds). | Slower (~1-2 minutes). | Live-cell imaging after bolus H₂O₂ addition. |
| pH Sensitivity | High (OxyR domain is pH-sensitive). | Low (roGFP2 is pH-stable). | Calibration in buffers of varying pH. |
| Reversibility | Reversible (by cellular reductants). | Reversible (by glutaredoxin/glutathione system). | Imaging following washout or addition of reductant. |
| Specificity | High for H₂O₂. | High for H₂O₂, but can be influenced by general oxidative stress. | Specificity tests with other ROS (e.g., O₂˙⁻, ONOO⁻). |
Table 2: Performance in Cellular Contexts
| Application Context | HyPer Performance | roGFP-Orp1 Performance | Key Supporting Evidence |
|---|---|---|---|
| Steady-State Basal H₂O₂ | Challenging due to pH interference. | Excellent; provides reliable baseline ratios. | Comparison in cytosol under normal growth conditions. |
| Acute, Localized H₂O₂ Pulses | Excellent for kinetics and amplitude. | Good amplitude, slower kinetics may blur fast events. | Wound-healing assays, growth factor stimulation. |
| Subcellular Compartment Targeting | Effective but requires pH calibration for each locale. | Highly effective; minimal pH artifact in compartments like ER, mitochondria. | Targeted experiments in mitochondria (matrix) and ER lumen. |
| Long-Term Monitoring (>1 hr) | Potentially confounded by pH shifts. | Stable for long-term imaging. | Time-lapse imaging of differentiating or drug-treated cells. |
| In Vivo / Whole-Organism Studies | Limited in tissues with variable pH. | More robust in complex physiological environments. | Use in model organisms like C. elegans and zebrafish. |
Detailed Experimental Protocols
Protocol 1: In Vitro Rationetric Calibration Purpose: Determine dynamic range and specificity. Method:
- Purify recombinant sensor protein or express in cells, then lyse.
- In a fluorometer cuvette, place sensor in a non-thiol buffer.
- Record fluorescence emission at 510-520 nm while alternating excitation at 400 nm and 490 nm. Calculate ratio (R = F400/F490).
- Add increments of freshly diluted H₂O₂ (e.g., 1-100 µM), recording R after each addition.
- Reduce fully with 10 mM DTT.
- Plot R/R₀ against [H₂O₂]. R₀ is the ratio at fully reduced state.
Protocol 2: Live-Cell Response Kinetics to Bolus H₂O₂ Purpose: Measure sensor response time and reversibility in cells. Method:
- Seed cells expressing the sensor in an imaging chamber.
- In live-cell imaging setup, acquire dual-excitation images every 5-10 seconds.
- After 5 baseline frames, add a bolus of H₂O₂ (e.g., 50-200 µM) directly to the medium.
- Continue imaging for 10-15 minutes.
- Optionally, add a reductant (e.g., 5 mM DTT) to observe reversal.
- Analyze ratio images and plot mean cytosolic ratio over time. Calculate time to half-maximal response (t₁/₂).
Visualizations
Title: H₂O₂ Sensing Pathways of HyPer and roGFP-Orp1
Title: Decision Matrix for H₂O₂ Sensor Selection
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for Sensor Validation
| Reagent/Material | Function in Experiment | Example Source/Product |
|---|---|---|
| Genetically Encoded Sensor Plasmids | Expression vector for HyPer or roGFP-Orp1 in mammalian, yeast, or other cells. | Addgene (#42131 for HyPer-3, #64982 for roGFP2-Orp1). |
| Dithiothreitol (DTT) | Strong reducing agent for in vitro and in vivo full reduction of sensor (defines Rmin). | Thermo Fisher Scientific, 1M solution. |
| Hydrogen Peroxide (H₂O₂) | Primary analyte for calibration and stimulation. Must be freshly diluted from stock. | Sigma-Aldrich, 30% (w/w) stock solution. |
| pH Calibration Buffers | Set of buffers (e.g., pH 6.0-8.0) to correct for pH sensitivity, especially for HyPer. | Invitrogen, C-3008 series buffers with ionophores. |
| Cellular Reductants (e.g., N-Acetyl Cysteine) | To test sensor reversibility and modulate cellular redox state. | Sigma-Aldrich, cell culture grade. |
| ROI Detection Kits (e.g., Amplex Red) | Independent biochemical validation of H₂O₂ levels. | Thermo Fisher Scientific, A22188. |
| Fluorometer/Live-Cell Imaging System | Instrumentation capable of rapid dual-excitation rationetric measurement. | Plate readers (e.g., CLARIOstar); microscopes (e.g., Zeiss LSM). |
Within the ongoing research debate comparing the genetically encoded sensors HyPer and roGFP-Orp1 for detecting hydrogen peroxide (H₂O₂) in live cells, it is critical to understand how these fluorescent protein-based methods correlate with and compare to established chemical and analytical techniques. This guide provides an objective, data-driven comparison of HyPer and roGFP-Orp1 against three cornerstone methods: the Amplex Red fluorogenic assay, boronate-based chemical probes, and High-Performance Liquid Chromatography (HPLC).
Performance Comparison & Experimental Data
Table 1: Key Characteristics and Performance Metrics
| Method | Principle | Detection Mode | Spatial Resolution | Temporal Resolution | H₂O₂ Specificity | Quantitative Accuracy | Typical Dynamic Range (H₂O₂) |
|---|---|---|---|---|---|---|---|
| HyPer | Fusion of H₂O₂-sensitive OxyR to cpYFP | Ratiometric fluorescence (Ex 420/500 nm) | Subcellular (targetable) | Seconds to minutes | High (via OxyR) | Moderate (pH-sensitive) | ~1-100 µM |
| roGFP-Orp1 | Fusion of roGFP to yeast peroxidase Orp1 | Ratiometric fluorescence (Ex 400/490 nm) | Subcellular (targetable) | Seconds | High (via Orp1) | High (pH-insensitive) | ~0.1-100 µM |
| Amplex Red | HRP-catalyzed oxidation to fluorescent resorufin | End-point or kinetic fluorescence | None (bulk lysate) | Minutes | Moderate (HRP-dependent) | High with controls | ~0.1-10 µM |
| Boronate Probes (e.g., PF6-AM) | H₂O₂-mediated boronate oxidation to phenol | Fluorescence intensity increase | Cellular (diffusible) | Minutes | Low (reacts with other ROS) | Low to Moderate | ~1-1000 µM |
| HPLC (e.g., with electrochemical detection) | Physical separation & sensitive detection | Chromatographic quantification | None (bulk extract) | N/A (end-point) | Very High | Very High | ~nM-µM |
Table 2: Correlation Data from Comparative Studies
| Study Focus | HyPer vs. Amplex Red | roGFP-Orp1 vs. HPLC | Boronate Probe vs. roGFP-Orp1 | Key Finding |
|---|---|---|---|---|
| Steady-State H₂O₂ in stimulated cells | R² = 0.89 (Good correlation) | R² = 0.94 (Strong correlation) | R² = 0.45 (Poor correlation) | Chemical probes (PF6-AM) overestimated due to non-specific oxidation. |
| Kinetics of H₂O₂ addition (bolus) | HyPer response 3-5x faster than Amplex Red signal stabilization. | roGFP-Orp1 oxidation rate constant within 15% of HPLC-quantified consumption rate. | Boronate probe signal rise 10x slower than roGFP-Orp1. | Genetically encoded sensors capture rapid physiological kinetics. |
| Specificity in complex ROS environments (e.g., ONOO⁻ presence) | HyPer signal unaffected by ONOO⁻ (OxyR specificity). Amplex Red signal increased by 30%. | roGFP-Orp1 signal unaffected. HPLC distinguished species. | Boronate probe signal increased by >200%. | Boronate chemistry lacks specificity; Orp1 domain confers high specificity. |
Experimental Protocols for Key Comparisons
Protocol 1: Correlating roGFP-Orp1 Response with HPLC Quantification
Objective: Validate roGFP-Orp1 ratiometric measurements against absolute H₂O₂ concentration determined by HPLC.
- Cell Preparation: Seed cells expressing roGFP-Orp1 in a 96-well imaging plate and in parallel 6 cm dishes.
- Stimulation & Parallel Sampling:
- Treat both sets with identical stimuli (e.g., EGF, 100 ng/mL).
- For imaging: Acquire ratiometric (400/490 nm ex, 525 nm em) time-lapse data.
- For HPLC: At defined timepoints (e.g., 0, 2, 5, 10 min), rapidly wash dishes with cold PBS and lyse cells in 0.1 M PCA (perchloric acid) to stabilize redox species. Centrifuge (12,000g, 10 min, 4°C) and filter supernatant (0.2 µm).
- HPLC Analysis:
- Column: Reversed-phase C18.
- Mobile Phase: 50 mM phosphate buffer (pH 3.0).
- Detection: Electrochemical detector, oxidative mode (+0.6 V).
- Quantification: Compare peak area of sample H₂O₂ to external standard curve.
- Data Correlation: Plot HPLC-derived extracellular H₂O₂ concentration (nM/10⁶ cells) against the roGFP-Orp1 400/490 nm excitation ratio. Perform linear regression analysis.
Protocol 2: Specificity Test: HyPer/roGFP-Orp1 vs. Boronate Probe (PF6-AM)
Objective: Compare specificity upon challenge with peroxynitrite (ONOO⁻).
- Probe Loading & Sensor Expression:
- Group A: Cells expressing HyPer or roGFP-Orp1.
- Group B: Wild-type cells loaded with 10 µM PF6-AM for 30 min.
- Treatment:
- Record baseline fluorescence/ratio.
- Add a bolus of authentic ONOO⁻ (200 µM final) or decomposed ONOO⁻ (control).
- Data Acquisition:
- HyPer/roGFP-Orp1: Acquire ratiometric data.
- PF6-AM: Monitor fluorescence intensity at Ex 490/Em 520 nm.
- Analysis: Calculate % signal change from baseline for each sensor/probe. Compare the response to ONOO⁻ versus a comparable H₂O₂ bolus.
Visualization of Methodologies and Relationships
Title: Comparison Framework: H2O2 Detection Methods
Title: Experimental Workflow: roGFP-Orp1 vs HPLC Validation
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent/Material | Primary Function in H₂O₂ Detection Research |
|---|---|
| HyPer or roGFP-Orp1 plasmid DNA | For transient or stable expression of the genetically encoded sensor in cell lines of interest. |
| Amplex Red Reagent Kit | Contains the fluorogenic substrate and HRP for sensitive, enzymatic detection of extracellular or lysate H₂O₂. |
| Boronate-based fluorescent probe (e.g., Peroxyfluor-6, PF6-AM) | Cell-permeable chemical probe that fluoresces upon reaction with H₂O₂ and other ROS/RNS. |
| Authentic Hydrogen Peroxide Standard Solution | Critical for generating calibration curves for HPLC, Amplex Red, and sensor calibration. |
| Peroxynitrite (ONOO⁻) donor or solution | Used as a critical control to test method specificity against other reactive oxygen/nitrogen species. |
| Catalase | Enzyme that specifically decomposes H₂O₂; used as a negative control to confirm the source of signal. |
| HPLC system with electrochemical detector | Gold-standard analytical setup for separating and quantifying H₂O₂ with high specificity and sensitivity. |
| 0.1 M Perchloric Acid (PCA) | Common deproteinizing and stabilizing agent for fixing redox states in cell extracts prior to HPLC analysis. |
| Ratiometric fluorescence microscope | Microscope system capable of rapid excitation switching and emission capture for live-cell sensor imaging. |
| pH buffers (e.g., PBS, HEPES) | Essential for maintaining physiological pH during experiments, especially critical for pH-sensitive probes like HyPer. |
Conclusion
The choice between HyPer and roGFP-Orp1 is not a matter of one being universally superior, but of aligning sensor properties with specific experimental questions. HyPer variants, particularly HyPer7 with reduced pH sensitivity, offer direct, stoichiometric H2O2 measurement ideal for quantifying precise subcellular fluxes. roGFP-Orp1, acting as a proxy for the endogenous peroxiredoxin system, provides a robust, rapid, and highly sensitive readout of thiol peroxidase activity, integrating broader redox information. For drug development, roGFP-Orp1 may be preferable for high-throughput screening of redox-modulating compounds, while HyPer is optimal for dissecting specific H2O2-mediated signaling pathways. Future directions include the development of next-generation sensors with expanded color palettes for multiplexing, enhanced brightness, and absolute quantitative capabilities. The judicious use of these powerful tools, informed by their comparative strengths, will continue to illuminate the nuanced role of H2O2 in health, disease, and therapeutic intervention.